Abstract
Purpose:
The American College of Medical Genetics and Genomics (ACMG) recommended that clinical laboratories performing next-generation sequencing analyze and return pathogenic variants for 56 specific genes it considered medically actionable. Our objective was to evaluate the clinical and economic impact of returning these results.
Methods:
We developed a decision-analytic policy model to project the quality-adjusted life-years and lifetime costs associated with returning ACMG-recommended incidental findings in three hypothetical cohorts of 10,000 patients.
Results:
Returning incidental findings to cardiomyopathy patients, colorectal cancer patients, or healthy individuals would increase costs by $896,000, $2.9 million, and $3.9 million, respectively, and would increase quality-adjusted life-years by 20, 25.4, and 67 years, respectively, for incremental cost-effectiveness ratios of $44,800, $115,020, and $58,600, respectively. In probabilistic analyses, returning incidental findings cost less than $100,000/quality-adjusted life-year gained in 85, 28, and 91%, respectively, of simulations. Assuming next-generation sequencing costs $500, the incremental cost-effectiveness ratio for primary screening of healthy individuals was $133,400 (<$100,000/quality-adjusted life-year gained in 10% of simulations). Results were sensitive to the cohort age and assumptions about gene penetrance.
Conclusion:
Returning incidental findings is likely cost-effective for certain patient populations. Screening of generally healthy individuals is likely not cost-effective based on current data, unless next-generation sequencing costs less than $500.
Genet Med 17 7, 587–595.
Similar content being viewed by others
Introduction
The advent of next-generation sequencing (NGS) technologies is enabling the increased use of whole-exome and whole-genome sequencing in clinical practice. As the range of applications for NGS continues to expand, so too does the potential to discover incidental findings (IFs). IFs are genetic results unrelated to the indication for sequencing but that may be clinically significant.1 How to address the potential discovery and disclosure of IFs has been a topic of intense debate among researchers, clinicians, and policy-makers grappling with how to implement genome sequencing in clinical practice.2
The American College of Medical Genetics and Genomics (ACMG) recently developed a policy statement recommending that all clinical laboratories performing NGS analyze and report any IFs identified from a list of 56 genes considered medically actionable and having a high probability of causing disease.3 Many commentators responded to these recommendations by raising concerns about the clinical and ethical implications of this policy.4,5,6,7,8,9 These concerns prompted a revision to the policy allowing patients to opt out of the analysis and return of IFs.10 However, there is concern that the return of IFs has the potential to increase the downstream health-care costs associated with diagnostic workup, surveillance, and prophylactic treatments while offering little, if any, benefit to patients.5,11 Largely absent from the debate, however, is formal evaluation of the patient and economic outcomes of returning the ACMG-recommended list of IFs.
Our aim was to assess the expected clinical and economic consequences of returning ACMG-recommended IFs to individuals receiving genomic sequencing. Although decision models are commonly used to evaluate the clinical and economic impact of returning a single genetic test result,12,13,14,15 the myriad results potentially generated from NGS required a more complex model structure. To that end, we developed a novel quantitative framework to examine the impact of returning ACMG-recommended IFs to patients. This framework uses decision-analytic techniques and synthesizes a complex array of data from the genomic, clinical, epidemiological, comparative-effectiveness, and patient outcomes literature. The findings from our study can inform the ongoing policy debate regarding the return of the ACMG-recommended list of IFs, as well as provide a framework for formally evaluating other policies regarding the return of genomic IFs.
Materials and Methods
Overall approach
Our goal was to develop an explicit, quantitative model assessing clinical and economic outcomes to inform policies regarding the return of genomic Ifs: the Genomic IFs Policy Model. We created a decision-analytic model to project the lifetime health outcomes and costs associated with the return of the ACMG-recommended list of genes from genomic sequencing. Because the clinical settings in which NGS will be used are rapidly evolving, we could not evaluate a representative group of patients who will receive NGS as part of clinical care. Instead, we evaluated three archetypal settings in which opportunistic screening with NGS may become part of routine clinical or preventative care: patients with hypertrophic or dilated cardiomyopathy; patients with colorectal cancer or polyposis; and generally healthy individuals receiving testing because family members have been identified with genomic risk factors or family history indicates risk of a specific disease. In these three cases, screening for IFs is considered opportunistic because patients were undergoing NGS for indications unrelated to the IFs. Importantly, we were not evaluating the cost-effectiveness of NGS in these settings but rather the policy of returning IFs in several hypothetical clinical scenarios.
