Abstract
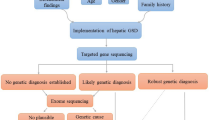
Purpose: Glycogen Storage Disease Type III (limit dextrinosis; Cori or Forbes disease) is an autosomal recessive disorder of glycogen metabolism caused by deficient activity of glycogen debranching enzyme in liver and muscle (Glycogen Storage Disease Type IIIa) or liver only (Glycogen Storage Disease Type IIIb). These two clinically distinct phenotypes are caused by mutations in the same gene (amylo-1,6-glucosidase or AGL). Although most patients with Glycogen Storage Disease Type III have private mutations, common mutations have been identified in some populations, and two specific mutations in exon 3, c.18_19delGA (p.Gln6HisfsX20) and c.16C>T (p.Gln6X), are associated with the Glycogen Storage Disease Type IIIb phenotype.
Methods: To further examine the heterogeneity found in Glycogen Storage Disease Type III patients, we have sequenced the AGL gene in 34 patients with a clinically and/or biochemically confirmed diagnosis of Glycogen Storage Disease Type III.
Results: We have identified 38 different mutations (25 novel and 13 previously reported) and have compiled a list of all mutations previously reported in the literature.
Discussion: We conclude that Glycogen Storage Disease Type III is a highly heterogeneous disorder usually requiring full gene sequencing to identify both pathogenic mutations. The finding of at least one of the two exon 3 mutations in all of the Glycogen Storage Disease Type IIIb patients tested allows for diagnosis of this subtype without the need for a muscle biopsy.
Similar content being viewed by others
Main
Glycogen Storage Disease Type III (GSD III), also known as Cori disease or Forbes disease (OMIM 232400), is an autosomal recessive disorder that results from deficiency of glycogen debranching enzyme (GDE). GDE is unusual in that it contains two catalytic activities—oligo-1,4–1,4-glucanotransferase (EC 2.4.1.25) and amylo-1,6-glucosidase (EC 3.2.1.33)—at two separate sites on a single polypeptide chain.1–3 GDE, together with phosphorylase, is responsible for the complete degradation of stored glycogen.4 Deficiency of GDE results in incomplete glycogenolysis and accumulation of abnormal glycogen, with shorter outer chains (resembling limit dextrin) in affected tissues.
GSD III is associated with a high degree of clinical variability in terms of organ involvement and disease severity. Symptoms of hepatomegaly and hypoglycemia are common to all patients, and many also have short stature and dyslipidemia. Approximately 85% of patients have myopathy, and some may develop cardiomyopathy. These patients are classified as having GSD IIIa. Approximately 15% of patients have only liver involvement and are classified as having GSD IIIb.4 In rare cases, selective loss of only one of the two GDE activities, glucosidase or transferase, results in GSD IIIc or GSD IIId, respectively.5,6 Although GSD IIIc and IIId can be distinguished from other subtypes of GSD III by complicated laboratory testing, these patients are clinically indistinguishable from GSD IIIa.
GSD III can be diagnosed by measurement of GDE activity in liver, muscle, and erythrocytes. Measurement of muscle GDE activity is required for distinguishing GSD IIIa from GSD IIIb because patients with GSD IIIa can have normal muscle function and creatine kinase activity well into adulthood and, thus, can be misclassified as GSD IIIb. Cloning and characterization of the AGL gene, which encodes GDE, have allowed development of DNA-based assays for this disorder and can also help to determine subtype,7–9 knowledge of which is important for clinical management. The AGL gene is composed of 35 exons spanning about 85 kb of genomic DNA.9 The major mRNA isoform in muscle and liver encodes a protein of 1532 amino acids.7,8 To date, 87 pathogenic mutations have been reported, reflecting a high degree of genetic heterogeneity. Most mutations have been reported in only one or a few patients, but there are some mutations that are common to specific ethnic groups,10,11 and two mutations in exon 3, c.16C>T (p.Gln6X) and c.18_19delGA (p.Gln6HisfsX20), previously described as c.17_18delAG, have been exclusively associated with GSD IIIb subtype.4,12,13
We have sequenced the AGL gene in 34 patients with a known or suspected diagnosis of GSD III to further investigate associations between AGL gene mutations and clinical presentation. Most of these GSD III patients have been followed up at the Metabolic Diseases Clinic at Duke University Medical Center, which has expertise in managing patients with this disorder.
