Abstract
Purpose: Pitt-Hopkins syndrome is characterized by severe mental retardation, characteristic dysmorphic features, and susceptibility to childhood-onset seizures and intermittent episodes of hyperventilation. This syndrome is caused by haploinsufficiency of TCF4, which encodes a basic helix-loop-helix transcription factor. Missense, nonsense, splice-site mutations, and gene deletions have been found in individuals with Pitt-Hopkins syndrome. Previous reports have suggested that the Pitt-Hopkins syndrome phenotype is independent of mutation or deletion type.
Methods: We screened 13,186 individuals with microarray-based comparative genomic hybridization. We also conducted a review of the literature and statistical analysis of the phenotypic features for all individuals with confirmed mutations or deletions of TCF4.
Results: We identified seven individuals with TCF4 deletions. All patients have features consistent with Pitt-Hopkins syndrome, although only three have breathing anomalies, and none has seizures. Our review of previously reported cases with TCF4 mutations and deletions showed that all patients with Pitt-Hopkins syndrome reported to date have severe psychomotor retardation, the onsets of seizures and hyperventilation episodes are limited to the first decade in most reported patients with Pitt-Hopkins syndrome, hyperventilation episodes are more common than seizures and are seen in the oldest patients, and individuals with missense TCF4 mutations are more likely to develop seizures.
Conclusions: On the basis of an analysis of published cases, we propose a genotype–phenotype correlation of increased seizure activity with missense TCF4 mutations.
Similar content being viewed by others
Main
Pitt-Hopkins syndrome (PTHS, OMIM 610954) was first described in 19781 in two individuals and is characterized by specific dysmorphic features (deep-set eyes, broad-beaked nose, wide mouth with bow-shaped upper lip, and widely spaced teeth), childhood-onset hyperventilation episodes, childhood-onset seizures, microcephaly, and severe psychomotor retardation with a happy disposition. It is an autosomal dominant condition caused by haploinsufficiency for TCF4 (transcription factor 4, OMIM 602272) on 18q21.2.2–4 The etiology of PTHS was determined when two groups concurrently screened individuals with PTHS for chromosomal deletions using microarray technology, one with array-based comparative genomic hybridization (aCGH)5 and the other with single-nucleotide polymorphism-based molecular karyotyping.3 Each group described a patient with a de novo deletion involving this gene. Further screening of individuals with PTHS or phenotypic overlap with PTHS found individuals with de novo missense and nonsense mutations in TCF4.3,5 Subsequently, more individuals have been published with disruptions, deletions, and mutations in TCF4, most with typical PTHS features.4,6–10
TCF4 is a transcription factor that belongs to the class I basic helix-loop-helix (bHLH) protein family, also known as the “E-protein” family. These proteins form homo- and heterodimers with other helix-loop-helix (HLH) proteins through their HLH domains and bind DNA through their basic domains.11,12 Missense, nonsense, frameshift, and splice-site mutations and larger deletions encompassing all or part of TCF4 have been found in patients with PTHS.3–5,7–10 Whole-gene and large intragenic deletions represent 15% of molecularly confirmed TCF4 alterations in the literature, with the smallest deletion described to date a 0.5-Mb deletion that results in a deletion of the 5′ half of TCF4.7 All but three of the missense mutations so far described have been in the bHLH region of TCF4.3–5,8 To date, the only genotype–phenotype correlation observed has been limited flexion and an absent flexion crease on the thumb in some patients with deletions.
We report seven individuals with TCF4 microdeletions detected by aCGH. We also show a genotype–phenotype analysis of these and all other TCF4 mutations in the literature, which revealed an increased incidence of seizures among individuals with missense mutations.
MATERIALS AND METHODS
Patient ascertainment
Between November 2007 and February 2009, we screened 13,186 samples with aCGH. For the individuals described here, informed consent was obtained to publish clinical descriptions and photographs.
