Abstract
Purpose:
Up to half of unique genetic variants in genomic evaluations of familial cancer risk will be rare variants of uncertain significance. Classification of rare variants will be an ongoing issue as genomic testing becomes more common.
Methods:
We modified standard power calculations to explore sample sizes necessary to classify and estimate relative disease risk for rare variant frequencies (0.001–0.00001) and varying relative risk (20–1.5), using population-based and family-based designs focusing on breast and colon cancer. We required 80% power and tolerated a 10% false-positive rate because variants tested will be in known genes with high pretest probability.
Results:
Using population-based strategies, hundreds to millions of cases are necessary to classify rare cancer variants. Larger samples are necessary for less frequent and less penetrant variants. Family-based strategies are robust to changes in variant frequency and require between 8 and 1,175 individuals, depending on risk.
Conclusion:
It is unlikely that most rare missense variants will be classifiable in the near future, and accurate relative risk estimates may never be available for very rare variants. This knowledge may alter strategies for communicating information about variants of uncertain significance to patients.
Genet Med 16 7, 529–534.
Similar content being viewed by others
Introduction
Up to half of unique genetic variants in evaluations of familial cancer risk are variants of uncertain significance.1,2,3 The number of rare missense variants identified increases linearly, proportionate with the length of DNA sequenced, at a rate of ˜0.008 rare variants identified per kilobase of exonic DNA sequence.4 New next-generation -sequencing–based clinical assays aimed at comprehensive evaluation of cancer risk genes are predicted to identify at least one rare missense variant in more than half of the individuals sequenced.5 These rare variants of uncertain significance can cause confusion and patient anxiety, so definitive classification of these variants is a high priority.6,7,8 Several frameworks have been proposed for classifying novel variants in known cancer genes, with ongoing debate about the level of evidence necessary to classify a novel variant in each category.9,10,11,12,13 For example, the framework outlined by Plon et al. (2008)9 suggests that a variant could be considered pathogenic if combined evidence from multiple sources indicates a 99% or greater probability that the variant causes the phenotype in question, and the Partners Laboratory for Molecular Medicine has multiple criteria for pathogenicity, including LOD score >3 (≥10 meioses).10 Similarly, a variant could be considered likely to be pathogenic if there is greater than 95% probability that the variant is pathogenic or if segregation is seen across more than three meiosis cycles along with other supporting evidence, depending on the classification framework.9,10 Several groups have detailed specific mechanisms that might be used to combine evidence from multiple sources to classify variants.14,15,16,17
Classification is important, but the information about risk of disease is what drives clinical decisions. This risk information intuitively appears implicit in classification; however, classification may or may not facilitate accurate risk prediction. For genes associated with familial cancer risk, understanding novel variants could be seen as a two-step process: (i) categorizing the variant in a broad class and (ii) estimating actual cancer risk conferred by the novel variant. This second step can be more challenging than simply classifying a variant, particularly for missense and splice-site mutations.
In practice, cancer risk is usually inferred from literature based on other variants with the same classification, and many methods for categorizing uncertain variants explicitly assume that risk for novel variants will be identical to that for previously described, highly penetrant variants.6,14 Grouping variants that clearly completely abrogate gene function, such as premature stop codons, early frameshift mutations, and large deletions, appears appropriate for many genes, particularly for highly penetrant cancer risk genes such as BRCA1 (breast cancer 1, early onset) or MLH1 (mutL homolog 1). However, all variants that alter activity, such as missense variants, leaky splice-site variants, or variants that occur near the 3′ end of a gene, may not confer similar risk.18,19,20 In this article, we will first illustrate the level of risk implied by classification groups, acknowledging that there is large uncertainty surrounding this implied risk. Then we will describe the magnitude of effort that would be necessary to better define cancer risk estimates for novel, rare variants in known cancer genes through power calculations of sample sizes necessary to generate minimally useful mutation-specific relative risk (RR) estimates for rare missense variants.
