Abstract
Purpose: To determine the sensitivity of whole-genome oligonucleotide array comparative genomic hybridization for the detection of mosaic cytogenetic abnormalities.
Methods: Mosaicism sensitivity was evaluated by testing artificially derived whole chromosome and segmental aneuploidies ranging from 0% to 100% abnormal and additional naturally occurring mosaic specimens.
Results: Using combined dye-reversed replicates and an unfiltered analysis, oligonucleotide array comparative genomic hybridization detected as low as 10% and 20–30% mosaicism from whole chromosome and segmental aneuploidies, respectively. To investigate discrepancies between cultured and uncultured specimens, array comparative genomic hybridization was performed on DNA from additional direct product of conception specimens with abnormal karyotypes in culture. Interestingly, 5 of 10 product of conception specimens with double trisomies on cultured cell analysis had only a single trisomy by array comparative genomic hybridization and quantitative polymerase chain reaction on DNA from the uncultured direct specimen, and all harbored the more commonly observed trisomy. Thus, oligonucleotide array comparative genomic hybridization revealed previously unidentified placental mosaicism in half of the products of conception with double-aneuploid conventional karyotypes.
Conclusion: Oligonucleotide array comparative genomic hybridization can detect low-level mosaicism for whole chromosome (∼10%) and segmental (∼20–30%) aneuploidies when using specific detection criteria. In addition, careful interpretation is required when performing array comparative genomic hybridization on DNA isolated from direct specimens as the results may differ from the cultured chromosome analysis.
Similar content being viewed by others
Main
One of the major causes of developmental delay, mental retardation, and multiple congenital anomalies are pathogenic genomic imbalances, which are routinely evaluated by conventional cytogenetic methods in clinical laboratories. However, genomic aberrations must be larger than 3–5 Mb to be detected by standard GTG-based chromosome banding. Molecular cytogenetic techniques, such as fluorescence in situ hybridization (FISH), have significantly higher resolution (∼100 kb) but are limited because relatively few loci can be interrogated in a single experiment. In contrast, microarray-based comparative genomic hybridization (aCGH) offers a markedly higher resolution and genome-wide assessment and is increasingly being used for clinical evaluation of patients with developmental delay, mental retardation, and multiple congenital anomalies.1–8 Recognition of the value of aCGH as a cytogenetic tool has resulted in reports detailing the validation and use of both bacterial artificial chromosome (BAC) and oligonucleotide-based microarray platforms for postnatal diagnostic evaluation.2,9–14
Recently, the American College of Medical Genetics15 and a European consortium16 published standards and guidelines for the use of aCGH in evaluating constitutional cytogenetic abnormalities. These guidelines detailed the proper use of analytic standards and assay validation for both BAC- and oligonucleotide-based platforms. Important components of aCGH validation included testing with abnormal controls that cover the genomic regions represented on the array and determining array sensitivity to assess its ability to detect mosaicism. Although multiple reports have validated aCGH platforms by thorough testing with control specimens,2,9–14 relatively few have reported on array sensitivity,17,18 particularly for oligonucleotide-based arrays. Furthermore, because aCGH can circumvent any tissue culture-based selection bias by testing DNA isolated directly from blood or tissue specimens, it is important to determine the lower limits of detection for the specific aCGH technology being used.
Given that oligonucleotide arrays can offer increased genome coverage compared with BAC arrays, we validated a previously reported whole-genome oligonucleotide array using clinical specimens representing a variety of constitutional cytogenetic abnormalities, including trisomies, monosomies, deletions, and duplications. Given the limited amount of literature on oligonucleotide aCGH and mosaicism detection, the sensitivity of this array to detect mosaicism was determined by testing artificially derived mosaic samples and additional naturally occurring mosaic specimens. Notably, combining dye-reversed replicates increased the sensitivity of oligonucleotide aCGH to detect mosaicism compared with individual uncombined aCGH experiments. In addition, discordant results were observed between aCGH performed on DNA isolated directly from an uncultured product of conception (POC) specimen and the mosaic karyotype obtained from the corresponding cultured specimen. Interestingly, further comparison of aCGH results from uncultured direct specimens to their cultured karyotypes in additional abnormal POC samples led to the detection of previously unidentified placental mosaicism in 5 of 10 (50%) specimens with double trisomies in culture.