MATERIALS AND METHODS
Patients
Twenty-five of the 34 subjects in this study had the diagnosis of GSD III based on clinical symptoms and biochemical measurement of elevated glycogen content, an altered ratio of glucose-1-phosphate/glucose, and evidence of debrancher enzyme (GDE) deficiency in biopsies of liver (n = 20; these subjects could have GSD IIIa or GSD IIIb) or muscle (n = 5; four IIIa and one IIIb). The remaining nine subjects had liver or muscle histology, suggesting the possibility of GSD III or clinical symptoms of GSD III, but GDE enzyme activity was not evaluated. GSD III subtype was determined based on presence or absence of GDE activity in muscle tissue sample and/or the presence of one of the two GSD IIIb-specific pathogenic mutations (c.16C>T [pGln6X] and c.17_18delAG [p.Gln6HisfsX25]) in a patient (n = 8), as has been described before.12,13,4 Our patient cohort is diverse, consisting of non-Hispanic Caucasian of mixed European ancestry (n = 17), Hispanic of Mexican and Latin American ancestry (n = 4), mixed European/Hispanic (n = 2), Southeast Asian of Indian and Pakistani ancestry (n = 6), African American (n = 2), Pacific Islander (n = 1), mixed African American/Korean (n = 1), and unknown ethnicity (n = 1). This study was conducted in accordance with the Institutional Review Board requirements of Duke University Medical Center.
AGL gene analysis
Most patients underwent full AGL gene sequencing (n = 32); however, two patients had exon 3 sequencing only but were lost to follow-up. All 35 coding exons and flanking intronic sequences of the AGL gene were amplified by polymerase chain reaction (PCR) using isolated genomic DNA. PCR was performed using HotStart-IT Taq Master Mix (2×; USB Corporation, Cleveland, OH) in a 30-μL reaction containing 50–500 ng genomic DNA and 0.3 μM primer mix. All PCR primers were tailed with M13 universal primers at the 5′ end (Table 1). Exons 7, 8, 17, 18, 21, 25, 29, and 31 were amplified by 95°C for 2 minutes; then 18 cycles of 94°C for 15 seconds, 54°C for 45 seconds, 72°C for 1 minute; followed by 22 cycles of 94°C for 15 seconds and 70°C for 1 minute; then 72°C for 10 minutes and 4°C hold. All other exons were amplified by 95°C for 2 minutes; 16 cycles of 94°C for 15 seconds, 58°C for 30 seconds, and 72°C for 1 minute; followed by 22 cycles of 94°C for 15 seconds and 70°C for 1 minute; then 72°C for 10 minutes and 4°C hold. Amplicons were treated with ExoSAP-IT (USB Corp.), and all except exon 27 were bidirectionally sequenced, using 0.6 μM M13 forward/−20 or M13 reverse/−27 primer with the Big Dye Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems, Foster City, CA). Exon 27 was sequenced only in the forward direction because of a poly-T stretch in intron 27. Sequence products were treated with the Big Dye XTerminator Purification Kit (Applied Biosystems) and electrophoresed on the ABI 3130xl Genetic Analyzer (Applied Biosystems). Chromatograms were base called, assembled, and aligned using SeqScape software v2.5 (Applied Biosystems). Sequences were compared with the AGL isoform 1 reference sequence (GenBank Accession: NM_000028.2).
To determine if any novel missense mutations identified in this study are common variants, we searched the NCBI single nucleotide polymorphism (SNP) database (http://www.ncbi.nlm.nih.gov/snp). The pathogenicity of novel missense mutations was further investigated by determining the level of conservation of the altered amino acid. Amino acid alignments were produced by ClustalW (http://www.ebi.ac.uk/Tools/clustalw2/index.html). PolyPhen and SIFT programs were used to predict the damaging effect of amino acid changes.14–16
To identify previously published mutations, a search of the PubMed database was performed. Only articles written in English were included. For mutations for which only the nucleotide change was reported, the effect on protein was determined.
Nomenclature used to describe the mutations identified in study subjects and those previously published in the literature follows published nomenclature guidelines.17,18 On the basis of the recommendation to assign a nucleotide change to the most 3′ position possible, the GSD IIIb mutation, previously described as c.17_18delAG,12 is described as c.18_19delGA.
RESULTS
Mutations identified in this study
In our 34 patients with a confirmed or suspected diagnosis of GSD III, we were able to identify 65 of the expected 68 pathogenic alleles. For three subjects, only one mutation could be identified. Two of these subjects had only targeted mutation analysis of exon 3 performed. These two patients were lost to follow-up, thus, could not be consented for full gene sequencing. The other patient had full gene sequencing performed, but only one mutation was identified. We assume that the second mutation is either a large deletion mutation that could not be detected by routine gene sequencing or that the mutation resides in an area of the gene not analyzed, such as the promoter or an intronic region.
