Abstract
Purpose:
Krabbe disease (KD) results from galactocerebrosidase (GALC) deficiency. Infantile KD symptoms include irritability, progressive stiffness, developmental delay, and death. The only potential treatment is hematopoietic stem cell transplantation. New York State (NYS) implemented newborn screening for KD in 2006.
Methods:
Dried blood spots from newborns were assayed for GALC enzyme activity using mass spectrometry, followed by molecular analysis for those with low activity (≤12% of the daily mean). Infants with low enzyme activity and one or more mutations were referred for follow-up diagnostic testing and neurological examination.
Results:
Of >1.9 million screened, 620 infants were subjected to molecular analysis and 348 were referred for diagnostic testing. Five had enzyme activities and mutations consistent with infantile KD and manifested clinical/neurodiagnostic abnormalities. Four underwent transplantation, two are surviving with moderate to severe handicaps, and two died from transplant-related complications. The significance of many sequence variants identified is unknown. Forty-six asymptomatic infants were found to be at moderate to high risk for disease.
Conclusions:
The positive predictive value of KD screening in NYS is 1.4% (5/346) considering confirmed infantile cases. The incidence of infantile KD in NYS is approximately 1 in 394,000, but it may be higher for later-onset forms.
Genet Med 18 3, 239–248.
Similar content being viewed by others
Introduction
Krabbe disease (KD; globoid cell leukodystrophy; OMIM 245200) is a rare, inherited lysosomal storage disorder (LSD) caused by deficiency of galactocerebrosidase (GALC), an enzyme essential for myelin turnover. Infantile KD comprises early infantile KD (EIKD; symptoms within 6 months of age) and late infantile KD (LIKD; symptom onset between 6 and 12 months) forms that reportedly account for 90% of patients.1,2,3 Early symptoms include irritability, unexplained fever, limb stiffness, seizures, feeding difficulties, reflux, and slowing of mental and motor development. Later symptoms include muscle weakness, spasticity, deafness, and blindness.2,4,5 Infants deteriorate and are typically severely compromised before age 2 years, requiring artificial feeding and constant care, with death during childhood. Individuals with later-onset forms of KD (symptoms after 1 year of age) and a more heterogeneous clinical phenotype have been described,6,7,8 and some exhibit a rapidly declining course.5,9
More than 140 disease-causing mutations and polymorphisms in the GALC gene have been reported,2 but most are rare. A 30-kb deletion (also called 502T/del) is the most common in populations of European ancestry, occurring in ~40% of infantile-onset KD alleles.2,10 Two point mutations (p.T513M and p.Y551S) are reported to be responsible for ~10–15% of the remaining infantile KD alleles in Europeans.2,11 Another, p.G270D, comprises a large proportion of alleles in patients with later-onset KD.2 Three common polymorphisms (p.R168C, p.D232N, p.I546T) attenuate GALC activity, complicating clinical interpretation of low enzyme activity, but do not cause disease (pseudodeficiency).2
There is growing interest in newborn screening (NBS) for LSDs,12,13,14,15,16,17 primarily because treatment is thought to be most effective when initiated before symptom onset. A multiplex tandem mass spectrometry (MS/MS) assay for five LSDs (KD, Gaucher, Pompe, Niemann-Pick, and Fabry diseases) was developed13 and optimized for use on dried blood spots.15,18,19 There are challenges to adding LSDs to NBS panels, including the need for expensive instrumentation and adequate quality control materials (especially dried blood spots from infants with KD),14,20 diagnostic laboratories for confirmatory testing with short turnaround times, infrastructure for rapid referral and urgent transplantation in newborns with EIKD, protocols for follow-up, and clinicians experienced in interpreting and relaying the full spectrum of NBS results, including variability in enzyme activity and identification of sequence variants of unknown significance or variable expressivity.21 Early studies showed that a small cohort of infantile KD patients who received umbilical cord blood transplantation from unrelated donors prior to symptom onset had preserved neurological function.22,23,24 However, even when transplantation is performed early, delays in development of gross motor skills have developed in most patients.22 It is critical that families be fully educated about benefits, risks, and limitations of treatment following a positive screen.
New York State (NYS) was mandated to implement NBS for KD in August 2006.25 Missouri has recently begun screening,26 and several other states have pending mandates to screen KD and/or other LSDs.16 Here, we describe our two-tiered enzyme and molecular screening assay and provide results from the first 1.9 million infants screened over the course of 8 years.