Taken together, these results not only provide a comprehensive analysis of the sensitivity of an oligonucleotide array for the detection of mosaic abnormalities but also emphasize the need for careful interpretation when performing aCGH on uncultured direct specimens as the results may differ from those of the cultured cell karyotypes. These findings have important implications for clinical cytogenetic laboratories as molecular methods begin to replace the existing conventional cytogenetic techniques.19,20
MATERIALS AND METHODS
Specimens and DNA isolation
Peripheral blood, POC, and cultured chorionic villi with abnormal karyotypes by conventional cytogenetic analyses were acquired from the Mount Sinai Cytogenetics and Cytogenomics Laboratory and served as abnormal controls for aCGH validation. For POC specimens, chromosome analysis was performed on cultured cells, which would be predominantly mesenchymal in origin, from chorionic villi that were carefully dissected from maternal decidua. In addition, abnormal fibroblast and lymphoblastoid cell lines were acquired from the Coriell Institute for Medical Research (Camden, NJ), and their catalogue numbers are listed in the appropriate Tables. The karyotypes of all commercial cell lines were confirmed in our laboratory; however, a small number of cell lines (noted in Table, Supplemental Digital Content 1, http://links.lww.com/GIM/A92) were purchased as DNA, and their karyotypes were obtained from the Coriell Institute catalogue.
For aCGH analysis, genomic DNA was isolated from specimens using the Puregene® DNA Purification or DNeasy kits (both from Qiagen, Valencia, CA) according to the manufacturer's instructions. For POC specimens, DNA was extracted from direct chorionic villi to reduce the likelihood of maternal decidual overgrowth in long-term culture. Although the direct chorionic villi are composed of both mesenchymal and trophoblast cells, the extracted DNA from direct villi would primarily originate from the more predominant trophoblasts.
Cytogenetic and FISH analyses
Karyotypes were determined on stimulated peripheral blood cultures, cultured POC, cultured chorionic villi, and established cell lines by routine GTG chromosome banding using standard protocols. In addition, some specimens were analyzed by FISH using commercially available probes according to the manufacturer's instructions (Abbott Molecular, Des Plaines, IL).
Array comparative genomic hybridization
The custom 44K Agilent Technologies oligonucleotide array (Agilent Technologies, Santa Clara, CA) used for this study has been described previously.12 This array has enriched subtelomere coverage with an average theoretical resolution of 5 kb in the subtelomeres covering approximately 1 Mb of sequence and a 125 kb resolution throughout the remaining genome. All aCGH experiments were performed according to the manufacturer's instructions. In brief, 500 ng of experimental and gender-mismatched reference DNAs (Promega, Madison, WI) were digested with AluI and RsaI restriction endonucleases (Promega) and fluorescently labeled with Cyanine 5-dCTP (Cy-5; experimental) and Cyanine 3-dCTP (Cy-3; reference) using the Genomic DNA Labeling kit (Agilent Technologies). A dye-reversal experiment (Cy-3 for experimental and Cy-5 for reference) was performed in parallel for selected samples. Labeled experimental and reference DNAs were purified, combined, denatured, preannealed with Cot-1 DNA (Invitrogen, Carlsbad, CA) and blocking reagent (Agilent Technologies), and hybridized to arrays in a rotating oven (20 rpm) at 65°C for 24 hours. After hybridization and recommended washes, the arrays were scanned at 5 μm resolution with a G2505B Agilent Microarray Scanner. Images were processed with Feature Extraction 9.5.1 Software, and the data were analyzed with DNA Analytics 4.0 Software (both from Agilent Technologies). All array data passed the following quality control metrics: derivative log ratio spread ≤ 0.30; signal to noise (both colors) ≥ 20; signal intensity (both colors) ≥ 50; and background intensity (both colors) ≤ 10. Data from forward and dye-reversed experiments were analyzed both independently and after combining the paired data in DNA analytics using the inverse probe log ratios of the dye-reversed experiments.
All aCGH analyses were performed using both the “Fuzzy Zero” and “Centralization” default algorithms of DNA Analytics. Aberrations were identified using the Aberration Detection Method-1 (ADM1) algorithm12–13,21 with a sensitivity threshold of 6.0 and a data filter that rejected aberrations that did not include at least three consecutive probes with a log2 ratio ± 0.25. Aberrations were crossreferenced with the Database of Genomic Variants (http://projects.tcag.ca/variation/) to identify imbalances that were likely to represent benign copy number variants (CNVs). To minimize the detection of benign variants and given that the majority of benign CNVs are <100 kb in size,22,23 an aberration size threshold of 125 kb was used when the imbalance was located outside a clinically relevant region and/or when it lacked pathogenic gene content.