In our base case, individuals entered our decision model at age 45 years (cardiomyopathy patients and healthy individuals) or age 50 years (colorectal cancer patients); however, we also explored different ages in sensitivity analyses. Details of how we modeled outcomes for each patient group are included in the Supplementary Materials and Methods online and outlined below. In brief, we recreated (cardiomyopathy) or obtained original (colorectal cancer) disease simulation models from previously published cost-effectiveness analyses (CEAs) of genetic testing in these conditions.16,17 The life expectancy of generally healthy individuals was estimated from life tables.
Genomic parameters
The ACMG identified 56 genes associated with 24 inherited conditions that they recommended be analyzed and returned to individuals receiving NGS for any clinical purpose. The conditions were selected because of their high penetrance and clinical actionability, and the variants were limited to those known or expected to be pathogenic. The probabilities of receiving specific IFs after genome sequencing were based on population-level estimates of prevalence from GeneReviews, a collection of expert-authored, peer-reviewed articles on the diagnosis and management of specific genetic diseases ( Table 1 ).18
Policy model structure
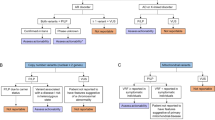
In the decision model, individuals undergoing genomic sequencing as part of clinical care can either receive or not receive the ACMG-recommended IFs ( Figure 1 ). If a patient has a pathogenic variant in one of the dominant genes, or in two for a recessive disorder, then disclosure could lead to additional diagnostic tests, increased surveillance, and/or prophylactic treatment. These early interventions generally reduce the increased risk of developing adverse outcomes such as cancer or sudden cardiac death. If patients have a pathogenic variant but do not receive IFs as part of genomic sequencing, then they are assumed to remain at increased risk for development of the condition for their lifetime.
Given the complexity of independently modeling 56 IFs, we devised an approach to capture the major impacts of returning these findings while providing a tractable and useful policy model. First, 7 of the 24 conditions represented by the ACMG list would encompass ~95% of the expected IFs. We therefore focused our analysis on these seven conditions and evaluated the incremental costs and quality-adjusted life years (QALYs) saved by the return of results for each individually. For five conditions, we derived estimates of costs and effectiveness primarily from previously published CEAs; we created novel decision models for the remaining two conditions. Second, given the very small population-level impact of the rare genetic conditions, and given the paucity of data regarding natural history progression and treatment effectiveness, we modeled these conditions collectively. Complete details regarding the modeling of each condition are described in the Supplementary Materials and Methods online. Third, we multiplied the individual-level estimates for costs and QALYs associated with returning an IF by the expected prevalence of identifying and returning those results to estimate the implications of returning IFs at the population level. We assumed that cardiac-related variants from the ACMG list would not be considered incidental for cardiomyopathy patients and that Lynch syndrome and BRCA variants would not be incidental for patients with colorectal cancer. Finally, we conducted sensitivity analyses in which we explored the impact of a wide range of estimates and assumptions for the remaining 17 rare conditions.
Overview of IF-specific modeling
We searched PubMed for previously published CEAs of genetic testing, particularly cascade or population-based testing strategies, for each of the seven most prevalent conditions using as search terms both the phenotype and pathogenic variants. We identified two CEAs for familial hypercholesterolemia19,20,21; however, given the high rate of lipid screening in the United States,22 we assumed in our base case that all individuals receiving an FH IF would already be identified from lipid screening. We found one CEA of cascade genetic testing for hypertrophic or dilated cardiomyopathy16 and long QT syndrome23; however, these CEAs did not publish their results in sufficient detail to be included in our model directly; therefore, we reconstructed these disease-specific decision models using Markov modeling techniques and, in so doing, confirmed the key model inputs that were used in the original publications with several clinical genetics experts (G.P.J., C.J.G., and W.B.). We identified two CEAs for hereditary breast and ovarian cancer24,25 that reported sufficient data regarding incremental costs and health effects to be included in our model. Finally, we identified four CEAs for Lynch syndrome testing.17,26,27,28 Upon request, the authors of one CEA provided us with their original model of genetic testing for Lynch syndrome17; the rationale and implications of choosing this CEA are discussed in the Supplementary Materials and Methods online. See Table 1 , Supplementary Table S1 online, and the Supplementary Materials and Methods online for complete details about how we modeled each IF.
Cost parameters
In our base case analysis, we did not include the cost of sequencing because we assumed it was ordered as part of the diagnostic workup; however, we included the cost of sequencing in a scenario analysis for generally healthy individuals as a preliminary exploration of primary screening with NGS. For all analyses, we assumed that each individual who received IFs would require additional pre-test genetic counseling to discuss the potential of returning IFs (15 minutes) and each IF would require review to determine pathogenicity (23 minutes).29 Finally, we assumed that each individual with a positive IF would receive an additional hour of posttest counseling in addition to condition-specific diagnostic testing, surveillance, and/or treatment.