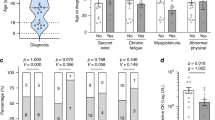
Thirty-eight different mutations were identified in our subjects; 13 of these have been previously identified, whereas 25 have not been reported before, to our knowledge (Fig. 1, Table, Supplemental Digital Content 1, http://links.lww.com/GIM/A102). Of these mutations, the majority, including nonsense mutations, and deletions and duplications that create a reading frameshift, are expected to disrupt production of GDE protein by creating premature termination codons. In addition, we identified four novel splice site mutations. Although the effect of these mutations on splicing has not been studied, they are all expected to disrupt normal splicing by altering the splice junction consensus sequences.19
Diagrammatic presentation of mutations found in our patient population (38 mutations), based on Ref. 4. Mutations denoted by “superscript a” were found in patients with two other pathogenic mutations. Mutations with “*” denote novel mutations. Three prominent domains in the GDE protein are potentially encoded by: Transferase domain, exons 6 and 13–15; Glucosidase domain, exons 26 and 27; Glycogen-binding domain, exons 31–34.3,4,7,45
Common mutations identified
The most common changes identified in our unrelated Caucasian non-Hispanic subjects (n = 17) were p.Arg864X and c.2309-1G>A, each of which had an allele frequency of 11.8% (n = 4 of 34 alleles). We also identified c.4260-12A>G (IVS32-12A>G) in 5.9% of alleles from the unrelated Caucasian non-Hispanic subjects. Two of the four unrelated Hispanic patients (one Mexican and one with unknown country of origin) were homozygous for c.1384delG (p.Val462X), indicating that this may be a common genetic change in the Hispanic population (allele frequency = 50%). This mutation was not found in the two mixed non-Hispanic/Hispanic Caucasian subjects. None of the Southeast Asian subjects (n = 6) in this study shared the same mutation, which suggests that a common mutation may not exist in this population. Eight patients were classified as GSD IIIb based on the clinical presentation or enzyme analysis and common mutation testing. Of these, two were heterozygous for c.16C>T (p.Gln6X), five were heterozygous for c.18_19delGA (p.Gln6HisfsX20), and one was homozygous for c.18_19delGA (p.Gln6HisfsX20).
Types of mutations identified
Missense mutations
Five novel missense mutations were found in this study (p.Gly138Glu, p.Ala253Pro, p.Leu400Pro, p.Arg494His, and p.Cys1515Arg). None of these substitutions was found in the NCBI SNP database (http://www.ncbi.nlm.nih.gov/snp). In all the cases, the altered amino acid is highly conserved between species, suggesting that the altered amino acids perform an important function (Fig. 2). The PolyPhen program14 predicted that p.Gly138Glu, p.Leu400Pro, p.Arg494His, and p.Cys1515Arg are “probably damaging,” and p.Ala253Pro is “possibly damaging,” and the SIFT program15,16 predicted all of these mutations to be “not tolerated.” Based on the clinical presentation and predicted pathogenic nature of these missense changes, and taking into account that the patients were either homozygous or compound heterozygotes for these mutations, we assume that these substitutions could be adversely affecting AGL function. Interestingly, the missense change p.Arg494His was identified in a patient with two other known pathogenic changes, and therefore the clinical effect of this mutation is unclear.
Amino acid sequences surrounding human AGL amino acid 138, 253, 400, 494, and 1515 and alignment with orthologous sequences in human (Homo sapiens), NP_000019; chimpanzee (Pan troglodytes), XP_524777; cow (Bos taurus), XP_595566; horse (Equus callabus), NP_001103778; rabbit (Oryctolagus cuniculus), NP_001075716; dog (Canis familiaris), NP_001041561; mouse (Mus musculus), NP_001074795; rat (Rattus norvegicus), NP_001102034; chicken (Gallus gallus), XP_422317; and zebra fish (Danio rerio), XP_696194, predicted proteins.
Intronic changes
In our subjects, we also identified six novel intronic changes (c.83-93C>G, c.406+81T>G, c.1612-24C>A, c.3260-19G>A, c.3260-20T>C, and c.4481+85G>T) none of which is listed in the SNP database.
Although these changes are thought to be benign because they are far into the intronic regions, the possibility that they affect splicing or gene expression cannot be ruled out without further studies. Numerous previously reported polymorphic variants were also found in our subjects (data not shown).
Variants founds in patients with two other pathogenic mutations
In two subjects (one from this study and one previously published12), c.966dup98 in exon 9 was found in addition to two known pathogenic variants. Both patients also carried c.2590C>T (p.Arg864X) and an additional mutation (c.4529insA in one patient and c.18_19delGA in the other). Finding c.966dup68 only in patients with c.2590C>T raises the possibility that these sequence changes exist in cis position. Results of parental studies, which could confirm this, are not available. We have not found the c.966dup68 alteration in any other patients or controls. Although this duplication is expected to cause a pathogenic frameshift change, the clinical effect cannot be determined because both subjects in whom it was found have two other pathogenic mutations.