Oligonucleotide aCGH validation
Validation of the whole-genome custom oligonucleotide array was carried out on DNA isolated from specimens with nonmosaic abnormal karyotypes that were assessed by conventional cytogenetic or FISH techniques (Table, Supplemental Digital Content 1, http://links.lww.com/GIM/A92). The specimen types analyzed included peripheral blood, POC, chorionic villi, and fibroblast and lymphoblastoid cell lines, and the spectrum of copy-number imbalances detected by aCGH included chromosomal aneuploidy, deletions, duplications, unbalanced translocations, supernumerary marker chromosomes, and ring chromosomes. Complete concordance was observed between the karyotype/FISH and aCGH results. The abnormal aberration sizes ranged from 0.794 (unbalanced translocation) to 242.6 (trisomy) Mb. No additional pathogenic rearrangements were identified in any of the specimens tested by aCGH, and the molecular karyotypes are listed in Table, Supplemental Digital Content 1, http://links.lww.com/GIM/A92. An average of three CNVs per sample were detected, which ranged in size from 0.125 to 2.388 Mb. In addition, five DNA samples from phenotypically normal individuals (four females and one male) were tested by aCGH which detected an average of two CNVs per sample that ranged in size from 0.198 to 1.022 Mb (data not shown).
Quantitative polymerase chain reaction
Selected copy-number aberrations were additionally interrogated using quantitative polymerase chain reaction (qPCR) and the comparative Ct method for relative quantitation.24 Chromosome specific and reference primers used for the study are summarized in Table, Supplemental Digital Content 2, http://links.lww.com/GIM/A94. Reactions were performed in 384-well plates on an ABI 7900HT Fast Real-Time PCR system using SYBR Green I PCR Master Mix (Applied Biosystems, Foster City, CA), which includes a ROX internal reference. Each qPCR was performed in triplicate in 10 μL containing 10 ng of DNA, 1X SYBR Green PCR Master Mix, and forward and reverse primers at optimized concentrations (Table, Supplemental Digital Content 2, http://links.lww.com/GIM/A94). Amplification consisted of an initial enzyme activation step at 95°C for 10 minutes followed by 40 cycles of denaturation at 95°C for 15 seconds and combined annealing and extension at 60°C for 1 minute. The specificity of amplification for each qPCR product was confirmed by the identification of a single dissociation peak by melting curve analysis and single bands by agarose gel electrophoresis. To allow for comparison between test and reference primer sets, all reactions had to amplify with comparable efficiency as determined by a log10 dilution series. The relative gene copy number was determined by the 2−ΔΔCt equation where ΔΔCt equals (CtTEST − CtREFERENCE)sample- (CtTEST− CtREFERENCE)calibrator, and the calibrator sample used for all analyses was commercial male reference DNA (Promega).
RESULTS
Detection of aneuploid mosaicism by aCGH
To determine the sensitivity of the oligonucleotide array to identify mosaic aneuploidy, DNA samples with trisomy 21 and monosomy X ranging from 0 to 100% abnormal were evaluated by aCGH. These samples were constructed by combining appropriate quantities of abnormal DNA and gender-matched reference DNA before aCGH analysis. The results of these aCGH experiments are summarized in Table 1 and representative images of their chromosome profiles are shown in Figure 1. Both gains and losses of whole chromosome material, as low as 10%, were detected by the ADM1 algorithm; however, such low-level mosaicism was only detected using the combined data from dye-reversed replicates and in the absence of the data filter (see Materials and Methods section). Without combining dye-reversed replicates, mosaic abnormalities from individual uncombined aneuploid samples (both Cy-5 and Cy-3 labeled) were detected when present as low as 20–30% (Table 1).
Chromosome illustrations of mosaic aneuploid samples. A, Trisomy 21 samples with 0–100% mosaicism were tested by aCGH, and the copy-number gain was detected from combined replicates when present as low as 10%. The lower levels of mosaicism are reflected in reduced log2 values along the x-axis compared with 100% trisomy 21. Average log2 ratios are noted below each chromosome. B, Monosomy X samples with 0–100% mosaicism were tested by aCGH, and the copy-number loss was detectable from combined replicates when present as low as 10%. The lower levels of mosaicism are reflected in reduced negative log2 values along the x-axis compared with 100% monosomy X. Average log2 ratios are noted below each chromosome.