Analyses
We calculated QALYs, lifetime health-care costs, and the incremental differences between strategies of disclosing or not disclosing the ACMG-recommended list of IFs. We calculated the incremental cost-effectiveness ratio (ICER) as the difference in costs divided by the difference in QALYs. We performed one-way sensitivity analyses for individual variables over their plausible ranges and propagated uncertainty in our model inputs using probabilistic sensitivity analysis with 10,000 Monte Carlo simulations. We used a lifetime horizon and a health-care payer perspective in 2013 US dollars; both costs and outcomes were discounted at 3% annually.30 Additional details regarding these analyses are included in the Supplementary Materials and Methods online.
Scenario analyses and assessment of assumptions
We conducted several scenario analyses to explore the assumptions in our model. First, because true prevalence and penetrance in unselected individuals are uncertain, we conducted sensitivity analyses to explore the impact that different estimates would have on our results. Second, we varied the age of the population in our model to provide a reasonable first approximation of the incremental costs and health outcomes associated with disclosing IFs across different patient populations with varying life expectancies. Third, we explored the clinical and economic implications of including family members who might be tested after an IF. Fourth, we explored how including FH results as potential IFs would impact our results across a range of assumptions regarding adherence. Finally, we varied the cost of sequencing in our scenario analysis evaluating primary screening of generally healthy individuals with NGS. See Supplementary Materials and Methods online for additional details.
Results
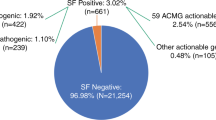
The expected probability of returning any pathogenic IF from the list of ACMG genes was 0.9% (1.2% including FH) for generally healthy individuals in our base case. Among cardiomyopathy and colorectal cancer patients, the expected probability of an IF was lower (0.6 and 0.4%, respectively) because certain IFs from the ACMG list would no longer be considered incidental in these populations. Table 2 shows the wide variation in the expected distribution of IFs in hypothetical cohorts of 10,000 individuals who received genome sequencing related to cardiomyopathy or colorectal cancer or as part of routine preventative medicine.
For an asymptomatic individual with a pathogenic variant, the expected individual-level benefit of returning condition-specific IFs ranged from 1.8 QALYs (hypertrophic or dilated cardiomyopathy) to 0.09 QALY (malignant hyperthermia susceptibility). For cardiomyopathy and colorectal cancer patients, the average expected individual-level benefit of returning each IF was somewhat lower than among generally healthy individuals because of the lower overall life expectancy given their condition ( Table 2 ). Among generally healthy individuals, the expected incremental costs of returning an IF ranged from being cost-saving for those with BRCA1 or BRCA2 pathogenic variants to ~$170,000 for those with an arrhythmogenic right ventricular dysplasia (ARVD) pathogenic variant; overall costs were somewhat attenuated in cardiomyopathy or colorectal cancer patients ( Table 2 ). Similarly, the ICERs of returning a specific IF ranged from being cost-saving (BRCA1/2) to more than $300,000 per QALY gained (ARVD).
In total, the incremental health benefits of returning the ACMG list of IFs to patients receiving NGS varied by the underlying clinical condition of the cohort ( Table 3 ). Generally healthy individuals who received NGS as part of a diagnostic workup would gain the most benefit from the return of IFs, ~67 QALYs (95% credible range (CR): 32−115) per 10,000 individuals tested at a cost of ~$3.9 million (95% CR: $2.0 million− $7.2 million), for an ICER of $58,600 (95% CR: $31,800–$129,100) per QALY gained. Cardiomyopathy patients were expected to gain substantially less benefit (20 QALYs per 10,000 patients tested; CR: 4.5−40) but also to be associated with lower downstream costs ($896,000, CR: $490,000–$1.9 million), for an ICER of $44,800 (95% CR: $18,300–$217,800). Finally, a cohort of 10,000 colorectal cancer patients was expected to gain 25.4 QALYs (95% CR: 12–66) at an incremental cost of $2.9 million (95% CR: $1.6 million−$6.3 million), for an ICER of $115,000 (95% CR: $49,700–$251,200) per QALY gained.