Current practice in risk estimation: implied risk from classification
Variant classification may imply different levels of risk to patients ( Figure 1 ). In the Plon framework,9 as implemented by current classification schemes, the “Definitely Pathogenic” classification implies risk similar to that reported in the literature for pathogenic mutations. For example, pathogenic variants in the most studied breast and colon cancer risk genes eliminate one functional copy of the gene; risk estimates from well defined cases that completely eliminate one functional copy of the gene represent a theoretical upper limit of risk conferred by a heterozygous variant in a specific gene. “Likely Pathogenic” classification implies that there is enough evidence to conclude that the RR of the variant is >1 and suggests that the RR may be similar to the upper limit defined by definitely pathogenic variants. The classification “Uncertain Significance” implies that the RR may be anywhere from slightly >1 to the upper limit of risk seen in pathogenic variants. The classification “Likely Benign” implies that there is enough evidence to conclude that the RR is unlikely to be as great as the risk for pathogenic variants and that the evidence suggests similar risk to that of the general population but that there is not enough evidence to definitively conclude that the risk is similar to the risk of the general population. The term “Benign” implies RR ˜1 (or very slightly <1 because these individuals lack risk conferred by reported variants). Explicitly illustrating this implied risk framework may be useful to genetic counselors for helping patients visualize and understand variant categories ( Figure 1 ).
Visualization of current standard-of-care-implied cancer relative risk (RR) from variant classification for dominant diseases with incomplete penetrance. Boxes indicate confidence intervals for RR. Solid vertical lines represent point estimates for RR for which data exist. Dotted vertical lines represent assumed point estimates not supported by independent, variant-specific studies. *High risk is specific to both disease and gene and is defined by variants that completely eliminate one functional copy of the gene; this is the theoretical upper limit of risk conferred by a heterozygous variant in a specific gene.
This display of implied risk illustrates how simple classification can be suboptimal for patient management because of the high degree of uncertainty in implied risk for missense and splice variants, even those classified as Likely Pathogenic or Likely Benign. Variant-specific RR estimates, beyond helping classification, allow quantitative estimates of outcome probabilities that are necessary for rational medical planning. Current studies indicate that risk conferred by different missense mutations can vary substantially.18,19,20 However, current classification systems often draw from many sources, including sources that provide no information about clinical outcomes or risk, such as in silico protein predictions, in vitro protein function studies, and cross-species sequence conservation.9,10,11 This is likely to lead to overestimates of risk for many rare variants and creates a genetic counseling dilemma because low-frequency missense variants may be grouped into risk categories before population- or family-based RR data are available.9,11 Furthermore, in the setting of novel genes that have been linked with prevalent and less common cancers, RR estimates may be unavailable for any variant, even those classified as known pathogenic. To ascertain the magnitude of this problem, we evaluated the sample sizes that might be necessary to generate a minimally accurate RR estimate for hypothetical rare variants of uncertain significance using the examples of breast and colon cancer risks.
Materials and Methods
Calculation of sample size needed for minimally useful risk estimates
Risk estimates often come from odds ratios (ORs) generated by case–control studies because OR and RR converge for rare diseases. Another strategy to evaluate variants is to use families with the investigated mutation. We modified standard power calculations to explore sample sizes necessary to determine whether the RR for a novel variant is >1.
We used standard formulas for calculating sample size from allele frequency, modified as described by Fleiss et al. to include continuity correction, and in the case of family data, to permit unequal numbers of affected and unaffected individuals.21,22 The R-script that we used for calculations of population- and family-based sample size is included as Supplementary Materials to facilitate additional power calculations across a wider spectrum of allele frequency, RR, desired power, and ascertainment parameters.
We specifically examined variant population frequencies of 0.1, 0.01, and 0.001%. We performed power calculations for population-based case–control studies and family-based linkage studies across several levels of cancer RR. We used RRs of 12, 6, 3, and 1.5 for breast cancer and 20, 10, 5, and 2.5 for colon cancer. From the literature, we identified 12 as the RR for established breast cancer genes (i.e., BRCA1, BRCA2) and 20 as the RR for established colon cancer genes (i.e., MLH1) and then used regular fractions of these to explore sample size over the spectrum of possible risk.23,24,25 We assumed breast cancer cumulative incidence of 0.08 and colon cancer cumulative incidence of 0.03 for individuals between the ages of 40 and 70 years, who were likely to be included in this type of study.