Detection of segmental aneuploid mosaicism by aCGH
To determine the sensitivity of the oligonucleotide array to identify mosaic segmental aneuploidy, DNA samples with segmental duplications and deletions ranging from 0 to 100% abnormal were evaluated by aCGH (Table 1). The duplication specimen [46,XX,dup(3)(q13q26)] was detected when present as low as 20% by the ADM1 algorithm using the combined data from dye-reversed replicates and in the absence of the data filter; however, the individual segmental aneuploid samples (both Cy-5 and Cy-3 labeled) were only detected as low as 40%. The duplicated region encompassed 56.1 Mb of sequence and 541 oligonucleotide aCGH probes. The deletion specimen [46,XY,del(15)(q11q13)] was detected when present as low as 30% by the ADM1 algorithm using the combined data from dye-reversed replicates and in the absence of the data filter; however, the individual segmental aneuploid samples labeled with Cy-5 and Cy-3 were only detected as low as 50% and 40%, respectively. The deleted region encompassed 5.0 Mb of sequence and 43 oligonucleotide aCGH probes.
Detection of aneuploid mosaicism in naturally occurring mosaic specimens
In addition, naturally occurring mosaic karyotypes ranging from 17 to 94% were tested by aCGH using dye-reversed replicates with unfiltered data analysis, and the results are summarized in Table 2. These specimens included four full aneuploidies and three partial aneuploidies. FISH was performed on selected cultured samples, and the percentage of aneuploid mosaicism in each was comparable with that of the cultured karyotype. Of note, a male POC specimen that displayed 40% mosaicism for trisomy 2 in cultured chorionic villi had an aCGH result that indicated a balanced male karyotype when performed on DNA isolated from the direct chorionic villi (Table 2). No evidence for maternal cell contamination was noted in the analysis of the cultured cells from this specimen or in its X and Y chromosome aCGH log2 ratios of the direct specimen. Given that the specimen was a POC, these results most likely represent type V mosaicism whereby the abnormality was absent or at very low levels in trophoblasts but present at detectable levels in the villus core and in the fetus, although no fetal tissue was available for confirmation.
Discordant aCGH results on specimens with double-aneuploid karyotypes
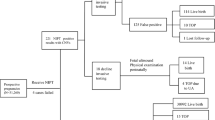
To investigate discrepancies between direct and cultured POC specimens, aCGH was performed on DNA from additional POC samples with abnormal karyotypes in culture. Interestingly, although more than 20 nonmosaic aneuploid POCs were confirmed by aCGH on direct specimens (Table, Supplemental Digital Content 1, http://links.lww.com/GIM/A92, and data not shown), additional discrepancies between direct and cultured specimens were observed among POCs with double-aneuploid karyotypes. The conventional and molecular karyotypes of the cultured and uncultured direct specimens, respectively, for 10 double-aneuploid POCs are summarized in Table 3. Notably, five specimens with double trisomy cultured karyotypes displayed only a single aneuploid abnormality when aCGH was performed on DNA isolated from the direct chorionic villi, and all had the more commonly observed trisomy (Figure, Supplemental Digital Content 3, http://links.lww.com/GIM/A93). These results suggested that placental mosaicism was present in half of the double aneuploidies. To confirm the aCGH findings, DNAs isolated directly from chorionic villi were analyzed by qPCR using primers specific for the relevant aneuploid chromosomes. Complete concordance between the aCGH and qPCR results was observed, and the qPCR copy-number quantitation is illustrated in Figure 2. As a negative control, a chromosomally balanced POC specimen by both aCGH and karyotype was analyzed by qPCR, and no change in copy number was observed at any of the tested loci (data not shown).
Quantitative PCR confirmation of POC specimens with discordant karyotype and aCGH results. Using the comparative Ct method of qPCR quantitation, samples with two and three total copies of tested autosomal loci have theoretical values of 1.0 and 1.5, respectively. Karyotypes derived from cultured cells are noted above qPCR quantitation graphs derived from corresponding direct DNA specimens. The cytoband of each primer set is listed on the x-axes, and error bars represent standard deviation. Gray and black bars represent chromosome regions that were disomic and trisomic, respectively, by aCGH.