AGL gene exon 3 mutations
Our one subject with an enzymatically confirmed diagnosis of GSD IIIIb had one of the exon 3 mutations (c.18_19delGA) previously associated with GSD IIIb.12 In addition, seven patients with an undefined GSD III subtype were also found to be heterozygous (n = 6) or homozygous (n = 1) for one of the common IIIb mutations. None of these patients has any evidence for muscle disease, including any sign of myopathy or elevated creatine kinase level, consistent with the GSD IIIb subtype.
Muscle GDE activity in subjects with GSD IIIb
Of note, muscle enzyme results in our subject with enzymatically confirmed GSD IIb, and in a previously reported subject,12 were 0.1 μmol/hour/g tissue (control range is 0.28 ± 0.08 μmol/hour/g tissue) and, thus, are about half of the normal level. This level of activity may reflect that these patients have one allele that is nonfunctional in muscle and one, with an exon 3 mutation, which is expressed, similar to being a carrier but on a tissue-specific level. In addition, glycogen content and glycogen structure, which are overtly abnormal in the muscle of patients with GSD IIIa, were normal or near normal in the muscle of these GSD IIb patients.
Glycogen content and structure were normal or near normal in the muscle of another two subjects, both of whom had an exon III (GSD IIIb) mutation. However, muscle GDE appeared to be deficient for both of these subjects. The internal control indicated poor sample quality for both subjects. These results suggest that it is important to consider glycogen content and structure, and GDE activity, when distinguishing between GSD III clinical subtypes.
Previously reported mutations in the AGL gene
Shen and Chen4 identified 31 different mutations in the AGL gene in their 2002 literature review. Since that time, the number of mutations identified and reported for AGL gene in the literature has risen to 87 (Table, Supplemental Digital Content 1, http://links.lww.com/GIM/A102). By adding the 25 novel mutations identified by this study to that, the total number of mutations identified in the AGL gene so far becomes 112. The mutations range from nonsense (n = 27), missense (n = 16), intronic splicing mutations (n = 19), small insertions and deletions causing frameshifts (n = 43), small inframe deletions/insertions (n = 3), and larger deletion/duplications/insertions/complex rearrangements (n = 4).
DISCUSSION
Cloning and molecular characterization of the AGL gene has allowed for investigations into the genetic heterogeneity in GSD type III. Our review of published mutations (Table, Supplemental Digital Content 1, http://links.lww.com/GIM/A102) indicates that a high degree of genetic heterogeneity exists in patients with GSD III. Most mutations in the AGL gene occur in only one or a couple of patients, except for a handful of mutations that are associated with a specific ethnic group or clinical subtype (GSD IIIb). In studies of patients from the United States, the most common mutations identified are p.Arg864X (10.3%), c.3964delT (6.7%), c.4260-12A>G (IVS32-12A>G; 5.5%), and p.Arg1228X (5.2%), but they account for only 28% of all alleles.4 We found the p.Arg864X and c.4260-12A>G mutations at similar frequencies (11.8% and 5.9% of alleles, respectively). Interestingly, we did not identify either the p.Arg1228X or c.3965delT mutations in any of our Caucasian subjects, possibly because of sample size. However, we did find that a novel splice site mutation, c.2309-1G>A, represented 11.8% of Caucasian non-Hispanic alleles and that c.1384delG (p.Val462X) represented ∼50% of Hispanic alleles, suggesting that these mutations may be common in these populations.
A previous study of North African Jewish families identified c.4455delT in all 12 patients studied (10 families),10 consistent with a proposed founder effect and possibly explaining the high incidence (1 in 5400) of GSD III in this population.4 Similarly, six children with GSD IIIa from the Faroe Islands were found to be homozygous for p.Arg408X, and 9 heterozygotes were identified out of 272 newborns screened. This founder mutation explains the high prevalence of GSD IIIa in the Faroe islands, 1 in 3100, which is the highest in the world.11 In Italian and Mediterranean patients, c.2681+G>A (IVS21+1G>A) and c.664+3A>G (IVS6+3A>G) have been found in up to 28% and 11.7% of patients, respectively.20–22 Recently, p.Trp1327X was reported as a founder mutation in Turkish patients with GSD III.23
Some mutations seem to be recurrent as they occur in patients of different ethnicities. For example, the c.4260–12G>A (IVS32–12G>A) mutation, which is common in North American patients,13 was also identified in a Japanese patient and a Chinese patient24,25; the p.Arg408X mutation that is common in the Faroese population was also identified in a Chinese patient26; and p.Trp1327X, which has been reported as a founder mutation in Turkish patients,23 has also been identified in a Tunisian,21 an Egyptian,27 a Canadian patient,28 and a German-Ukraine family.29 In all of these cases, haplotype analysis suggests that each of these mutations arose independently at least twice. One of the Asian Indian subjects in this study is homozygous for a mutation, c.2681+1G>A (IVS21+1G>A), which has been previously published in many patients from different ethnicities and, thus, may be a recurrent change.30,31,21,20 Haplotype analysis has not been performed to confirm this. As some mutations appear to be recurrent in different population groups, these may be mutation hotspots. Further work needs to be done to explore this possibility. Because AGL is a recently cloned and large gene, this information is not currently available.