Scenario analyses
When we assumed the penetrance of IFs was reduced, the expected incremental benefits associated with disclosure also decreased (Supplementary Table S2 online). For example, when penetrance was reduced by 30%, the QALYs gained per 10,000 individuals tested were reduced to 15, 20, and 51 for cardiomyopathy, colorectal cancer, and otherwise healthy individuals, respectively, whereas costs remained relatively constant. The total costs and QALYs gained from returning IFs increased approximately proportionally with increasing prevalence because the incremental costs and benefits associated with returning IFs were primarily attributable to surveillance and prophylactic treatment, which increased proportionally as prevalence increased, rather than additional counseling or review of variants for pathogenicity.
Our results were particularly sensitive to the age of the cohort (Supplementary Table S2 and Figure S1 online). For example, a generally healthy individual who is 65 years old at the time of sequencing is expected to gain only one-sixth as many QALYs as a 25-year-old because the treatments generally provide less absolute improvement in life expectancy for older individuals. However, older individuals still receive similar initial treatments and therefore incur a substantial fraction of the total costs: a 65-year-old incurs more than half of the total lifetime costs for treatment and surveillance of a 25-year-old. Among generally healthy individuals, the ICER ranged from $45,700 to $120,500 for 25- to 65-year-olds, respectively. Similarly wide ranges were seen for individuals with cardiomyopathy ($31,100–$215,100 per QALY) or colorectal cancer ($75,600–$190,100 per QALY). The probability that returning IFs to patients would be cost-effective at various willingness-to-pay thresholds decreases substantially with increasing age in each clinical setting (Supplementary Figure S1 online).
The cost-effectiveness of returning IFs improved when we also included the clinical and economic implications for family members, particularly children. If we assumed that 55-year-olds had one 25-year-old child who was willing to undergo targeted genetic testing after an IF, the cost-effectiveness of returning IFs improved from ~$81,600 to $43,200 per QALY gained for cardiomyopathy patients and from $129,000 to $90,500 for colorectal cancer patients. With two children, the ICERs improved to $33,700 and $80,100, respectively.
The probability that sequencing and returning the ACMG list of IFs would be cost-effective varied across a range of sequencing costs and ages for primary screening (Supplementary Figure S2 online). At a cost of $500, genome sequencing and return of IFs for a 45-year-old was not cost-effective (ICER: $133,400; 95% CR: $82,600–$265,500) but approached being cost-effective for younger adults (ICER: $95,800 for 25-year-old individuals; 95% CR: $62,600–$179,900).
We also conducted a series of sensitivity analyses to explore the robustness of our results to various assumptions. First, there was considerable uncertainty regarding the treatment patterns and benefits of early treatment for ARVD; however, if all individuals with an ARVD pathogenic variant received an implantable cardioverter defibrillator (ICD) and the ICD offered no risk reduction, the overall ICER increased to $74,800 and $151,300 for generally healthy and colorectal cancer patients. Second, if we included the incremental costs and health benefits associated with returning IFs related to familial hypercholesterolemia, the ICER improved to $49,400 and $80,100 for generally healthy and colorectal cancer patients, respectively. These findings were robust to fairly conservative assumptions regarding suboptimal adherence; the ICER improved even if we assumed patients accrued the full cost of treatment but only 40 or 25% of the projected benefits from disclosure of FH findings for generally healthy individuals or patients with colorectal cancer. Finally, when we tripled the costs associated with rare conditions (to $162,000), the ICER increased to $65,800 and $69,100 per QALY for generally healthy individuals and cardiomyopathy patients.
Discussion
We developed a policy model to project the health outcomes and costs associated with returning the ACMG-recommended list of IFs. We found that carriers whose pathogenic variants were returned after NGS were expected to gain between 0.25 and 0.57 QALYs on average, depending on their underlying clinical condition, as compared with carriers whose pathogenic variants remained unknown. Returning IFs was expected to increase lifetime health-care costs by ~$90 on average for each cardiomyopathy patient, $325 for each colorectal cancer patient, and $440 for each generally healthy individual who received genomic sequencing. Therefore, our results suggest that returning the ACMG list of IFs will probably improve health outcomes across a range of clinical settings and that the cost-effectiveness of policies to return IFs will depend on the specific patient population.
Implications of our findings
The ACMG Working Group invested a considerable amount of time and effort into developing the initial list of IFs it felt should be returned to all patients receiving genomic sequencing for an underlying clinical condition. Such careful selection of specific genes and variants is a critical first step in developing a policy for the return of results. Our results indicate that by restricting the list of variants to those that are unequivocally pathogenic, lead to disease with very high probability, and for which some evidence supports the benefits of early intervention, returning results could improve health outcomes at a reasonable cost in certain patient populations. Although the return of each IF that we individually modeled provided some clinical benefit to patients, these benefits came at widely varying costs. To improve transparency and consistency, we recommend that future iterations of the IFs guidelines provide an explicit estimate of the benefits and costs for recommending disclosure of specific IFs.