DISCUSSION
The increasing use of aCGH in clinical cytogenetic laboratories requires careful validation of this technology in its ability to detect clinically important chromosomal abnormalities. Although oligonucleotide aCGH has greater resolution than traditional cytogenetic techniques, its sensitivity for detecting mosaicism has not been fully assessed. In addition to testing control samples representing the genomic regions covered on arrays, both the American College of Medical Genetics15 and the European best practice guidelines16 have recommended that clinical laboratories determine the sensitivity of their microarray for detecting mosaicism as part of their validation. Although few aCGH reports have documented mosaicism sensitivity, two recent studies using BAC-based platforms have shown that aCGH may be more likely to detect low-level mosaicism than traditional cytogenetic techniques and that mosaicism may be more common than previously appreciated.17,18 Herein, we extend these studies to oligonucleotide aCGH where we detected 10% mosaicism from trisomy 21- and monosomy X-derived samples, and 20–30% mosaicism from interstitial duplication- and deletion-derived samples. The trisomy 21 and monosomy X results were consistent with aneuploid mosaicism detected using BAC arrays (∼7%),17,18 suggesting that these two platforms may have similar sensitivity profiles for detecting whole chromosome aneuploidy. Moreover, BAC-based arrays are often run using dye-reversed replicates,7,18,25,26 which may account for the observed similarity in sensitivities between our study and those performed using BAC arrays.
Our detection of 10% aneuploid mosaicism by oligonucleotide aCGH was only observed using the unfiltered combined data from dye-reversed experiments, highlighting the importance of analyzing data with specific aberration criteria for mosaicism, and performing dye-reversed replicates when low-level mosaicism is suspected in clinical specimens. Although lowering the algorithm thresholds of individual experiments may also aid in the detection of low-level mosaicism, the increased sensitivity of this analysis typically results in an unacceptable decrease in specificity. In addition, although most software programs automatically detect statistically significant aberrations, careful examination of the data are warranted given that 10% mosaicism only slightly deviates from balanced log2 ratios. Of note, as unfiltered data results in false-positive copy-number aberrations, prospective specimens with a clinical indication for mosaicism, such as body asymmetry and/or pigmentary skin anomalies and clinical features strongly suggestive of a known aneuploidy syndrome, should be independently analyzed for aneuploid mosaicism using the described criteria (dye-reversed replicates and an unfiltered analysis) in addition to a routine filtered data analysis. However, given that clinical information is not always available and that dye-reversed replicates increase the cost of clinical aCGH testing, the decision to perform a dye-reversed combined analysis is up to the individual cytogenetics laboratory.
There was a tendency for the individual experimental aCGH samples labeled with Cy-3 to detect lower level mosaicism than the corresponding experimental sample labeled with Cy-5. This was presumably due to unique characteristics of the Cy-5 and Cy-3 fluorophores, which are known to have different incorporation and quantum efficiencies, and are detected by array scanners with different efficiencies.27–29 In addition, the target nucleic acid sequence is known to influence the direct incorporation of Cy-5 and Cy-3 labeled nucleotides,28,30 and Cy-5 is more sensitive to photobleaching than Cy-3.27 Together, these variables may have affected the ability of Cy-5 and Cy-3 labeled samples to individually detect lower level aberrations and thus provides support for the use of a dye-reversal aCGH strategy when mosaicism is suspected in clinical specimens.
By examining the results from both the individual and combined Cy-5 and Cy-3 labeled experiments, the ability to detect mosaicism was somewhat greater in the monosomy X and trisomy 21 specimens, followed by the duplication 3q and deletion 15q specimens. The size of their aberrations was 151.7, 36.8, 56.1, and 5.0 Mb, and the number of oligonucleotide probes per aberration was 1319, 519, 541, and 43, respectively. Although it is tempting to conclude that larger mosaic aberrations are more readily detectable than smaller ones, further testing is necessary to confirm this hypothesis.
An unexpected finding during our study was the discordant results between aCGH on uncultured POC chorionic villi and the karyotype from their cultured cells. Although this was initially observed in a mosaic specimen, the percentage of mosaicism (40%) was well within the previously determined detectable range of the oligonucleotide array. To further investigate this discrepancy, we analyzed a number of abnormal POCs and found that 5 of 10 specimens with double trisomies had only a single trisomy when the direct DNAs from uncultured chorionic villi were analyzed by both aCGH and qPCR. Of note, all discordant uncultured villi harbored the more commonly observed trisomy. Given that uncultured and cultured POC chorionic villi are primarily composed of cytotrophoblasts and mesenchymal cells, respectively, the discrepant findings likely represent previously unidentified placental mosaicism between the two cell populations. Although karyotype discordance between trophoblast and mesenchymal cells has been reported previously, to our knowledge, this is the first study to describe placental mosaicism between direct and cultured chorionic villi in double-trisomy POC specimens. The incidence of double trisomies in spontaneously aborted pregnancies has been reported to range from 0.21 to 2.8%.31,32 Because double trisomies tend to be aborted earlier in gestation than single trisomies and can often be associated with an empty sac, it is unclear whether our findings of mosaicism are confined to the placenta or reflect true fetal mosaicism.31 Although we hypothesize that a mosaic conceptus could progress further in a pregnancy before being spontaneously aborted than a nonmosaic double-trisomy conceptus, further studies and fetal tissue are necessary to support this conclusion.