Numerous polymorphisms have also been identified in the AGL gene4 (NCBI SNP database). Most of these variants are intronic, but some are located in exons and result in amino acid substitutions. Of note, with increasing knowledge, some amino acid changes that were originally classified as mutations have been reclassified as benign changes. For example, p.Thr38Ala was found in homozygous form in a patient with no other pathogenic sequence changes and was believed to be a causative mutation.21 This is now thought to be a benign variant based on the current SNP database and prediction that it is “benign” by the Polyphen program.14
Location of mutations/AGL domain structure
Although many of the mutations in the AGL gene are expected to result in production of no protein, missense mutations and small inframe deletions and insertions may lead to production of a polypeptide, but one with reduced or no function because of poor folding of essential domain structures. Only 16 missense mutations, two small inframe deletions, and one small inframe insertion have been reported in the AGL gene (Table, Supplemental Digital Content 1, http://links.lww.com/GIM/A102). However, these amino acid sequence changes can help elucidate the role that specific regions of the GDE protein play in its function and regulation.
Putative transferase and glucosidase catalytic domains and carbohydrate-binding domains (CBDs) have been identified for GDE (Fig. 1).4 A recent study reported the in vitro effects of four mutations in GDE. The p.Leu620Pro mutation in the transferase domain leads to complete loss of this activity but retention of residual glucosidase activity, whereas the p.Arg1147Gly mutation in the glucosidase domain primarily affects glucosidase function.23,32 Two mutations in the CBD (p.Tyr1445ins and p.Gly1448Arg) have a more severe effect on GDE function, abolishing carbohydrate binding and both the enzymatic activities.32 Mutations in the CBD lead to rapid degradation of GDE indicating that the CBD may stabilize GDE. CBD mutations were also shown to result in increased ubiquitination. Thus, the CBD is thought to have a role in coordinating the activities of GDE and in its regulation by the ubiquitin-protease system.32
Many missense mutations identified in GSD III patients are outside the transferase domain, glucosidase domain, and CBDs. These amino acid changes could affect protein function by altering protein tertiary structure and stability in general.
GSD IIIb and mutations in exon 3 of the AGL gene
GSD IIIa and IIIb have previously been shown to be allelic disorders, with two mutations in exon 3 (c.16C>T and c.18_19delGA) being strongly associated with GSD IIIb.4,12,13 Neither of the exon 3 mutations has been identified in patients with GSD IIIa or GSD IIId, suggesting that they are exclusively associated with GSD IIIb. Evidence to date shows that most, if not all, patients with GSD IIIb will have one of these mutations.4 In the initial report, 12 of 13 patients had one of these mutations (10 with c.18_19delGA and 2 with c.16C>T). The one patient with neither mutation was later reclassified as having GSD IIIa.13 Another patient was reported to have GSD IIIb, but no exon 3 mutation was identified.25 Although this patient had no clinical evidence for muscle involvement, muscle GDE activity was not measured, and thus a diagnosis of GSD IIIa could not be excluded. As this patient was homozygous for a milder mutation, c.4260-12A>G (IVS32-12A>G), it is possible that he may develop muscle symptoms later in life.4 In our study, the allele frequency of the two common GSD IIIb mutations was similar to what has been previously reported.4
The finding of an exon 3 mutation (c.18_19delGA) in our subject with enzymatically confirmed GSD IIIb further indicates that this mutation confers the GSD IIIb phenotype. The exclusive association of the exon 3 mutations with the GSD IIIb phenotype is useful as it precludes the need for a muscle biopsy in determining subtype. The second mutation in patients with GSD IIIb can occur anywhere else in the gene. Many of the second mutations found in patients with GSD IIIb have also been found in patients with GSD IIIa, suggesting that these mutations do not affect subtype.