Materials and Methods
GALC activity is quantified from dried blood spots using synthetic substrate and analyzed by MS/MS.25 Activity (µmol/hour/l) is converted to percent activity based on the daily mean activity (DMA). Specimens with ≤20% DMA are retested in duplicate using new punches from the original dried blood spots, and specimens with mean GALC ≤12% DMA (initial plus two repeat punches) are subjected to molecular GALC analysis. Infants with at least one potentially clinically relevant variant/mutation are considered screen-positive and referred to a NYS-accredited inherited metabolic disease Specialty Care Center for diagnostic testing, genetic counseling, and clinical/neurological examination.21 Details regarding GALC enzyme and molecular assays, the diagnostic assay, and follow-up procedures have been published21,22,25 or are included in Supplementary Methods and Supplementary Tables S1 and S2 online. Specimens collected from infants born during the first 8 years of screening are included in this study (through 7 August 2014). Infants who were born earlier but had a repeat specimen collected during this period are also included.
Results
Screening algorithm tier 1: GALC activity
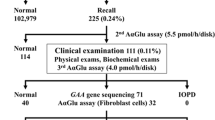
As of 7 August 2014, 2,090,910 specimens from 1,968,568 infants had been screened using the two-tiered NYS KD algorithm. More than 99.9% of all specimens tested were screen-negative ( Table 1 ). Overall, 10,199 specimens (4.9 per 1,000) with ≤20% GALC activity were retested in duplicate, and 620 (0.3 per 1,000) were confirmed to have average activity of ≤12% and were reflexed to molecular analysis. Screen-positive results were obtained for 348 (0.17 per 1,000) infants who were referred to a Specialty Care Center for diagnostic workup.
Table 2 shows GALC activity overall and stratified by screening results, specimen type, and demographic data collected over a representative sample (first 5 years of NBS). There were differences in enzyme activity by age at specimen collection, gender, and race/ethnic group. Activity was markedly higher (11.54 ± 9.88) in specimens taken within the first 24 hours of life (“day of birth (DOB) specimens”; primarily collected from premature or sick infants) compared with those collected from infants at least 24 hours of age (4.01 ± 2.78). Per standard practice, repeat specimens are requested for all DOB specimens; however, if GALC activity in a DOB specimen was ≤12% and at least one mutation was detected, then the infant would be referred. To date, no DOB specimens have had GALC activity ≤12%.
Screening algorithm tier 2: molecular GALC analysis
Infants with low GALC activity (≤12%) were screened for GALC variants by complete sequencing and deletion analysis (30- and 7-kb).
Infants with polymorphisms only
Overall, 272 of 620 infants with low enzyme activity (43.9%) carried polymorphisms only, indicating that molecular analysis substantially reduces the number of referrals. Variants considered to be non-disease-causing include -511C>T and -457T>A, just upstream of GALC; c.-345G>A, -335G>A, c.-238G>A, c.-85C>T, c.-59T>A, and c.-59T>G in the promoter region; synonymous polymorphisms p.G9=, p.D94=, p.L117=, p.S434=, p.Q312=, p.T524=, and p.V550=; known activity-attenuating polymorphisms p.R168C, p.D232N, and p.I546T; p.V320M; a haplotype always found with p.D232N that includes c.-196T>C, c.-7G>C, and p.A5P; and c.1786+5C>G and p.A625T. In the absence of other potentially disease-causing variants, infants found to carry only variants listed above are considered screen-negative. Six infants who carried either p.D94= (three infants) or p.V320M (three infants) and no other mutations were referred before these variants were considered polymorphisms, and all were found to be not at risk. In this report, p.D94= and p.V320M are classified as polymorphisms.
Screen-positive referral population
Three-hundred forty-eight newborns screened positive for KD and were referred for confirmatory diagnostic testing. Supplementary Table S5 online shows GALC variants and associated frequencies among the referred infants. Common variants detected included the 30-kb deletion (48 infants), p.T96A (96 infants), and p.Y303C (39 infants). We also detected mutations previously reported in infantile-onset KD patients, such as p.T513M, p.Y551S, p.R380W, and p.G537R,2,4,27 and in later-onset KD patients, including p.G49=, p.G95S, p.D171V, p.G270D, p.Y303C, and p.L618S.3,6,28,29 However, many of the sequence variants detected in the referral population were novel variants of unknown significance. Several of the novel variants of unknown significance were found in more than one individual, but most were seen only once. Supplementary Table S3 online lists haplotype backgrounds for variants detected among referred infants.