A previous study of POC specimens comparing G-banded karyotypes and aCGH results identified additional submicroscopic abnormalities in ∼10% of POC specimens when analyzed with BAC-based arrays.33 Similar to our findings, this study also identified one aneuploid specimen with discordant aCGH and karyotype results. However, in contrast to our study, they performed aCGH on DNA isolated from the same cultured cells used for karyotyping, whereas our study performed aCGH on DNA isolated directly from the chorionic villi of the POC specimen. By analyzing cultured cells that are largely mesenchymal in origin, their analyses were presumably unable to detect additional mosaic POCs given that direct chorionic villi specimens are primarily composed of trophoblasts, which do not propagate well in culture.34 In addition, other studies have noted that because there is a high rate of culture failure and maternal decidua can selectively overgrow in long-term culture, examination of the uncultured villi from POC specimens may actually be preferable.33,35,36
Using conventional CGH, discordant results between trophoblast and chorionic stroma have previously been reported in placentas derived from infants that were viable and without obvious malformations.37 However, these cases represent confined placental mosaicism as no aneuploidy was observed in the corresponding amniotic membranes. In addition, conventional CGH identified confined placental mosaicism in placental tissue from complicated pregnancies, including those with intrauterine growth restriction.38 These reports and the placental mosaicism identified in our study underscore the need for careful interpretation when performing aCGH on uncultured specimens as the results may differ from the chromosome analysis of cultured cells. For example, discrepancies have been observed between aCGH of DNA from peripheral blood and the FISH and karyotype results from phytohemagglutinin-stimulated T-cells,18 suggesting that selection for or against specific abnormalities may occur during culture and that chromosomal abnormalities may be over- or underrepresented in certain cell types. Alternatively, although unlikely, it is noteworthy that some discrepancies between direct and cultured cell analyses may be the result of culture artifact, especially if an abnormality arises early in the establishment of the culture and represents a significant portion of the total cells.
In conclusion, we have determined the sensitivity of our validated whole-genome oligonucleotide array to detect mosaic abnormalities and have identified an increased ability to detect mosaicism by using unfiltered combined dye-reversed replicates. aCGH will likely become an invaluable tool for the detection of clinically relevant mosaic abnormalities as evidenced by the recent identification of postnatal mosaic tetrasomy 12p39 and trisomy 14,40 and prenatal mosaic trisomy 8q.41 In addition, our finding of previously unidentified placental mosaicism in POC specimens with double trisomies using oligonucleotide aCGH underscores the importance of careful interpretation when performing aCGH on DNA from uncultured direct specimens. Given that aCGH is currently unable to detect balanced translocations, inversions, and most polyploidy,42 traditional cultured chromosome analyses will likely continue to be used in conjunction with aCGH for diagnostic evaluation. Because this approach may intermittently interrogate distinct cellular populations depending on the type of specimen and chromosomal abnormality, it is important to recognize that discordant results between the two testing strategies may occur.
REFERENCES
Veltman JA, Schoenmakers EF, Eussen BH, et al. High-throughput analysis of subtelomeric chromosome rearrangements by use of array-based comparative genomic hybridization. Am J Hum Genet 2002; 70: 1269–1276.
Cheung SW, Shaw CA, Yu W, et al. Development and validation of a CGH microarray for clinical cytogenetic diagnosis. Genet Med 2005; 7: 422–432.
de Vries BB, Pfundt R, Leisink M, et al. Diagnostic genome profiling in mental retardation. Am J Hum Genet 2005; 77: 606–616.
Schoumans J, Ruivenkamp C, Holmberg E, Kyllerman M, Anderlid BM, Nordenskjold M . Detection of chromosomal imbalances in children with idiopathic mental retardation by array based comparative genomic hybridisation (array-CGH). J Med Genet 2005; 42: 699–705.
Bejjani BA, Shaffer LG . Application of array-based comparative genomic hybridization to clinical diagnostics. J Mol Diagn 2006; 8: 528–533.
Fan YS, Jayakar P, Zhu H, et al. Detection of pathogenic gene copy number variations in patients with mental retardation by genomewide oligonucleotide array comparative genomic hybridization. Hum Mutat 2007; 28: 1124–1132.