Population-based sample size calculation
Because variants of clinical interest will be in known genes and will presumably have in silico data available, we assume that in silico data in known cancer genes is equivalent to a pretest probability of 0.9, and we use a Bayesian approach to define thresholds for power calculations, similar to approaches used for variant classification in previous studies.14 Hence, we used desired power of 0.8 and an α of 0.1, which would be consistent with a posttest probability of pathogenicity equaling 99% for a pathogenic variant. We used a one-tailed test to calculate sample size because we are assuming that alleles increase cancer risk. We purposefully used these liberal assumptions, which result in low sample size estimates, because we are considering the situation in which we desire definitive classification and a reasonable independent estimate of RR for rare variants in established cancer genes. The estimated RR will have some degree of error. More conservative assumptions would obviously result in larger sample requirements and more precise RR estimates, which may be desirable in certain clinical or research scenarios. If the measured RR is extremely high, this is not a major concern because the practical upper limit of risk is defined by well studied, highly pathogenic variants. Similarly, the statistical lower bound for RR is 0, but the practical lower limit is 1 because only elevated cancer risk is clinically actionable.
Family-based sample size calculation
For family-based variant classification analysis, several strategies have been proposed to generate likelihood ratios that can be used for multifactorial classification of rare variants.26,27,28,29 Variant classification strategies usually favor genotyping individuals with extreme phenotypes such as distant relatives with cancer at a young age. This strategy takes advantage of the fact that identifying a shared rare allele in an unlikely clinical situation can generate very large likelihood ratios with minimal genotyping. Although this strategy may work well for classifying a variant as pathogenic, it does not create information that can be used to define the RR conferred by the variant. Likelihood-based classification studies may dramatically overestimate risk (the winner’s curse). However, for extremely rare variants, it is unlikely that unrelated carriers can be identified. Despite its drawbacks, a family-based approach may be the only way to estimate RR. However, to mitigate the probability of dramatically overestimating risk, studies of families with novel mutations should recruit individuals related to previously identified carriers of variants without regard for disease status. As noted above for population-based studies, extreme overestimates can be avoided by capping risk at the level defined by common, highly pathogenic variants.
One efficient way to gather the most informative individuals for RR estimates in a family would be to iteratively genotype close relatives of individuals carrying the rare allele starting with the proband (but excluding the proband in calculations to avoid ascertainment bias). Case–family and case–family–control methods have been described previously.29,30,31,32 One would recruit all available family members of appropriate age and gender who are likely to carry the variant of interest, regardless of personal cancer history. The strategy would be to ascertain genotype data for all available first- and second-degree relatives of the proband and repeat this process for newly identified carriers, branching to new first- and second-degree relatives while gathering data on disease status but not skewing recruitment based on these data. When a variant is very rare, first-degree relatives have a 50% chance and second-degree relatives have a 25% chance of being carriers. Alternatively, one could recruit only first-degree relatives of identified carriers, which would require additional iterations of testing, or recruit both near and distant relatives, resulting in a lower variant frequency but potentially fewer stages of iterative recruiting. Regardless of strategy, it should be emphasized that for accurate RR estimates, relatives must be recruited without regard for disease status. To calculate RR, one must phenotype enough individuals (i) with and without the variant and (ii) with and without the disease to generate a meaningful risk ratio.
Confidence intervals for the risk estimates could be computed using linear mixed models to account for within-family genotype and environmental cancer risk correlation.29,33,34 For simplicity in our calculations, we assumed that nongenetic factors influencing cancer risk are uncorrelated and that genetic cancer risk beyond the variant of interest is negligible. We also assumed that the baseline cancer risk in a family is independent of and identical to the risk in a population. This allows definition of the lower bound of sample size for risk estimates with confidence intervals small enough to classify the variant without knowing clinical details about specific families. For our analysis, we assumed that equal numbers of first- and second-degree relatives would be genotyped, resulting in an overall rare variant frequency of 37.5% in the genotyped cohort. We used assumptions similar to those we used for population-based studies: one-sided α of 0.1 and 80% power. As with population-based studies, these are low estimates of sample size because correlation between family members would widen confidence intervals. More accurate power estimates would require more specific disease models, and sample sizes required for adequate power may be substantially higher.