Comparison with other studies
Two previous studies have evaluated the expected prevalence of IFs at the population level.29,31 Ding et al.31 used an approach similar to ours to estimate the rate of IFs and reported results that were, in general, consistent with our projections; however, our estimated rates for MUYTH-associated mutations are several orders of magnitude lower because we restricted IFs to individuals with biallelic mutations, whereas Ding et al. also included heterozygous mutations. Our decision to only include biallelic mutations follows the ACMG recommendations.3 Furthermore, including heterozygous mutations in our model would not have importantly changed our results because only individuals with biallelic mutations are recommended to receive increased surveillance.32 Dorschner et al.29 also evaluated the expected prevalence of IFs at the population level using data from 1,000 participants in the Exome Sequencing Project and found pathogenic or likely pathogenic variants from the ACMG list in 20 participants. Based on this early empirical evaluation of population-based prevalence, our projected rate of IFs may represent a lower boundary; however, a lower prevalence of pathogenic variants is conservative with respect to cost-effectiveness.
Limitations
This work has a number of limitations that warrant discussion. First, we derived estimates of incremental costs and effectiveness from previously published CEAs for several of the clinical conditions in our model and, as such, did not have as much control of or flexibility in the models, particularly in conducting sensitivity analyses. Another limitation was the lack of CEAs of genetic testing for FH that incorporate potentially suboptimal adherence to statins; however, in scenario analyses we found that even under conservative assumptions regarding treatment adherence, including FH IFs would slightly improve the overall cost-effectiveness of returning IFs. We also needed to make several assumptions regarding the costs and benefits of treatment for the rare conditions on the ACMG list given the varying quality of evidence and considerable uncertainty associated with these conditions. Our approach was pragmatic and efficient, and it was intended to provide a first approximation within a reasonable time frame to inform the ongoing debate and policy development for the return of IFs. We assessed the potential impact of uncertainty in our results by performing scenario and multivariate probabilistic sensitivity analyses, which showed fairly robust findings.
Second, the clinical conditions that could prompt exome or genome sequencing are heterogeneous and evolving. We therefore did not consider the clinical and economic implications of returning IFs to any patient who might receive NGS, but rather presented the results for three archetypal patient populations that may receive NGS and that illustrate the range of downstream clinical and economic consequences associated with returning IFs across a wide range of settings. It is also important to note that, except for the exploratory scenario analysis of primary screening with NGS, we have not assessed the cost-effectiveness of NGS, but rather that of the policy of returning IFs after NGS. The costs of adopting NGS could be much higher than those associated with returning IFs if this technology is more expensive than alternative genetic testing strategies. Whether these increased costs are warranted will depend not only on the return of IFs but also on the improved diagnosis of the primary genetic conditions, which will need to be evaluated in each clinical scenario.
Third, current estimates of prevalence and penetrance are derived primarily from selected high-risk populations and may not appropriately reflect estimates in unselected populations.32,33 Begg et al.32 found substantial between-family residual variation in the risk for development of breast cancer among those with BRCA1 or BRCA2 pathogenic variants that was attributable to unobserved risk factors. They suggested that the reduced risk seen between families of probands with contralateral versus unilateral breast cancer—reduced up to one-third by age 80 in probands with BRCA2 pathogenic variants—might mirror the reduced risk seen between carriers identified because of family history and those identified from unselected populations. We found that reducing the penetrance had a moderate impact on our results; however, the ICER remained below $80,000 per QALY gained for healthy individuals and cardiomyopathy patients across a range of plausible estimates for reduced penetrance.
Finally, it is important to note that our policy model, like the ACMG guidelines, is only a first iteration. We anticipate frequent revisions and updates as new evidence emerges or as policies are developed regarding the use of NGS in clinical care.34 In particular, only the CEA of genetic testing for BRCA1/2 explicitly incorporated the disutility of a positive result. Furthermore, patient preferences for Lynch syndrome outcomes suggest that although patients value genetic information on average, there is substantial heterogeneity in preferences.35 Thus, it will be important to incorporate additional data regarding utility for genetic information once such data are available and to explore whether and how differences in personal utility provide a rationale for more personalized return of results policies.36 The ACMG Working Group recently updated its recommendations to allow patients to opt out of the return of IFs; however, it is not clear whether such all-or-nothing disclosure policies will adequately address the heterogeneity in patient preferences for genomic information.37 Our policy model provides a timely and relevant evaluation of the long-term health and economic implications of the current ACMG recommendations and, importantly, also provides a framework for evaluating future iterations of these guidelines.