As part of the diagnostic workup, blood from the 348 referred infants was sent to the Lysosomal Diseases Testing Laboratory at Sidney Kimmel Medical College at Thomas Jefferson University for confirmatory diagnostic GALC enzyme activity testing.21 From 2006 to 2011, risk was categorized based on diagnostic GALC activity analysis (high ≤0.15, moderate = 0.16–0.29, low = 0.3–0.5, no risk >0.5). The definition of risk categories was determined from results of >40 years of diagnostic Krabbe disease testing by the Lysosomal Diseases Testing Laboratory and was implemented by consensus of the Krabbe Consortium (see Supplemental Text Box S1 online).21 In 2012, the low-risk category was eliminated and infants with activity between 0.3 and 0.5 are now classified as not at risk, unless they have two potentially disease-associated mutations, in which case they are classified as being at moderate risk. From 2006 to 2011, 16 infants had activity from 0.3 to 0.5 and two potentially disease-associated mutations; since 2012, there have been five such infants (Supplementary Table S4 online). Infants at high, moderate, and low (prior to 2012) risk are reported/classified as abnormal and infants at moderate to high risk are monitored using the reported KD follow-up algorithm and schedule.21,22 Fourteen referred infants were classified as being at high risk for KD and 37 were classified as being at moderate risk. There were 92 infants at low risk, and the remaining 203 infants referred were categorized as not at risk/no risk. These 295 infants are no longer followed in the clinic. Two infants (<1%) were not tested because they were lost to follow-up, consistent with referrals lost to follow-up for other disorders in NYS. In the absence of diagnostic testing results, we could not categorize these infants based on risk; however, neither carried two disease-associated mutations.
Infants at high risk for KD
Following confirmatory testing, 14 infants with the lowest activity were categorized as being at high risk for KD ( Table 3 ; five confirmed EIKD and nine asymptomatic infants). Symptoms and clinical findings consistent with EIKD in five infants led to recommendation for hematopoietic stem cell transplantation based on previously described criteria.21 All five had at least one copy of the 30-kb deletion. Among the five, one underwent transplantation at 1 month of age and is doing well, although gross motor delays are apparent; another had delays getting to the transplant procedure due to extenuating social circumstances and underwent transplantation at 2 months of age and has severe developmental and motor delays and ongoing issues with spasticity; two died as a result of transplant-related complications; and the family of the fifth infant decided against transplantation. At several months of age, KD was clinically evident; this child has died. Detailed clinical characteristics and outcomes for these infants will be described elsewhere (unpublished data).
The remaining nine infants at high risk did not satisfy the criteria for recommending transplantation ( Table 3 ).21 All nine carried two variants/mutations. None carried two mutations unequivocally linked to infantile-onset KD and, with the exception of case 9, the genotypes detected in these asymptomatic cases have not been reported in symptomatic individuals. Follow-up of these infants continues, and none has exhibited KD symptoms, although additional neurodiagnostic studies have not been performed for most.
Genotypes and phenotypes
Many of the variants/mutations detected in our referral population were novel and individually rare and are classified as variants of unknown significance. Therefore, it is difficult to draw conclusions linking genotypes and phenotypes, even after 8 years of KD NBS. Classification of pathogenic versus nonpathogenic variants will take years of follow-up and cannot be accurately distinguished at this time. Among variants with potential clinical significance, the 30-kb deletion, p.T96A, and p.Y303C were recurrent at appreciable frequencies (Supplementary Table S5 online). Overall, 13.8% of infants referred carried the 30-kb deletion (5 high, 4 moderate, and 39 no/low risk). Consistent with previous reports,10,30 p.R168C was always carried in cis, and no other variants were detected on the deleted allele.
p.T96A was carried by more than one-quarter of infants referred (96/348), representing by far the most common potentially pathogenic mutation detected among the referral population. Eight were homozygous, 21 were compound heterozygous for p.T96A and another variant, and 67 infants were heterozygous for p.T96A. On all 104 alleles, p.T96A was carried on the p.D232N haplotype background. Infants carrying p.T96A were classified diagnostically in moderate, low, and no-risk categories, but none was classified as being at high risk, and p.T96A has not been detected in confirmed cases identified via NBS ( Table 4 ). This variant has been reported previously in two patients with adult-onset KD (one with p.D171V on the other chromosome,7 and the other with p.P138R3). As part of a study to determine the frequency of GALC variants in the general population, we sequenced >240 specimens with >20% GALC activity, and p.T96A was not detected. In an expanded institutional review board–approved screen (NYS Department of Health), we genotyped p.T96A in initial specimens from approximately 3,400 infants accessioned by the program over a 4-day period without regard to GALC activity and with a similar race/ethnic distribution as the general population. We found the carrier frequency of p.T96A was 0.62% (no homozygotes detected; minor allele frequency = 0.31%). p.T96A may be another pseudodeficiency allele; however, it has been reported in two KD patients with adult-onset disease, so we cannot rule it out as a late-onset mutation.
Overall, 39 individuals carried at least one copy of p.Y303C, which was also always found on the p.D232N haplotype. p.Y303C carriers were found in all risk categories ( Table 4 ). This variant has been reported in several individuals compound heterozygous for p.Y303C and the 30-kb deletion, with onset ranging from 5 to 23 years,3 and in another individual compound heterozygous for p.Y303C and a splice site mutation, with onset at 29 months.5
Each infant in the high-risk group was either homozygous or compound heterozygous for two potentially pathogenic variants. In the moderate-risk group, 78.4% (29/37) carried variants/mutations on both alleles. Excluding nine infants with ≤8% activity who carried only polymorphisms, in the no-risk/low-risk groups (see Supplementary Methods online) 7.3% (21/286) carried a potentially pathogenic mutation on both alleles and 92.7% were heterozygous for a single variant/mutation (265/286).