It is not fully understood how the exon 3 mutations result in functional GDE in muscle but not in liver. Expression from the GSD IIIb allele must somehow be sufficient to prevent muscle symptoms. One possible explanation is the use of alternative exons in the AGL gene that allow the mutation to be bypassed.12 For example, exons 3′ and 4′ are alternative exons in the AGL gene, which are present in minor isoforms in both liver and muscle. Similar to exon 3, these exons contain a transcription initiation site and a Kozak sequence in frame with the rest of the GDE-coding sequence. It is possible that the GSD IIIb mutations in exon 3 somehow activate translation beginning at one of these downstream ATGs. Why the effects should be different in liver and muscle is not known. Use of these alternative exons would result in a small N-terminal deletion of GDE, but it is possible that this would not affect activity.4 Another possibility is that highly truncated peptides are produced by the c.16C>T and c.18_19delGA alleles and they somehow activate the expression of muscle-specific isoforms of GDE.4,12
Genotype/phenotype correlation
Other than the association between the exon 3 mutations (c.16C>T and c.18_19delGA) and GSD IIIb, few genotype/phenotype correlations have been made. Because most subjects are compound heterozygotes, it is not possible to predict the clinical effect of most mutations. However, previous reports suggest that some mutations are associated with a severe phenotype including c.3965delT13 and c.4529insA.33 Other mutations, such as c.4260-12A>G (IVS32–12A>G) appear to confer milder symptoms.13 In addition, clinical variability has been noted between patients who have the same genotype, suggesting that other genetic and environmental factors also affect clinical presentation.22 Future work may help to uncover additional correlations between specific AGL mutations and disease severity. For example, although liver disease usually improves with age in patients with GSD III, there are reports of patients with liver cirrhosis, with some progressing to liver failure, and cases of adenoma and hepatocellular carcinoma (HCC).34–40 Mutations have been identified in two patients with HCC39 (Table, Supplemental Digital Content 1, http://links.lww.com/GIM/A102). Whether these mutations are a predisposing factor in the development of HCC is not yet known, but such knowledge would be of great value in providing an appropriate level of screening based on risk.
Predicted effects of mutations in the AGL gene
The majority of mutations identified in the AGL gene (all nonsense and frameshift and some splice site mutations) are expected to result in premature termination codons and, thus, may be subject to nonsense-mediated mRNA decay.41 This is in agreement with previous results of Western blot analysis in GSD III patients, which found no detectable GDE protein.5
Although we have classified the mutations in Table, Supplemental Digital Content 1, http://links.lww.com/GIM/A102 according to the expected effect on mRNA processing and amino acid sequence, we appreciate that predictions for most of these mutations have been made using computer models available to date and have not been tested in vitro. In addition to these predicted effects, exonic splice enhancers (ESEs)42,43 seem to be common, and alterations to these elements may also affect splicing. Although most of the mutations reported in Table 1 are predicted to be frameshift, missense, or nonsense mutations, it is possible that they could alter ESEs, as has already been reported in GSD III; the mutation, c.853C>T in exon 8 (p.Arg285X), was shown to alter the recognition sequence for a SF2/ASF ESE and cause skipping of exon 8.44
CONCLUSION
GSD III is caused by mutations in the AGL gene resulting in GDE deficiency in liver and muscle (GSD IIIa) or liver only (GSD IIIb). GSD III is a highly genetically heterogeneous condition with >100 different mutations now identified. As a result of the high level of genetic heterogeneity, full gene sequencing is typically required to identify both mutations in the AGL gene. Most of the reported mutations in the AGL gene are expected to result in a premature stop codon, making mRNAs a target for nonsense-mediated decay. Patients with GSD IIIb have either the c.16C>T (p.Gln6X) or c.18_19delGA (p.Gln6Hisfx20) mutation in exon 3 of the AGL gene, along with a second mutation elsewhere in the AGL gene. These two exon 3 mutations are specifically associated with GSD IIIb, allowing for diagnosis of GSD IIIb without the need for a muscle biopsy.
References
Gordon RB, Brown DH, Brown BI . Preparation and properties of the glycogen-debranching enzyme from rabbit liver. Biochim Biophys Acta 1972; 289: 97–107.
Bates EJ, Heaton GM, Taylor C, Kernohan JC, Cohen P . Debranching enzyme from rabbit skeletal muscle; evidence for the location of two active centres on a single polypeptide chain. FEBS Lett 1975; 58: 181–185.
Liu W, Madsen NB, Braun C, Withers SG . Reassessment of the catalytic mechanism of glycogen debranching enzyme. Biochemistry 1991; 30: 1419–1424.
Shen JJ, Chen YT . Molecular characterization of glycogen storage disease type III. Curr Mol Med 2002; 2: 167–175.
Ding JH, de Barsy T, Brown BI, Coleman RA, Chen YT . Immunoblot analyses of glycogen debranching enzyme in different subtypes of glycogen storage disease type III. J Pediatr 1990; 116: 95–100.
Van Hoof F, Hers G . The subgroups of type 3 glycogenosis. Eur J Biochem 1967; 2: 265–270.
Yang BZ, Ding JH, Enghild JJ, Bao Y, Chen YT . Molecular cloning and nucleotide sequence of cDNA encoding human muscle glycogen debranching enzyme. J Biol Chem 1992; 267: 9294–9299.
Bao Y, Yang BZ, Dawson TL Jr, Chen YT . Isolation and nucleotide sequence of human liver glycogen debranching enzyme mRNA: identification of multiple tissue-specific isoforms. Gene 1997; 197: 389–398.