Conclusion
The release of the ACMG recommendations for the return of genomic IFs prompted considerable debate. Although many criticized the recommendations for being premature because of a lack of evidence, often policies are needed despite the absence of conclusive data to provide guidance for practicing clinicians. We developed a pragmatic and efficient policy model to project the health and economic consequences of the ACMG recommendation and evaluated the impact of key uncertainties on our results. Our findings suggest that the ACMG policy for returning results could provide substantial health benefits to certain patient populations receiving genomic sequencing in a cost-effective manner. Finally, our findings emphasize the importance of having a careful and rigorous process for guiding the return of genomic IFs to patients.
Disclosure
The authors declare no conflict of interest.
References
Wolf SM, Lawrenz FP, Nelson CA, et al. Managing incidental findings in human subjects research: analysis and recommendations. J Law Med Ethics 2008;36:219–248, 1.
Wright CF, Middleton A, Burton H, et al. Policy challenges of clinical genome sequencing. BMJ 2013;347:f6845.
Green RC, Berg JS, Grody WW, et al.; American College of Medical Genetics and Genomics. ACMG recommendations for reporting of incidental findings in clinical exome and genome sequencing. Genet Med 2013;15:565–574.
Townsend A, Adam S, Birch PH, Friedman JM . Paternalism and the ACMG recommendations on genomic incidental findings: patients seen but not heard. Genet Med 2013;15:751–752.
Vayena E, Tasioulas J . Genetic incidental findings: autonomy regained? Genet Med 2013;15:868–870.
Holtzman NA . ACMG recommendations on incidental findings are flawed scientifically and ethically. Genet Med 2013;15:750–751.
Rosenblatt DS . Who’s on first in exome and whole genome sequencing? Is it the patient or the incidental findings? Mol Genet Metab 2013;110:1–2.
Allyse M, Michie M . Not-so-incidental findings: the ACMG recommendations on the reporting of incidental findings in clinical whole genome and whole exome sequencing. Trends Biotechnol 2013;31:439–441.
Evans BJ . Minimizing liability risks under the ACMG recommendations for reporting incidental findings in clinical exome and genome sequencing. Genet Med 2013;15:915–920.
American College of Medical Genetics and Genomics. ACMG Updates Recommendation on “Opt Out” for Genome Sequencing Return of Results. https://www.acmg.net/docs/Release_ACMGUpdatesRecommendations_final.pdf. Accessed 15 April 2014.
Haverfield AD, Fuller A, Smaoui N, et al. Evaluation of ACMG Recommended Incidental Findings in Clinical Whole Exome Sequencing. Abstract presented at ASHG. http://www.ashg.org/2013meeting/abstracts/fulltext/f130122357.htm. Accessed 7 September 2014.
Ademi Z, Watts GF, Pang J, et al. Cascade screening based on genetic testing is cost-effective: evidence for the implementation of models of care for familial hypercholesterolemia. J Clin Lipidol 2014;8:390–400.
Ingles J, McGaughran J, Scuffham PA, Atherton J, Semsarian C . A cost-effectiveness model of genetic testing for the evaluation of families with hypertrophic cardiomyopathy. Heart 2012;98:625–630.
Reed SD, Dinan MA, Schulman KA, Lyman GH . Cost-effectiveness of the 21-gene recurrence score assay in the context of multifactorial decision making to guide chemotherapy for early-stage breast cancer. Genet Med 2013;15:203–211.
Elkin EB, Marshall DA, Kulin NA, et al. Economic evaluation of targeted cancer interventions: critical review and recommendations. Genet Med 2011;13:853–860.
Wordsworth S, Leal J, Blair E, et al. DNA testing for hypertrophic cardiomyopathy: a cost-effectiveness model. Eur Heart J 2010;31:926–935.
Mvundura M, Grosse SD, Hampel H, Palomaki GE . The cost-effectiveness of genetic testing strategies for Lynch syndrome among newly diagnosed patients with colorectal cancer. Genet Med 2010;12:93–104.
Pagon RA, Adam MP, Bird TD, et al. (eds). GeneReviews. University of Washington, Seattle: Seattle, WA, 1993–2014. http://www.ncbi.nlm.nih.gov/books/NBK1116/. Accessed 18 March 2014.