Results
Population-based case–control sample size necessary to define risk for a low-frequency variant of uncertain significance
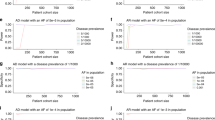
Population-based studies to categorize additional variants that may confer cancer risk will need to be increasingly large as both variant frequency and RR decrease ( Tables 1 and 2 ). When there is very high RR of disease, case–control studies will yield sufficient cases to prove that the variant is pathogenic, but with insufficient controls to accurately calculate ORs because of the extremely low frequency of the mutation in controls; therefore, using case-only analysis and known population disease frequencies from larger samples in the denominator might yield more accurate ORs.
Family-based sample size necessary to define risk for a low-frequency variant of uncertain significance
Family-based studies to categorize additional variants that may confer cancer risk will need to be increasingly large as RR decreases but are robust to changes in variant frequency because rare variant carrier frequency in families is a direct function of relationship to identified carriers ( Tables 3 and 4 ). Note that in the unrelated and familial study designs, the two groups being compared are orthogonal: in population-based studies, the carrier frequency is compared between cases and controls. In families, the affected frequency is compared between carriers and noncarriers.
GALNT12 example
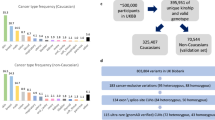
GALNT12 has recently been identified as a colorectal cancer risk gene, but the RRs have not been established for GALNT12 variants. There are 69 exonic missense variants identified by the Exome Variant Server project in ˜6,500 individuals sequenced; 33 of these were missense variants at a frequency <0.001, of which 28 had a frequency <0.0002.35 Some of these rare variants may have been oversampled due to chance and may have actual population frequencies that are much lower.
We recently identified an individual with a GALNT12 D303N mutation. This is present at a frequency of ˜0.001 in both the 1000 Genomes and Exome Variant Server databases. There is limited literature suggesting that this variant may be pathogenic.36,37 However, the literature does not indicate what the RR or OR might be for this variant but suggests that risk may be lower than that for known pathogenic mutations in well defined hereditary nonpolyposis colon cancer genes.36,37 If the RR of colon cancer conferred by this variant is 5, we would expect that a case–control study with ˜1,089 cases and 1,089 cancer-free controls would have a reasonable likelihood of definitively categorizing the variant and generating a reasonable RR estimate ( Table 2 ). We would expect such a study to identify approximately five individuals carrying the variant among cases and one with the variant among the controls. As noted above, using a larger control data set, such as the Exome Variant Server database, may allow more accurate estimates of the OR.35 If we were to calculate risk from family-based studies, and if we can successfully sample relatives of the proband such that 37.5% of genotyped individuals carry the variant in question and are old enough to be at risk of cancer, we would need to genotype 137 relatives of the proband to classify the variant and define a reasonable independent RR estimate. In this process, we would identify ˜11 individuals who have had colon cancer, of whom, 8 would be expected to carry the variant of interest and 3 would be incidental cancer cases. Despite the substantial RR, only a portion of the ˜52 (8 + 44) related individuals carrying the risk variant would have developed cancer ( Table 4 ).
Discussion
Some patients, physicians, and genetic counselors may have the hope that many variants of uncertain significance will be classified in the near future.6 However, despite the liberal assumptions resulting in lower bounds on sample size estimates that we report, it appears unlikely that most very rare missense variants will be classifiable in the near future. Furthermore, accurate RR estimates are more challenging from an epidemiological perspective. Unfortunately, based on sample sizes necessary, independent RR estimates may not be available for most rare variants anytime in the foreseeable future. Functional studies will probably improve and may help with Bayesian classification of some variants; however, because functional assays are usually targeted at specific domains and typically generate likelihood ratios between 1.5 and 10, functional assays for many variants may not be available, and even when these functional assays exist, some epidemiological evidence would probably be required as additional support.17,38,39
Efforts to build large shared databases of cases and population-based controls are promising and may make it possible to classify and estimate risk for the highest-risk variants, i.e., those with nearly 0.1% frequency in the population, such as the GALNT12 variant described above. However, the use of population-based RR calculations may not be feasible for most rare variants. It is unlikely that the enormous research funds required could be made available to do adequate population-based surveys to classify extremely rare variants, but some data may become available from pooled results obtained from clinical testing institutions that are early adopters of genomic methodologies for cancer risk testing.