Bao Y, Dawson TL Jr, Chen YT . Human glycogen debranching enzyme gene (AGL): complete structural organization and characterization of the 5′ flanking region. Genomics 1996; 38: 155–165.
Parvari R, Moses S, Shen J, Hershkovitz E, Lerner A, Chen YT . A single-base deletion in the 3′-coding region of glycogen-debranching enzyme is prevalent in glycogen storage disease type IIIA in a population of North African Jewish patients. Eur J Hum Genet 1997; 5: 266–270.
Santer R, Kinner M, Steuerwald U, et al. Molecular genetic basis and prevalence of glycogen storage disease type IIIA in the Faroe Islands. Eur J Hum Genet 2001; 9: 388–391.
Shen J, Bao Y, Liu HM, Lee P, Leonard JV, Chen YT . Mutations in exon 3 of the glycogen debranching enzyme gene are associated with glycogen storage disease type III that is differentially expressed in liver and muscle. J Clin Invest 1996; 98: 352–357.
Shaiu WL, Kishnani PS, Shen J, Liu HM, Chen YT . Genotype-phenotype correlation in two frequent mutations and mutation update in type III glycogen storage disease. Mol Genet Metab 2000; 69: 16–23.
Ramensky V, Bork P, Sunyaev S . Human non-synonymous SNPs: server and survey. Nucleic Acids Res 2002; 30: 3894–3900.
Kumar P, Henikoff S, Ng PC . Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat Protoc 2009; 4: 1073–1081.
Ng PC, Henikoff S . SIFT: predicting amino acid changes that affect protein function. Nucleic Acids Res 2003; 31: 3812–3814.
Antonarakis SE . Recommendations for a nomenclature system for human gene mutations. Nomenclature Working Group. Hum Mutat 1998; 11: 1–3.
den Dunnen JT, Antonarakis SE . Mutation nomenclature extensions and suggestions to describe complex mutations: a discussion. Hum Mutat 2000; 15: 7–12.
Shapiro MB, Senapathy P . RNA splice junctions of different classes of eukaryotes: sequence statistics and functional implications in gene expression. Nucleic Acids Res 1987; 15: 7155–7174.
Lucchiari S, Fogh I, Prelle A, et al. Clinical and genetic variability of glycogen storage disease type IIIa: seven novel AGL gene mutations in the Mediterranean area. Am J Med Genet 2002; 109: 183–190.
Lucchiari S, Donati MA, Parini R, et al. Molecular characterisation of GSD III subjects and identification of six novel mutations in AGL. Hum Mutat 2002; 20: 480.
Lucchiari S, Santoro D, Pagliarani S, Comi GP . Clinical, biochemical and genetic features of glycogen debranching enzyme deficiency. Acta Myol 2007; 26: 72–74.
Aoyama Y, Ozer I, Demirkol M, et al. Molecular features of 23 patients with glycogen storage disease type III in Turkey: a novel mutation p.R1147G associated with isolated glucosidase deficiency, along with 9 AGL mutations. J Hum Genet 2009; 54: 681–686.
Horinishi A, Okubo M, Tang NL, et al. Mutational and haplotype analysis of AGL in patients with glycogen storage disease type III. J Hum Genet 2002; 47: 55–59.
Okubo M, Horinishi A, Nakamura N, et al. A novel point mutation in an acceptor splice site of intron 32 (IVS32 A-12–>G) but no exon 3 mutations in the glycogen debranching enzyme gene in a homozygous patient with glycogen storage disease type IIIb. Hum Genet 1998; 102: 1–5.
Lam CW, Lee AT, Lam YY, et al. DNA-based subtyping of glycogen storage disease type III: mutation and haplotype analysis of the AGL gene in Chinese. Mol Genet Metab 2004; 83: 271–275.
Endo Y, Fateen E, Aoyama Y, et al. Molecular characterization of Egyptian patients with glycogen storage disease type IIIa. J Hum Genet 2005; 50: 538–542.
Endo Y, Horinishi A, Vorgerd M, et al. Molecular analysis of the AGL gene: heterogeneity of mutations in patients with glycogen storage disease type III from Germany, Canada, Afghanistan, Iran, and Turkey. J Hum Genet 2006; 51: 958–963.
Schoser B, Glaser D, Muller-Hocker J . Clinicopathological analysis of the homozygous p.W1327X AGL mutation in glycogen storage disease type 3. Am J Med Genet A 2008; 146A: 2911–2915.
Hadjigeorgiou GM, Comi GP, Bordoni A, et al. Novel donor splice site mutations of AGL gene in glycogen storage disease type IIIa. J Inherit Metab Dis 1999; 22: 762–763.
Uotani S, Yamasaki H, Takino H, et al. Identification of a 5′ splice junction mutation in the debranching enzyme gene in a Japanese patient with glycogen storage disease type IIIa. J Inherit Metab Dis 2000; 23: 527–528.