Lu X, Shaw CA, Patel A, et al. Clinical implementation of chromosomal microarray analysis: summary of 2513 postnatal cases. PLoS ONE 2007; 2: e327.
Pickering DL, Eudy JD, Olney AH, et al. Array-based comparative genomic hybridization analysis of 1176 consecutive clinical genetics investigations. Genet Med 2008; 10: 262–266.
Ballif BC, Sulpizio SG, Lloyd RM, et al. The clinical utility of enhanced subtelomeric coverage in array CGH. Am J Med Genet A 2007; 143A: 1850–1857.
Shen Y, Irons M, Miller DT, et al. Development of a focused oligonucleotide-array comparative genomic hybridization chip for clinical diagnosis of genomic imbalance. Clin Chem 2007; 53: 2051–2059.
Thorland EC, Gonzales PR, Gliem TJ, Wiktor AE, Ketterling RP . Comprehensive validation of array comparative genomic hybridization platforms: how much is enough?. Genet Med 2007; 9: 632–641.
Toruner GA, Streck DL, Schwalb MN, Dermody JJ . An oligonucleotide based array-CGH system for detection of genome wide copy number changes including subtelomeric regions for genetic evaluation of mental retardation. Am J Med Genet A 2007; 143A: 824–829.
Baldwin EL, Lee JY, Blake DM, et al. Enhanced detection of clinically relevant genomic imbalances using a targeted plus whole genome oligonucleotide microarray. Genet Med 2008; 10: 415–429.
Zhang ZF, Ruivenkamp C, Staaf J, et al. Detection of submicroscopic constitutional chromosome aberrations in clinical diagnostics: a validation of the practical performance of different array platforms. Eur J Hum Genet 2008; 16: 786–792.
Shaffer LG, Beaudet AL, Brothman AR, et al. Microarray analysis for constitutional cytogenetic abnormalities. Genet Med 2007; 9: 654–662.
Vermeesch JR, Fiegler H, de Leeuw N, et al. Guidelines for molecular karyotyping in constitutional genetic diagnosis. Eur J Hum Genet 2007; 15: 1105–1114.
Ballif BC, Rorem EA, Sundin K, et al. Detection of low-level mosaicism by array CGH in routine diagnostic specimens. Am J Med Genet A 2006; 140: 2757–2767.
Cheung SW, Shaw CA, Scott DA, et al. Microarray-based CGH detects chromosomal mosaicism not revealed by conventional cytogenetics. Am J Med Genet A 2007; 143A: 1679–1686.
Edelmann L, Hirschhorn K . Clinical utility of array CGH for the detection of chromosomal imbalances associated with mental retardation and multiple congenital anomalies. Ann N Y Acad Sci 2009; 1151: 157–166.
Gijsbers AC, Lew JY, Bosch CA, et al. A new diagnostic workflow for patients with mental retardation and/or multiple congenital abnormalities: test arrays first. Eur J Hum Genet 2009; 17: 1394–1402.
Hester SD, Reid L, Nowak N, et al. Comparison of comparative genomic hybridization technologies across microarray platforms. J Biomol Tech 2009; 20: 135–151.
Iafrate AJ, Feuk L, Rivera MN, et al. Detection of large-scale variation in the human genome. Nat Genet 2004; 36: 949–951.
Perry GH, Ben-Dor A, Tsalenko A, et al. The fine-scale and complex architecture of human copy-number variation. Am J Hum Genet 2008; 82: 685–695.
Schmittgen TD, Livak KJ . Analyzing real-time PCR data by the comparative C(T) method. Nat Protoc 2008; 3: 1101–1108.
Van den Veyver IB, Patel A, Shaw CA, et al. Clinical use of array comparative genomic hybridization (aCGH) for prenatal diagnosis in 300 cases. Prenat Diagn 2009; 29: 29–39.
Sahoo T, Cheung SW, Ward P, et al. Prenatal diagnosis of chromosomal abnormalities using array-based comparative genomic hybridization. Genet Med 2006; 8: 719–727.
van Hal NL, Vorst O, van Houwelingen AM, et al. The application of DNA microarrays in gene expression analysis. J Biotechnol 2000; 78: 271–280.
Tseng GC, Oh MK, Rohlin L, Liao JC, Wong WH . Issues in cDNA microarray analysis: quality filtering, channel normalization, models of variations and assessment of gene effects. Nucleic Acids Res 2001; 29: 2549–2557.