Other than those homozygous for the 30-kb deletion (three infants), p.T96A (eight infants), or p.Y303C (eight infants; Table 4 ), seven infants homozygous for potentially pathogenic mutations were identified, including one p.V320M homozygote (diagnostic activity = 1.90; not at risk), one p.M101V homozygote (diagnostic activity = 0.18; moderate risk), and five who carried two copies of p.L618S. Seventeen p.M101V carriers were found in all risk groups in our population. p.M101V and p.M101L have been reported in later-onset patients.6,31 Each of the four infants who carried at least one copy of p.V320M were found to be not at risk. This variant is no longer considered to be pathogenic and probably represents a neutral variant or an activity-modifying polymorphism or is only associated with reduced activity because it is allelic with p.I546T (three infants) or p.I546T+p.R380W (one infant). Functional studies would be necessary to determine whether p.V320M and others (such as p.D94=) are indeed pseudodeficiency alleles. p.L618S was found in cis with several other variants in our population (Supplementary Table S3 online), and those who carried two copies of p.L618S were categorized as being at high (1; case 9, Table 3 ), moderate (3; cases 18, 21, 38, Supplementary Table S4 online), and low risk (1; case 52, Supplementary Table S4 online). p.L618S has been previously reported in the compound heterozygous state or homozygous state in at least three symptomatic patients, with onset ranging from 8 months to adulthood.3,28,29,32
Nearly all of the 348 infants referred carried at least one of the three common activity-attenuating polymorphisms (p.R168C, p.D232N, p.I546T). The frequency of each was elevated in the referral population compared with published data.2 For example, the minor allele frequency of the p.R168C+p.I546T haplotype among referrals was 28.6%, but it is reported to represent 2% of alleles in the unaffected general population.2 Slight differences in enzyme activity among the race/ethnic groups were probably due, at least in part, to different frequencies of activity-modifying polymorphisms and other variants in the race/ethnic groups.
Discussion
NYS was the first, and is currently one of only two states, to have active KD NBS.21,25,26 Although the Secretary’s Advisory Committee on Heritable Disorders in Newborns and Children advised against NBS for KD, citing insufficient knowledge about the accuracy of screening, diagnostic strategy, benefits and harms of treatment, and long-term prognosis,33 screening for KD was mandated in NYS in 2006. Screening for KD is on our panel by NYS regulation, and the program has identified infantile Krabbe disease cases. We are unaware of any infantile cases of Krabbe disease that have occurred and were not detected by screening. After 8 years, nearly 2 million infants have been screened, with a screen-positive rate of 0.018%. Fifty-one infants considered to be at moderate to high risk of disease have been identified; among them, five cases of EIKD have been confirmed. All infants classified as high-risk either have been diagnosed with EIKD or have passed the cutoff age (onset <1 year) for early/late infantile KD diagnosis. The positive predictive value of screening is 1.4% for confirmed infantile KD, although this does not take into account potential later-onset forms.
The estimated infantile KD incidence in NYS, at 1 in 394,000, is lower than previous estimates of 1 in 100,000. It was also expected that the infantile form would represent 90% of all KD.1,2 Some proportion of the asymptomatic but at-risk infants identified may, at some point, manifest disease; these infants are now “patients-in-waiting,” which can have a substantial psychosocial impact on families. In addition, the assessment of risk is under evaluation and the changes are under consideration, such as use of psychosine testing and repeating enzyme testing; this should reduce the number of at-risk infants. If all infants determined to be at high risk for disease were included, then the calculated NYS KD incidence would be 1 in 140,000; including all 51 infants considered to be at moderate to high risk, the incidence would be 1 in 39,000. Including all infants in the moderate- to high-risk groups who have two potentially pathogenic mutations in GALC, representing a more conventional recessive disease definition in which both alleles have potentially disease-causing mutations and there is an abnormal (in this case, biochemical) phenotype, the incidence would be 1 in 46,000. The residual enzyme activity makes it unlikely that infants classified as being at low risk or not at risk will develop disease, although 21 infants in these two groups were homozygous or compound heterozygous for two variants/mutations, suggesting that at least some of these variants are nonpathogenic.