Family-based analysis requires the same sample size regardless of variant frequency; therefore, despite substantial limitations, this may be the best strategy for classifying extremely rare missense variants, particularly if RR is predicted to be high. However, it will be necessary to identify many distant relatives or multiple apparently unrelated families to classify and estimate RR for most rare variants using families. This may be challenging because average family size has been decreasing in much of the world, knowledge of family medical history is often limited, and obtaining additional family history can be difficult due to geography, family communication, and limited availability of older records. Unfortunately, the probability of finding more than one independent family carrying a rare variant is directly proportional to variant frequency. Although this type of family-based analysis might be feasible in a research setting for highly penetrant genes, in the current funding environment, it is highly unlikely that grant funding will become available for classification of private mutations in already well-characterized genes. From a clinical perspective, identifying enough family members to classify and estimate risk for most rare variants will constitute a heroic genetic counseling effort, and insurance coverage for such testing would be difficult to justify.
The GALNT12 D303N mutation example presented herein is illustrative. Although this specific variant is common enough that it may be definitively classified relatively soon, the risk conferred by this variant may remain unclear even after the variant is definitively classified. Dozens of other rare GALNT12 missense variants have already been identified in fewer than 0.5% of individuals sequenced for this gene.35 It is clear from recent population-based exome- and genome-sequencing projects that the number of rare variants with potential clinical implications identified in the future will increase with the number of individuals receiving genomic testing.4,40
We demonstrate that generating clinically actionable estimates of RR for rare missense variants will be very challenging even after extensive efforts to categorize these as Likely Pathogenic or Pathogenic. This demonstrates a significant limitation to personalized cancer risk estimates based on genetic information.
Disclosure
The authors declare no conflict of interest.
References
Peltomäki P, Vasen H . Mutations associated with HNPCC predisposition – Update of ICG-HNPCC/INSiGHT mutation database. Dis Markers 2004;20:269–276.
Frank TS, Deffenbaugh AM, Reid JE, et al. Clinical characteristics of individuals with germline mutations in BRCA1 and BRCA2: analysis of 10,000 individuals. J Clin Oncol 2002;20:1480–1490.
Goldgar DE, Easton DF, Byrnes GB, Spurdle AB, Iversen ES, Greenblatt MS ; IARC Unclassified Genetic Variants Working Group. Genetic evidence and integration of various data sources for classifying uncertain variants into a single model. Hum Mutat 2008;29:1265–1272.
Tennessen JA, Bigham AW, O’Connor TD, et al.; Broad GO; Seattle GO; NHLBI Exome Sequencing Project. Evolution and functional impact of rare coding variation from deep sequencing of human exomes. Science 2012;337:64–69.
Walsh T, Lee MK, Casadei S, et al. Detection of inherited mutations for breast and ovarian cancer using genomic capture and massively parallel sequencing. Proc Natl Acad Sci USA 2010;107:12629–12633.
Miller-Samuel S, MacDonald DJ, McDonald DJ, et al. Variants of uncertain significance in breast cancer-related genes: real-world implications for a clinical conundrum. Part one: clinical genetics recommendations. Semin Oncol 2011;38:469–480.
O’Neill SC, Rini C, Goldsmith RE, Valdimarsdottir H, Cohen LH, Schwartz MD . Distress among women receiving uninformative BRCA1/2 results: 12-month outcomes. Psychooncology 2009;18:1088–1096.