Bilban M, Buehler LK, Head S, Desoye G, Quaranta V . Normalizing DNA microarray data. Curr Issues Mol Biol 2002; 4: 57–64.
Taniguchi M, Miura K, Iwao H, Yamanaka S . Quantitative assessment of DNA microarrays—comparison with Northern blot analyses. Genomics 2001; 71: 34–39.
Reddy KS . Double trisomy in spontaneous abortions. Hum Genet 1997; 101: 339–345.
Menasha J, Levy B, Hirschhorn K, Kardon NB . Incidence and spectrum of chromosome abnormalities in spontaneous abortions: new insights from a 12-year study. Genet 2005 Med; 7: 251–263.
Schaeffer AJ, Chung J, Heretis K, Wong A, Ledbetter DH, Lese Martin C . Comparative genomic hybridization-array analysis enhances the detection of aneuploidies and submicroscopic imbalances in spontaneous miscarriages. Am J Hum Genet 2004; 74: 1168–1174.
Niazi M, Coleman DV, Loeffler FE . Trophoblast sampling in early pregnancy. Culture of rapidly dividing cells from immature placental villi. Br J Obstet Gynaecol 1981; 88: 1081–1085.
Bell KA, Van Deerlin PG, Haddad BR, Feinberg RF . Cytogenetic diagnosis of “normal 46,XX” karyotypes in spontaneous abortions frequently may be misleading. Fertil Steril 1999; 71: 334–341.
Lomax B, Tang S, Separovic E, et al. Comparative genomic hybridization in combination with flow cytometry improves results of cytogenetic analysis of spontaneous abortions. Am J Hum Genet 2000; 66: 1516–1521.
Lestou VS, Desilets V, Lomax BL, et al. Comparative genomic hybridization: a new approach to screening for intrauterine complete or mosaic aneuploidy. Am J Med Genet 2000; 92: 281–284.
Amiel A, Bouaron N, Kidron D, Sharony R, Gaber E, Fejgin MD . CGH in the detection of confined placental mosaicism (CPM) in placentas of abnormal pregnancies. Prenat Diagn 2002; 22: 752–758.
Theisen A, Rosenfeld JA, Farrell SA, et al. aCGH detects partial tetrasomy of 12p in blood from Pallister-Killian syndrome cases without invasive skin biopsy. Am J Med Genet A 2009; 149A: 914–918.
Shinawi M, Shao L, Jeng LJ, et al. Low-level mosaicism of trisomy 14: phenotypic and molecular characterization. Am J Med Genet A 2008; 146A: 1395–1405.
Wood E, Dowey S, Saul D, et al. Prenatal diagnosis of mosaic trisomy 8q studied by ultrasound, cytogenetics, and array-CGH. Am J Med Genet A 2008; 146A: 764–769.
Ballif BC, Kashork CD, Saleki R, et al. Detecting sex chromosome anomalies and common triploidies in products of conception by array-based comparative genomic hybridization. Prenat Diagn 2006; 26: 333–339.
Author information
Authors and Affiliations
Corresponding author
Additional information
S.A.S. and T.B. were the recipients of Biochemical/Molecular Genetics Fellowships from the Genzyme Corporation.
Disclosure: The authors declare no conflict of interest.Supplemental digital content is available for this article. Direct URL citations appear in the printed text and are provided in the HTML and PDF versions of this article on the journal's Web site (www.geneticsinmedicine.org).
Rights and permissions
About this article
Cite this article
Scott, S., Cohen, N., Brandt, T. et al. Detection of low-level mosaicism and placental mosaicism by oligonucleotide array comparative genomic hybridization. Genet Med 12, 85–92 (2010). https://doi.org/10.1097/GIM.0b013e3181cc75d0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1097/GIM.0b013e3181cc75d0
Keywords
This article is cited by
-
Phenotypic findings and pregnancy outcomes of fetal rare autosomal aneuploidies detected using chromosomal microarray analysis
Human Genomics (2022)
-
Cytogenomic identification and long-read single molecule real-time (SMRT) sequencing of a Bardet–Biedl Syndrome 9 (BBS9) deletion
npj Genomic Medicine (2018)
-
Analyses of karyotype by G-banding and high-resolution microarrays in a gender dysphoria population
Genes & Genomics (2018)
-
Setleis syndrome due to inheritance of the 1p36.22p36.21 duplication: evidence for lack of penetrance
Journal of Human Genetics (2015)
-
The accuracy of chromosomal microarray testing for identification of embryonic mosaicism in human blastocysts
Molecular Cytogenetics (2014)