The paucity of early-onset (within 6 months of age) and late-onset (6–12 months) infantile cases and overabundance of potential later-onset cases are not explained. The low incidence of EIKD may be due, in part, to the relatively high frequency of births among non-Europeans in NYS. However, none of the confirmed cases identified via NBS is of European decent, highlighting demographic differences between NYS and the general US population. Population-based screening for low GALC activity could have revealed a broader spectrum of penetrance or expressivity for later-onset KD that has not been previously recognized. Fabry disease incidence in cases identified by NBS has also been reported to be elevated compared with diagnosis in symptomatic individuals.34 The incidence of later-onset cases may be higher than reported, as suggested by the number of two-variant/mutation infants identified by NBS, and may be higher than reported in at least one other population.35
After 8 years of NBS in a state with an annual birth rate of approximately 250,000 per year, an average of 44 infants per year screened positive for KD and were referred for follow-up, corresponding to 18 per 100,000 births. This includes one asymptomatic case classified as being high-risk and possibly one confirmed infantile-onset case each year. From the laboratory perspective, including both initial and repeat heel sticks, NYS conducts ~5,100 assays, 25 retests (in duplicate), and ~1.5 reflexes to molecular analysis per week. From the follow-up perspective, one infant is referred per week (18 per 100,000 births). Other programs using comparable algorithms and methodology could expect screen-positives proportional to the size and characteristics of the population screened. Programs testing for multiple LSDs will be able to assess sample quality issues affecting enzyme activity, which could result in fewer specimens requiring molecular analysis or referral.
The NBS GALC activity assay is sensitive. However, the assay measures activity on synthetic, rather than natural, substrate. We have found variation in activity due to biological and seasonal factors lowering activity, resulting in false positives. Based on seasonal fluctuation of enzyme activity,25 we speculate that temperature plays a role, so cutoffs based on a DMA rather than using a fixed cutoff were implemented. The threshold (≤12% activity) was conservatively set to minimize false negatives. To the best of our knowledge, no KD case has been missed by screening. The MS/MS assay also identifies pseudodeficient infants. To minimize the false-positive rate, molecular GALC analysis was included as a second-tier test. We have found that molecular analysis reduces the referral rate by ~43.8% by identifying pseudodeficient infants with nonpathological low activity, sparing families distress and anxiety associated with referral for a devastating disease with limited treatment options.36
It is advantageous for NBS laboratories to have the required expertise, infrastructure, and funding to conduct timely molecular analyses. Samples cannot be batched and must be analyzed immediately on detection of low GALC activity for the infant to be seen by a specialist and to undergo clinical and neurological testing if warranted and still provide families with time to make a decision regarding treatment. The benefit of having this information substantially outweighs the cost to the families and health-care system because 43.8% of families are never brought to medical attention. If NBS programs do not provide it, then many physicians would order genetic testing following a KD referral, increasing the time for this information to be considered along with the clinical assessment. The referral rate could be reduced further if only infants with two mutations were referred; however, the risk of missing an infant with a second undetected mutation remains a concern because we do not know the frequency of other deletions not detectable by sequencing.37 Furthermore, families with a child with one mutation can be appropriately counseled with respect to carrier testing and risk for future pregnancies.
Several additional factors are associated with either large (age at sampling) or subtle (race/ethnicity and gender) variation in the NBS assay ( Table 2 ). For example, the average activity in DOB specimens is more than twice that found in specimens sampled after the first 24 hours of life. NYS considers DOB specimens unsuitable for most NBS tests, and this elevation in activity could mask true KD cases, resulting in false negatives. Thus far, all six infants with DOB specimens subsequently referred based on results from repeat specimens were classified after confirmatory testing as being at low risk/not at risk; we do not know the impact of DOB sampling on true positives. Furthermore, although inclusion of DOB specimens in the DMA calculation has minor effects on the NYS average, the mean could be inflated in laboratories that test a small number of samples or receive a large number of unsuitable DOB specimens.
The NBS MS/MS assay detects infants with low GALC activity but, as previously noted,25 there is little correlation between activity measured by the NBS assay and the diagnostic laboratory. Normalization to white blood cell protein, use of β-galactosidase enzyme activity for specimen quality monitoring, and variability in inhibitors probably contribute to differences. When white cell lysates isolated from diagnostic samples are split and tested using both assays, results are more comparable (data not shown). The combination of activity with molecular analysis more closely aligns with diagnostic activity, because the majority of newborns at high (100%) and moderate (78.4%) risk carried two GALC mutations/variants, whereas most of the infants at low risk/not at risk were heterozygous for one mutation/variant or carried polymorphisms only (92.9%).