Culver J, Brinkerhoff C, Clague J, et al. Variants of uncertain significance in BRCA testing: evaluation of surgical decisions, risk perception, and cancer distress. Clin Genet 2013;17:12097.
Plon SE, Eccles DM, Easton D, et al.; IARC Unclassified Genetic Variants Working Group. Sequence variant classification and reporting: recommendations for improving the interpretation of cancer susceptibility genetic test results. Hum Mutat 2008;29:1282–1291.
Rhem HL . Laboratory for Molecular Medicine Variant Classification Rules. Partners Laboratory for Molecular Medicine: Cambridge, MA, 2012. http://pcpgm.partners.org/sites/default/files/LMM/Resources/LMM_VariantClassification_05.26.11.pdf. Accessed 17 July 2013.
Bell J, Bodmer D, Sistermans E, Ramsden SC . Practice guidelines for the interpretation and reporting of unclassified variants (UVs) in clinical molecular genetics. UK Clinical Molecular Genetics Society; 11 January 2008: UK Clinical Molecular Genetics Society and Dutch Society of Clinical Genetics Laboratory Specialists. Unclassified variants good practice meeting, Manchester, UK, 2007.
Lindor NM, Goldgar DE, Tavtigian SV, Plon SE, Couch FJ . BRCA1/2 sequence variants of uncertain significance: a primer for providers to assist in discussions and in medical management. Oncologist 2013;18:518–524.
Dorschner MO, Amendola LM, Turner EH, et al.; National Heart, Lung, and Blood Institute Grand Opportunity Exome Sequencing Project. Actionable, pathogenic incidental findings in 1,000 participants’ exomes. Am J Hum Genet 2013;93:631–640.
Thompson BA, Goldgar DE, Paterson C, et al.; Colon Cancer Family Registry. A multifactorial likelihood model for MMR gene variant classification incorporating probabilities based on sequence bioinformatics and tumor characteristics: a report from the Colon Cancer Family Registry. Hum Mutat 2013;34:200–209.
Bayrak-Toydemir P, McDonald J, Mao R, et al. Likelihood ratios to assess genetic evidence for clinical significance of uncertain variants: hereditary hemorrhagic telangiectasia as a model. Exp Mol Pathol 2008;85:45–49.
Spurdle AB . Clinical relevance of rare germline sequence variants in cancer genes: evolution and application of classification models. Curr Opin Genet Dev 2010;20:315–323.
Lindor NM, Guidugli L, Wang X, et al. A review of a multifactorial probability-based model for classification of BRCA1 and BRCA2 variants of uncertain significance (VUS). Hum Mutat 2012;33:8–21.
Zhang B, Beeghly-Fadiel A, Long J, Zheng W . Genetic variants associated with breast-cancer risk: comprehensive research synopsis, meta-analysis, and epidemiological evidence. Lancet Oncol 2011;12:477–488.
Vink GR, van Asperen CJ, Devilee P, Breuning MH, Bakker E . Unclassified variants in disease-causing genes: nonuniformity of genetic testing and counselling, a proposal for guidelines. Eur J Hum Genet 2005;13:525–527.
Nieuwenhuis MH, Vasen HF . Correlations between mutation site in APC and phenotype of familial adenomatous polyposis (FAP): a review of the literature. Crit Rev Oncol Hematol 2007;61:153–161.
Ziegler A, Konig IR . A Statistical Approach to Genetic Epidemiology. Wiley-VCH Verlag & Co: Weinheim, Germany, 2006.
Fleiss JL, Levin B, Paik MC . Statistical Methods for Rates and Proportions, 3rd edn. Wiley-Interscience: Hoboken, NJ, 2003.
Antoniou A, Pharoah PD, Narod S, et al. Average risks of breast and ovarian cancer associated with BRCA1 or BRCA2 mutations detected in case Series unselected for family history: a combined analysis of 22 studies. Am J Hum Genet 2003;72:1117–1130.
Chen S, Parmigiani G . Meta-analysis of BRCA1 and BRCA2 penetrance. J Clin Oncol 2007;25:1329–1333.