Marks D, Wonderling D, Thorogood M, Lambert H, Humphries SE, Neil HA . Screening for hypercholesterolaemia versus case finding for familial hypercholesterolaemia: a systematic review and cost-effectiveness analysis. Health Technol Assess 2000;4:1–123.
Marks D, Wonderling D, Thorogood M, Lambert H, Humphries SE, Neil HA . Cost effectiveness analysis of different approaches of screening for familial hypercholesterolaemia. BMJ 2002;324:1303.
Nherera L, Calvert NW, Demott K, et al. Cost-effectiveness analysis of the use of a high-intensity statin compared to a low-intensity statin in the management of patients with familial hypercholesterolaemia. Curr Med Res Opin 2010;26:529–536.
Centers for Disease Control and Prevention. Prevalence of Cholesterol Screening and High Blood Cholesterol Among Adults—United States, 2005, 2007, and 2009. Morbidity and Mortality Weekly Report (MMWR) 2012;61(35):697–702. http://www.cdc.gov/mmwr/preview/mmwrhtml/mm6135a2.htm. Accessed 7 September 2014.
Perez MV, Kumarasamy NA, Owens DK, Wang PJ, Hlatky MA . Cost-effectiveness of genetic testing in family members of patients with long-QT syndrome. Circ Cardiovasc Qual Outcomes 2011;4:76–84.
Holland ML, Huston A, Noyes K . Cost-effectiveness of testing for breast cancer susceptibility genes. Value Health 2009;12:207–216.
National Institute for Health and Clinical Excellence. Full cost effectiveness evidence review & reports, 2013. Familial breast cancer: classification and care of women at risk of familial breast cancer and management of breast cancer and related risks in people with a family history of breast cancer. Developed for NICE by the National Collaborating Centre for Cancer. http://www.nice.org.uk/guidance/cg164/resources/cg164-familial-breast-cancer-cost-effectiveness-evidence-review2. Accessed 3 May 2014.
Ladabaum U, Wang G, Terdiman J, et al. Strategies to identify the Lynch syndrome among patients with colorectal cancer: a cost-effectiveness analysis. Ann Intern Med 2011;155:69–79.
Dinh TA, Rosner BI, Atwood JC, et al. Health benefits and cost-effectiveness of primary genetic screening for Lynch syndrome in the general population. Cancer Prev Res (Phila) 2011;4:9–22.
Wang G, Kuppermann M, Kim B, Phillips KA, Ladabaum U . Influence of patient preferences on the cost-effectiveness of screening for lynch syndrome. J Oncol Pract 2012;8(suppl 3):e24s–e30s.
Dorschner MO, Amendola LM, Turner EH, et al.; National Heart, Lung, and Blood Institute Grand Opportunity Exome Sequencing Project. Actionable, pathogenic incidental findings in 1,000 participants’ exomes. Am J Hum Genet 2013;93:631–640.
Garrison LP Jr, Mansley EC, Abbott TA 3rd, Bresnahan BW, Hay JW, Smeeding J . Good research practices for measuring drug costs in cost-effectiveness analyses: a societal perspective: the ISPOR Drug Cost Task Force report–Part II. Value Health 2010;13:8–13.
Ding LE, Burnett L, Chesher D . The impact of reporting incidental findings from exome and whole-genome sequencing: predicted frequencies based on modeling. Genet Med 2014; e-pub ahead of print 31 July 2014.
Begg CB, Haile RW, Borg A, et al. Variation of breast cancer risk among BRCA1/2 carriers. JAMA 2008;299:194–201.
Burke W, Matheny Antommaria AH, Bennett R, et al. Recommendations for returning genomic incidental findings? We need to talk! Genet Med 2013;15:854–859.
Berg JS, Amendola LM, Eng C, et al.; Members of the CSER Actionability and Return of Results Working Group. Processes and preliminary outputs for identification of actionable genes as incidental findings in genomic sequence data in the Clinical Sequencing Exploratory Research Consortium. Genet Med 2013;15:860–867.
Kuppermann M, Wang G, Wong S, et al. Preferences for outcomes associated with decisions to undergo or forgo genetic testing for Lynch syndrome. Cancer 2013;119:215–225.
Foster MW, Mulvihill JJ, Sharp RR . Evaluating the utility of personal genomic information. Genet Med 2009;11:570–574.
Bennette CS, Trinidad SB, Fullerton SM, et al. Return of incidental findings in genomic medicine: measuring what patients value–development of an instrument to measure preferences for information from next-generation testing (IMPRINT). Genet Med 2013;15:873–881.