Prior to the onset of screening, little had been published on the KD mutation spectrum and prevalence, especially in North America. Contrary to our expectations, several mutations expected to be prevalent either have not been detected in the first 8 years of screening (c.1424delA) or were detected in only one or a few infants (e.g., p.G270D, p.R380W). Other potentially pathogenic mutations such as p.T96A and p.Y303C are relatively common in the NYS referral population. Except for variants that clearly abolish protein function, such as the 30-kb deletion and other truncation, frameshift, nonsense, and splicing mutations, it is difficult to infer genotype–phenotype correlations because most variants are novel and/or rare. It is fairly well accepted that an individual homozygous for the 30-kb deletion will develop EIKD, and an individual carrying p.G270D in conjunction with any other mutation will develop later-onset KD.2 Others have found little association between GALC activity and disease onset or severity,1 and there is still limited knowledge of the effects of specific mutations on GALC activity. It is unclear whether novel variants detected in our population represent pathogenic mutations or are rare variants that slightly modify GALC activity in a manner similar to other more common polymorphisms such as p.I546T and p.R168C but are not sufficient to cause disease. Variants such as p.T96A and p.Y303C have been reported or observed in a few patients,2,3,5,7 but none of the 21 infants homozygous or compound heterozygous for p.T96A and/or p.Y303C or any of the infants compound heterozygous for p.T96A or p.Y303C and another known disease-associated mutation is known to have been diagnosed with KD. These two variants are probably associated with later-onset disease or nonpathological low GALC activity, or may cause disease only when coexistent with specific variants/mutations. Functional data from each of the mutant alleles will be an important step in establishing pathogenicity (functional studies are being completed as a joint collaboration in the NBS (M.C.) and diagnostic testing (D.A.W.) laboratories), which is complicated by variability in genetic background. We expect that as KD NBS continues and expands to other states, screening and referral algorithms may be refined and tailored, especially if phenotypes associated with specific mutations are identified.
Three key modifications to the original testing and referral algorithm have been made. First, although the NBS assay was rigorously tested using known positive, carrier, and negative controls before screening was initiated,25 the low activity cutoff was increased from 10 to 12% when an infant screened positive at 9.9% and was confirmed to have EIKD. This point emphasizes the importance of using age-matched controls in establishing cutoff values. Second, infants with ≤8% activity are no longer immediately referred. We had initially taken a cautious approach by referring infants with very low activity as quickly as possible, but the delay due to second-tier molecular analysis has been minimized. Furthermore, five infants with ≤8% activity carried only activity-modifying polymorphisms and were determined to be at low risk (one infant) or not at risk (four infants) for KD after the diagnostic testing laboratory measured GALC activities that were >0.35 nmol/hour/mg. Third, we eliminated the low-risk category, so any newborn with activity that is ≥0.3 nmol/hour/mg is placed in the no-risk/not-at-risk category, with the exception of infants carrying two variants, who are considered to be at moderate risk. Screening, referral, and follow-up algorithms continue to be reviewed by the NBS program and Specialty Care Centers.
KD NBS in other states is imminent.16 Although hematopoietic stem cell transplantation performed during the first month of life may provide benefit to some infants with infantile-onset KD,23,24,38 it is challenging for Specialty Care Centers to diagnose, evaluate, and refer infants to qualified transplant centers within the first few weeks of life. Even when transplantation is performed early, delays in development of gross motor skills have developed in most patients, including our two surviving patients.22 Later-onset forms detected by screening can also benefit from transplantation, even after mild symptoms develop.25 Still, transplantation may be most beneficial when performed before irreversible neurological damage occurs, so its true success rate may depend on identification and transplantation of patients before clinical presentation. Counseling families in the absence of apparent disease is challenging because, in the absence of two pathogenic mutations that clearly abolish protein function, neither enzyme activity nor molecular analysis predicts which high-risk or moderate-risk infants will develop symptoms during their lifetime. Identification and use of other biochemical markers of disease such as psychosine may eventually aid in establishing a diagnosis and/or risk assessment.39,40 Health-care professionals must be highly skilled in communicating information about KD prognosis and benefits, risks, and long-term outcomes associated with transplantation so that families are empowered to make the difficult decision of whether their child should undergo transplantation.
Disclosure
R.W.E. received research support from Synageva BioPharma, Corp. M.L.E. is a consultant for Shire Human Genetics, Biomarin, Synageva, Medicinova, Alexion, Abeona Therapeutics, Stem Cells, and principal investigator of a clinical trial for Shire and Synageva. J.E.P. receives research grant money from Genzyme and Shire. The other authors declare no conflict of interest.
References
Duffner PK, Jalal K, Carter RL. The Hunter’s Hope Krabbe family database. Pediatr Neurol 2009;40:13–18.
Wenger DA, Suzuki K, Suzuki Y. Galactosylceramide lipidosis: globoid cell leukodystrophy (Krabbe disease). In: Valle D, Beaudet AL, Vogelstein B, et al. (eds). The Online Metabolic and Molecular Bases of Inherited Disease. McGraw-Hill: New York, 2013.
Debs R, Froissart R, Aubourg P, et al. Krabbe disease in adults: phenotypic and genotypic update from a series of 11 cases and a review. J Inherit Metab Dis 2013;36:859–868.
Duffner PK, Barczykowski A, Jalal K, Yan L, Kay DM, Carter RL. Early infantile Krabbe disease: results of the World-Wide Krabbe Registry. Pediatr Neurol 2011;45:141–148.