Quehenberger F, Vasen HF, van Houwelingen HC . Risk of colorectal and endometrial cancer for carriers of mutations of the hMLH1 and hMSH2 gene: correction for ascertainment. J Med Genet 2005;42:491–496.
Petersen GM, Parmigiani G, Thomas D . Missense mutations in disease genes: a Bayesian approach to evaluate causality. Am J Hum Genet 1998;62:1516–1524.
Thompson D, Easton DF, Goldgar DE . A full-likelihood method for the evaluation of causality of sequence variants from family data. Am J Hum Genet 2003;73:652–655.
Mohammadi L, Vreeswijk MP, Oldenburg R, et al. A simple method for co-segregation analysis to evaluate the pathogenicity of unclassified variants; BRCA1 and BRCA2 as an example. BMC Cancer 2009;9:211.
Zhao LP, Aragaki C, Hsu L, et al. Integrated designs for gene discovery and characterization. J Natl Cancer Inst Monographs 1999;71–80.
Cui JS, Spurdle AB, Southey MC, et al. Regressive logistic and proportional hazards disease models for within-family analyses of measured genotypes, with application to a CYP17 polymorphism and breast cancer. Genet Epidemiol 2003;24:161–172.
Jenkins MA, Croitoru ME, Monga N, et al. Risk of colorectal cancer in monoallelic and biallelic carriers of MYH mutations: a population-based case-family study. Cancer Epidemiol Biomarkers Prev 2006;15:312–314.
Jenkins MA, Baglietto L, Dowty JG, et al. Cancer risks for mismatch repair gene mutation carriers: a population-based early onset case-family study. Clin Gastroenterol Hepatol 2006;4:489–498.
Zhang Z, Ersoz E, Lai CQ, et al. Mixed linear model approach adapted for genome-wide association studies. Nat Genet 2010;42:355–360.
Kang HM, Sul JH, Service SK, et al. Variance component model to account for sample structure in genome-wide association studies. Nat Genet 2010;42:348–354.
Project NGES. NHLBI Exome Sequencing Project (ESP). Exome Variant Server. Seattle, WA. University of Washington: Seattle, GO, 2013 (updated 7 June 2013; v.0.0.20 (Exome Variant Server). http://evs.gs.washington.edu/EVS/. Accessed 20 August 2013.
Clarke E, Green RC, Green JS, et al. Inherited deleterious variants in GALNT12 are associated with CRC susceptibility. Hum Mutat 2012;33:1056–1058.
Guda K, Moinova H, He J, et al. Inactivating germ-line and somatic mutations in polypeptide N-acetylgalactosaminyltransferase 12 in human colon cancers. Proc Natl Acad Sci USA 2009;106:12921–12925.
Bouwman P, van der Gulden H, van der Heijden I et al. A high-throughput functional complementation assay for classification of BRCA1 missense variants. Cancer Discov 2013;23:23.
Tram E, Savas S, Ozcelik H . Missense variants of uncertain significance (VUS) altering the phosphorylation patterns of BRCA1 and BRCA2. PLoS ONE 2013;8:e62468.
1000 Genomes Project Consortium; Abecasis GR, Altshuler D, Auton A,et al. A map of human genome variation from population-scale sequencing. Nature 2010;467:1061–73.
Acknowledgements
This study was supported in part by the following grants: U01HG006507 (G.P.J.), U01HG006375 (G.P.J.), U01HG007307 (G.P.J.), and HG004960 (B.L.B.)
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Materials
(DOC 27 kb)
Rights and permissions
About this article
Cite this article
Shirts, B., Jacobson, A., Jarvik, G. et al. Large numbers of individuals are required to classify and define risk for rare variants in known cancer risk genes. Genet Med 16, 529–534 (2014). https://doi.org/10.1038/gim.2013.187
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2013.187
Keywords
This article is cited by
-
Family Studies for Classification of Variants of Uncertain Classification: Current Laboratory Clinical Practice and a New Web‐Based Educational Tool
Journal of Genetic Counseling (2016)
-
Pragmatic and Ethical Challenges of Incorporating the Genome into the Electronic Health Record
Current Genetic Medicine Reports (2014)