Cheng A, Zhang M, Okubo M, Omichi K, Saltiel AR . Distinct mutations in the glycogen debranching enzyme found in glycogen storage disease type III lead to impairment in diverse cellular functions. Hum Mol Genet 2009; 18: 2045–2052.
Shen J, Bao Y, Chen YT . A nonsense mutation due to a single base insertion in the 3′-coding region of glycogen debranching enzyme gene associated with a severe phenotype in a patient with glycogen storage disease type IIIa. Hum Mutat 1997; 9: 37–40.
Labrune P, Trioche P, Duvaltier I, Chevalier P, Odievre M . Hepatocellular adenomas in glycogen storage disease type I and III: a series of 43 patients and review of the literature. J Pediatr Gastroenterol Nutr 1997; 24: 276–279.
Matern D, Starzl TE, Arnaout W, et al. Liver transplantation for glycogen storage disease types I, III, and IV. Eur J Pediatr 1999; 158( suppl 2): S43–S48.
Shimizu J, Shiraishi H, Sakurabayashi S, et al. [A report on an adult case of type III glycogenosis with primary liver cancer and liver cirrhosis]. Nippon Shokakibyo Gakkai Zasshi 1982; 79: 2328–2332.
Siciliano M, De Candia E, Ballarin S, et al. Hepatocellular carcinoma complicating liver cirrhosis in type IIIa glycogen storage disease. J Clin Gastroenterol 2000; 31: 80–82.
Haagsma EB, Smit GP, Niezen-Koning KE, Gouw AS, Meerman L, Slooff MJ . Type IIIb glycogen storage disease associated with end-stage cirrhosis and hepatocellular carcinoma. The Liver Transplant Group. Hepatology 1997; 25: 537–540.
Demo E, Frush D, Gottfried M, et al. Glycogen storage disease type III-hepatocellular carcinoma a long-term complication?. J Hepatol 2007; 46: 492–498.
Cosme A, Montalvo I, Sanchez J, et al. [Type III glycogen storage disease associated with hepatocellular carcinoma]. Gastroenterol Hepatol 2005; 28: 622–625.
Silva AL, Romao L . The mammalian nonsense-mediated mRNA decay pathway: to decay or not to decay! Which players make the decision?. FEBS lett 2009; 583: 499–505.
Liu HX, Cartegni L, Zhang MQ, Krainer AR . A mechanism for exon skipping caused by nonsense or missense mutations in BRCA1 and other genes. Nat Genet 2001; 27: 55–58.
Cartegni L, Chew SL, Krainer AR . Listening to silence and understanding nonsense: exonic mutations that affect splicing. Nat Rev Genet 2002; 3: 285–298.
Lucchiari S, Pagliarani S, Salani S, et al. Hepatic and neuromuscular forms of glycogenosis type III: nine mutations in AGL. Hum Mutat 2006; 27: 600–601.
Okubo M, Horinishi A, Takeuchi M, et al. Heterogeneous mutations in the glycogen-debranching enzyme gene are responsible for glycogen storage disease type IIIa in Japan. Hum Genet 2000; 106: 108–115.
Acknowledgements
Funding for this project was provided, in part, by the Association of Glycogen Storage Disease, USA, and by the National Institutes of Health through the Duke Clinical Research Center. We thank Dr. Y-T Chen and Denise Peterson for their continued support and endless help in diagnosing our GSD III patients. We also thank the patients and their families for their participation in our research studies.
Author information
Authors and Affiliations
Corresponding author
Additional information
Disclosure: The authors declare no conflicts of interest.
Supplemental digital content is available for this article. Direct URL citations appear in the printed text and are provided in the HTML and PDF versions of this article on the journal's Web site (www.geneticsinmedicine.org).
Rights and permissions
About this article
Cite this article
Goldstein, J., Austin, S., Boyette, K. et al. Molecular analysis of the AGL gene: Identification of 25 novel mutations and evidence of genetic heterogeneity in patients with Glycogen Storage Disease Type III. Genet Med 12, 424–430 (2010). https://doi.org/10.1097/GIM.0b013e3181d94eaa
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1097/GIM.0b013e3181d94eaa
Keywords
This article is cited by
-
The biallelic novel pathogenic variants in AGL gene in a chinese patient with glycogen storage disease type III
BMC Pediatrics (2022)
-
Intron retention is among six unreported AGL mutations identified in Malaysian GSD III patients
Genes & Genomics (2019)
-
Genetic analysis and clinical assessment of four patients with Glycogen Storage Disease Type IIIa in China
BMC Medical Genetics (2018)
-
Hepatic Manifestations in Glycogen Storage Disease Type III
Current Pathobiology Reports (2018)
-
Crystal structure of glycogen debranching enzyme and insights into its catalysis and disease-causing mutations
Nature Communications (2016)