Duffner PK, Barczykowski A, Kay DM, et al. Later onset phenotypes of Krabbe disease: results of the world-wide registry. Pediatr Neurol 2012;46:298–306.
De Gasperi R, Gama Sosa MA, Sartorato EL, et al. Molecular heterogeneity of late-onset forms of globoid-cell leukodystrophy. Am J Hum Genet 1996;59:1233–1242.
Luzi P, Rafi MA, Wenger DA. Multiple mutations in the GALC gene in a patient with adult-onset Krabbe disease. Ann Neurol 1996;40:116–119.
Köhler W. Leukodystrophies with late disease onset: an update. Curr Opin Neurol 2010;23:234–241.
Lyon G, Hagberg B, Evrard P, Allaire C, Pavone L, Vanier M. Symptomatology of late onset Krabbe’s leukodystrophy: the European experience. Dev Neurosci 1991;13:240–244.
Rafi MA, Luzi P, Chen YQ, Wenger DA. A large deletion together with a point mutation in the GALC gene is a common mutant allele in patients with infantile Krabbe disease. Hum Mol Genet 1995;4:1285–1289.
Kleijer WJ, Keulemans JL, van der Kraan M, et al. Prevalent mutations in the GALC gene of patients with Krabbe disease of Dutch and other European origin. J Inherit Metab Dis 1997;20:587–594.
Duffey TA, Bellamy G, Elliott S, et al. A tandem mass spectrometry triplex assay for the detection of Fabry, Pompe, and mucopolysaccharidosis-I (Hurler). Clin Chem 2010;56:1854–1861.
Li Y, Scott CR, Chamoles NA, et al. Direct multiplex assay of lysosomal enzymes in dried blood spots for newborn screening. Clin Chem 2004;50:1785–1796.
Adam BW, Orsini JJ Jr, Martin M, et al. The preparation and storage of dried-blood spot quality control materials for lysosomal storage disease screening tests. Clin Biochem 2011;44:704–710.
Metz TF, Mechtler TP, Orsini JJ, et al. Simplified newborn screening protocol for lysosomal storage disorders. Clin Chem 2011;57:1286–1294.
Hunter’s Hope. 2015. http://www.huntershope.org/site/PageServer?pagename=unbs_state. Accessed 26 October 2015.
Matern D, Gavrilov D, Oglesbee D, Raymond K, Rinaldo P, Tortorelli S. Newborn screening for lysosomal storage disorders. Semin Perinatol 2015;39:206–216.
Zhang XK, Elbin CS, Chuang WL, et al. Multiplex enzyme assay screening of dried blood spots for lysosomal storage disorders by using tandem mass spectrometry. Clin Chem 2008;54:1725–1728.
la Marca G, Casetta B, Malvagia S, Guerrini R, Zammarchi E. New strategy for the screening of lysosomal storage disorders: the use of the online trapping-and-cleanup liquid chromatography/mass spectrometry. Anal Chem 2009;81:6113–6121.
De Jesus VR, Zhang XK, Keutzer J, et al. Development and evaluation of quality control dried blood spot materials in newborn screening for lysosomal storage disorders. Clin Chem 2009;55:158–164.
Duffner PK, Caggana M, Orsini JJ, et al. Newborn screening for Krabbe disease: the New York State model. Pediatr Neurol 2009;40:245–52; discussion 253.
Duffner PK, Caviness VS Jr, Erbe RW, et al. The long-term outcomes of presymptomatic infants transplanted for Krabbe disease: report of the workshop held on July 11 and 12, 2008, Holiday Valley, New York. Genet Med 2009;11:450–454.
Escolar ML, Poe MD, Martin HR, Kurtzberg J. A staging system for infantile Krabbe disease to predict outcome after unrelated umbilical cord blood transplantation. Pediatrics 2006;118:e879–e889.
Escolar ML, Poe MD, Provenzale JM, et al. Transplantation of umbilical-cord blood in babies with infantile Krabbe’s disease. N Engl J Med 2005;352:2069–2081.
Orsini JJ, Morrissey MA, Slavin LN, et al. Implementation of newborn screening for Krabbe disease: population study and cutoff determination. Clin Biochem 2009;42:877–884.
Hopkins PV, Campbell C, Klug T, Rogers S, Raburn-Miller J, Kiesling J. Lysosomal storage disorder screening implementation: findings from the first six months of full population pilot testing in Missouri. J Pediatr 2015;166:172–177.
Tappino B, Biancheri R, Mort M, et al. Identification and characterization of 15 novel GALC gene mutations causing Krabbe disease. Hum Mutat 2010;31:E1894–E1914.
Xu C, Sakai N, Taniike M, Inui K, Ozono K. Six novel mutations detected in the GALC gene in 17 Japanese patients with Krabbe disease, and new genotype-phenotype correlation. J Hum Genet 2006;51:548–554.