Bacterial artificial chromosome microarray analysis
aCGH was initially performed on Patients 1, 3, and 5 with a bacterial artificial chromosome (BAC) microarray, the SignatureChip Whole Genome® (Signature Genomic Laboratories, Spokane, WA).13 Results were visualized using our laboratory-developed computer software Genoglyphix® (http://www.signaturegenomics.com/genoglyphix.html). Each BAC clone was verified by fluorescence in situ hybridization (FISH) for its chromosomal location before microarray construction and validated for use in FISH to visualize chromosome abnormalities identified by the microarray. Microarray analysis was performed as previously described.13
Oligonucleotide aCGH
Oligonucleotide-based microarray analysis was performed on all patients using a 105K-feature whole-genome microarray, the SignatureChip Oligo Solution®, made for Signature Genomic Laboratories by Agilent Technologies (Santa Clara, CA) with one probe every 10 kb in targeted regions—microdeletion/microduplication syndromes, pericentromeric regions, subtelomeres, and genes in important developmental pathways—and an average probe spacing of one probe every 35 kb throughout the rest of the genome. Genomic DNA was extracted from the peripheral blood using a Qiagen M48 Biorobot automated DNA extraction system. Purified genomic DNA was then sonicated and labeled with Alexa Fluor dyes 555 or 647 using a BioPrime Total DNA labeling kit (Invitrogen Corp, Carlsbad, CA). Array hybridization and washing were performed as specified by the manufacturer (Agilent Technologies). Arrays were scanned using an Axon 4000B scanner (Molecular Devices, Sunnyvale, CA) and analyzed using Agilent Feature Extraction software v9.5.1 and Agilent CGH Analytics software v3.5.14. Results were visualized using Genoglyphix.
Genome database analysis
Computational analysis of 18q21.2 was performed using the annotated March 2006 assembly of the human genome on the UCSC genome browser (http://genome.ucsc.edu; build hg18). The locations of TCF4 introns and exons were determined using the consensus CDS database available from NCBI (http://www.ncbi.nlm.nih.gov/CCDS/CcdsBrowse.cgi).
Fluorescence in situ hybridization
Metaphase FISH analysis with the BAC probe RP11–353I8 or RP11–746K23 was used to confirm deletions of the TCF4 locus in Patients 1 to 3.14
Literature review
Searches of the PubMed database (http://www.ncbi.nlm.nih.gov/pubmed/) using the terms “Pitt Hopkins,” “TCF4,” “18q deletion,” and “18q monosomy” were performed to find all the reports of patients with PTHS and cytogenetically defined deletions encompassing 18q21.2.
Statistical analyses
Fisher exact test calculations were performed using a publicly available program (http://www.langsrud.com/fisher.htm). Analysis was performed for the frequency of each phenotypic feature listed in Table 1 for predicted haploinsufficient mutations in comparison with missense mutations. The correction for multiple comparisons was made using Bonferroni adjustment (http://www.quantitativeskills.com/sisa/calculations/bonfer.htm).
RESULTS
Case reports
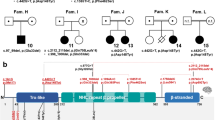
Of the 13,186 samples screened during the study period, we identified seven individuals with microdeletions of 18q21.2. We characterized the deletions using a high-density 105K oligonucleotide microarray. In these individuals, microarray analysis identified single-copy losses of the region containing TCF4 that ranged in size from 6.1 Mb to 63 kb (Fig. 1). Figure 2 shows the location of the intragenic gene deletions. Gene deletion in Patient 5 is predicted to cause a frameshift, whereas precise breakpoints are unknown in Patients 4, 6, and 7. It is possible that Patients 6 and 7 have in-frame deletions that leave the bHLH domain intact, whereas gene deletion in Patient 4 likely abolishes this domain.
Analysis of individuals with deletions encompassing TCF4 on 18q21.2. Oligonucleotide microarray profiles for (A) a single-copy loss of 6.1 Mb (chr18:45,564,835–51,677,777) in Patient 1, (B) a single-copy loss of 2.4 Mb (chr18:49,229,755–51,677,916) in Patient 2, (C) a single-copy loss of 2.9 Mb (chr18:49,884,520–52,773,507) in Patient 3, (D) a single-copy loss of 63 kb (chr18:51,050,140–51,113,108) in Patient 4, (E) a single-copy loss of 101 kb (chr18:51,136,052–51,237,535) in Patient 5, (F) a single-copy loss of 76 kb (chr18:51,059,650–51,136,312) in Patient 6, and (G) a single-copy loss of 64 kb (chr18:51,078,120–51,142,195) in Patient 7. For the microarray plots, probes are ordered on the x-axis according to physical mapping positions with distal 18p to the left and distal 18q to the right. Blue bars at the bottom of the plots represent the deletion sizes. (H) Summary of the deletion sizes in individuals with microdeletions encompassing TCF4. Green bars indicate the approximate deletion sizes in Patients 1 to 7. Black notches at the top of the diagram indicate distances, in Mb, from the chromosome 18p telomere. The TCF4 gene is indicated by a blue box.