Centers for Disease Control and Prevention. FastStats, Inpatient Surgery. http://www.cdc.gov/nchs/fastats/insurg.htm. Accessed 18 March 2014.
Malignant Hyperthermia Association of the United States. FAQs: Stocking an MH Cart. http://www.mhaus.org/faqs/stocking-an-mhcart. Accessed 18 March 2014.
Dantrium IV . Direct order form from JHP Pharmaceuticals. http://www.dantrium.com/dantriumiv/dantriumiv_eform.pdf. Accessed 14 April 2014.
Malignant Hyperthermia Association of the United States. Anesthesia Workstation Preparation. http://www.mhaus.org/healthcare-professionals/mhaus-recommendations/anesthesia-workstation-preparation. Accessed 18 March 2014.
Dynasthetics Vapor-Clean. Online catalogue. http://www.dynasthetics.com/store/vapor-clean.html. Accessed 18 March 2014.
Hirshey Dirksen SJ, Larach MG, Rosenberg H, Brandom BW, Parness J, Lang RS, et al. Special article: Future directions in malignant hyperthermia research and patient care. Anesth Analg 2011;113:1108–1119.
Maron BJ, Olivotto I, Spirito P, Casey SA, Bellone P, Gohman TE, et al. Epidemiology of hypertrophic cardiomyopathy-related death: revisited in a large non-referral-based patient population. Circulation 2000;102:858–864.
Marcus GM, Glidden DV, Polonsky B, Zareba W, Smith LM, Cannom DS, et al. Efficacy of antiarrhythmic drugs in arrhythmogenic right ventricular cardiomyopathy: a report from the North American ARVC Registry. J Am Coll Cardiol 2009;54:609–615.
Groeneveld PW, Matta MA, Suh JJ, Heidenreich PA, Shea JA . Costs and quality-of-life effects of implantable cardioverter-defibrillators. Am J Cardiol 2006;98:1409–1415.
Healthcare Bluebook. CAREOperative, 2014. http://www.healthcarebluebook.com/page_Default.aspx. Accessed 18 March 2014.
Guzauskas GF, Boudreau DM, Villa KF, Levine SR, Veenstra DL . The cost-effectiveness of primary stroke centers for acute stroke care. Stroke 2012;43:1617–1623.
Unroe KT, Greiner MA, Hernandez AF, Whellan DJ, Kaul P, Schulman KA, et al. Resource use in the last 6 months of life among medicare beneficiaries with heart failure, 2000-2007. Arch Intern Med 2011;171:196–203.
Brandle M, Davidson MB, Schriger DL, Lorber B, Herman WH . Cost effectiveness of statin therapy for the primary prevention of major coronary events in individuals with type 2 diabetes. Diabetes Care 2003;26:1796–1801.
Nherera L, Calvert NW, Demott K, Humphries SE, Neil HA, Minhas R, et al. Cost-effectiveness analysis of the use of a high-intensity statin compared to a low-intensity statin in the management of patients with familial hypercholesterolaemia. Curr Med Res Opin 2010;26:529–536.
Acknowledgements
This work was supported by grant U01 HG0006507-01 from the National Human Genome Research Institute, by an R36 Award (1R36HS023340-01) from the Agency for Healthcare Research and Quality, a doctoral dissertation grant from the National Science Foundation (award 1424250), and by a grant from the University of Washington Northwest Institute of Genetic Medicine awarded by the Washington State Life Sciences Discovery Funds (grant 265508). The funding sources had no role in the design, conduct, or interpretation of the study or the preparation, review, or approval of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
(PDF 780 kb)
Rights and permissions
About this article
Cite this article
Bennette, C., Gallego, C., Burke, W. et al. The cost-effectiveness of returning incidental findings from next-generation genomic sequencing. Genet Med 17, 587–595 (2015). https://doi.org/10.1038/gim.2014.156
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2014.156
Keywords
This article is cited by
-
Breast Cancer Screening Should Embrace Precision Medicine: Evidence by Reviewing Economic Evaluations in China
Advances in Therapy (2023)
-
Early Cost Effectiveness of Whole-Genome Sequencing as a Clinical Diagnostic Test for Patients with Inoperable Stage IIIB,C/IV Non-squamous Non-small-Cell Lung Cancer
PharmacoEconomics (2021)
-
Estimating the costs of genomic sequencing in cancer control
BMC Health Services Research (2020)
-
Population genetic screening efficiently identifies carriers of autosomal dominant diseases
Nature Medicine (2020)
-
Primary care provider perspectives on using genomic sequencing in the care of healthy children
European Journal of Human Genetics (2020)