Furuya H, Kukita Y, Nagano S, et al. Adult onset globoid cell leukodystrophy (Krabbe disease): analysis of galactosylceramidase cDNA from four Japanese patients. Hum Genet 1997;100:450–456.
Luzi P, Rafi MA, Wenger DA. Characterization of the large deletion in the GALC gene found in patients with Krabbe disease. Hum Mol Genet 1995;4:2335–2338.
Sharp ME, Laule C, Nantel S, et al. Stem cell transplantation for adult-onset krabbe disease: report of a case. JIMD Rep 2013;10:57–59.
Satoh JI, Tokumoto H, Kurohara K, et al. Adult-onset Krabbe disease with homozygous T1853C mutation in the galactocerebrosidase gene. Unusual MRI findings of corticospinal tract demyelination. Neurology 1997;49:1392–1399.
Kemper AR, Knapp AA, Green NS, Comeau AM, Metterville DR, Perrin JM. Weighing the evidence for newborn screening for early-infantile Krabbe disease. Genet Med 2010;12:539–543.
Spada M, Pagliardini S, Yasuda M, et al. High incidence of later-onset fabry disease revealed by newborn screening. Am J Hum Genet 2006;79:31–40.
Fiumara A, Barone R, Arena A, et al. Krabbe leukodystrophy in a selected population with high rate of late onset forms: longer survival linked to c.121G>A (p.Gly41Ser) mutation. Clin Genet 2011;80:452–458.
Hewlett J, Waisbren SE. A review of the psychosocial effects of false-positive results on parents and current communication practices in newborn screening. J Inherit Metab Dis 2006;29:677–682.
Tanner AK, Chin EL, Duffner PK, Hegde M. Array CGH improves detection of mutations in the GALC gene associated with Krabbe disease. Orphanet J Rare Dis 2012;7:38.
Escolar ML, Yelin K, Poe MD. Neurodevelopmental outcome of children with infantile Krabbe disease treated with umbilical cord blood transplantation: 10 years of follow-up. Curr Med Lit Lysosomal Storage Dis 2006;6:71–9.
Turgeon CT, Orsini JJ, Sanders KA, et al. Measurement of psychosine in dried blood spots–a possible improvement to newborn screening programs for Krabbe disease. J Inherit Metab Dis 2015;38:923–929.
Chuang WL, Pacheco J, Zhang XK, et al. Determination of psychosine concentration in dried blood spots from newborns that were identified via newborn screening to be at risk for Krabbe disease. Clin Chim Acta 2013;419:73–76.
Richards S, Aziz N, Bale S, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 2015;17:405–424.
Acknowledgements
This work was supported in part by the New York State Department of Health and a grant from the Legacy of Angels Foundation (to D.A.W.). Other members of the New York State Krabbe Disease Consortium include Mary R. Andriola, Carl J. Crosley, Aleksandra Djukic, Thomas J. Langan, Thomas Naidich, James M. Provenzale, Stanley Rothman, Lissette Estrella, Laura Jacobsen, Elaine Philipson, Ryan Miller, and Roberta Salveson. We thank Susan Hsu of the Red Cross in Philadelphia, PA; Colleen Stevens of the Wadsworth Center for performing HLA analyses; the Centers for Disease Control and Prevention for providing substrate, internal standard, and quality control materials; and Dietrich Matern and Silvia Tortorelli for generously sharing a protocol that multiplexed Krabbe disease with adrenoleukodystrophy. We also thank laboratory and follow-up staff; staff at Specialty Care Centers for the care of screen-positive newborns; and the New York State Krabbe Consortium for helpful discussions.
Author information
Authors and Affiliations
Consortia
Corresponding author
Supplementary information
Supplementary Methods
(DOC 74 kb)
Supplementary Tables
(DOC 519 kb)
Supplemental Text Box S1
(DOC 30 kb)
Rights and permissions
About this article
Cite this article
Orsini, J., Kay, D., Saavedra-Matiz, C. et al. Newborn screening for Krabbe disease in New York State: the first eight years’ experience. Genet Med 18, 239–248 (2016). https://doi.org/10.1038/gim.2015.211
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2015.211
Keywords
This article is cited by
-
Identifying altered developmental pathways in human globoid cell leukodystrophy iPSCs-derived NSCs using transcriptome profiling
BMC Genomics (2023)
-
Probability of high-risk genetic matching with oocyte and semen donors: complete gene analysis or genotyping test?
Journal of Assisted Reproduction and Genetics (2022)
-
Consensus guidelines for newborn screening, diagnosis and treatment of infantile Krabbe disease
Orphanet Journal of Rare Diseases (2018)
-
Emptying the stores: lysosomal diseases and therapeutic strategies
Nature Reviews Drug Discovery (2018)
-
New in Newborn Screening
Current Genetic Medicine Reports (2017)