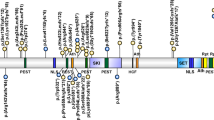
Diagram of TCF4 showing all reported mutations. Intron sizes are not to scale. White exons (1 and 20) are noncoding. The lightly shaded Exon 18 contains the basic helix-loop-helix (bHLH) domain. M, missense; trans, site of balanced translocation; ss, splice site; fs, frameshift creating a longer coding sequence; X, nonsense or frameshift mutation leading to premature protein truncation. Nine of the missense mutations in Exon 18 have been within the basic portion of the bHLH domain, with six in the codon 576/580, two in the codon 574/578, and one in the codon 572/576. One reported nonsense mutation in this exon was at the beginning of the bHLH domain, in the codon 563/567. The other missense mutation in this exon is downstream of the bHLH domain, in the codon 610/614, which codes for an amino acid that is highly conserved across species.3–6,8,10 Also shown are intragenic deletions of cases presented here. Gene deletion in Patient 5 extends from Intron 5 to Intron 8. Gene deletion in Patient 4 extends from Intron 8 to at least Intron 17, and it may extend into Exon 20. Gene deletion in Patient 6 extends from Intron 8 to at least Intron 15, and it may extend into Intron 17. Gene deletion in Patient 7 extends from Intron 8 to at least Intron 13, and it may extend into Intron 15.
Large deletions in Patients 1 to 3 were confirmed by FISH, whereas deletions in Patients 4 to 7 were below the resolution for FISH. Parental testing confirmed a de novo origin in Patients 2, 3, and 7. All other parental samples were unavailable, except the mother of Patient 5, who did not have the deletion found in her daughter.
Clinical information was available on Patients 1 to 7 and is summarized in Table 2. Images of some of the physical characteristics seen in these patients are shown in Figure 3.
Facial features of Patient 1 (A, B), Patient 2 (C, D), Patient 3 (E), and Patient 6 (F). Note the wide mouth with a protruding philtrum and bow-shaped upper lip, widely spaced teeth, deeply set eyes, thin eyebrows, broad and beaked nose with a high bridge, and thick, dysplastic ears. Strabismus is present in Patients 2 and 6 (C, F). Blue sclerae seen in Patient 3 (E) is due to osteogenesis imperfecta. Single palmar crease (G), mild 2 to 3 toe syndactyly, and toe clinodactyly (H) were present in Patient 1.
Review of cases with TCF4 mutations or deletions in the literature
A total of 52 cases with confirmed mutations or deletions of TCF4 have been reported.2–10 A summary of the clinical features of these individuals, categorized by the type of mutation or deletion, is listed in Table 1. Figure 2 shows the locations of all reported mutations in TCF4. In addition, 12 cases of terminal or interstitial deletions of 18q specifically involving 18q21.2 have been described, although the inclusion of TCF4 in these deletions has not been confirmed.15–20 Therefore, these cytogenetically described deletions were excluded from further comparison.
Statistical analyses
Individuals reported in the literature with missense mutations were compared with individuals reported in the literature with mutations predicted to lead to haploinsufficiency. Functional studies are necessary to determine the exact molecular effects of each of these mutations; however, the truncating mutations, including splice-site mutations, nonsense mutations, and intragenic deletions, were grouped within the category of “haploinsufficiency” for phenotypic analysis. There is no significant difference in the ages of these two groups (unpaired t test, P = 0.85). The Fisher exact two-tailed test was used to test for significant differences in the incidence of any phenotypic features (Table 1). A significant difference was observed in the incidence of seizures in patients with missense mutations (P = 0.0003). This remains significant after a correction for multiple comparisons; the Bonferroni adjustment for 18 tests (the number of categories assessed in this analysis) instructs to lower α to 0.0028 to maintain a 95% confidence level. No other significant differences were found.
DISCUSSION
We report seven individuals with microdeletions of 18q21.2 encompassing or within the TCF4 gene, haploinsufficiency of which causes PTHS. During the study period, our laboratory also detected 38 Williams syndrome deletions and 16 Smith-Magenis syndrome deletions. Using a comparison to the population frequency of 1 of 7,500 for Williams syndrome21 and 1 of 15,000 for Smith-Magenis syndrome,22 our calculation for an estimated frequency of PTHS deletions is between 1 of 34,000 and 1 of 41,000.
Most cases in the literature with TCF4 mutations or deletions have the typical PTHS neurologic and dysmorphic features. Our patients reported here all demonstrate the characteristic PTHS facial features and, with the exception of the youngest patient, severe psychomotor delay. Three experience breathing abnormalities, and none has a history of seizures. This is similar to what has been reported in the literature. The phenotypic incidences determined after our review are approximately 56% for hyperventilation and 35% for seizures and are likely underestimates for lifetime risk because some of the patients in the analysis are younger than the average age of onset for these features. Most cases in the literature who presented with seizures or hyperventilation episodes had an onset within the first decade of life. However, there has been a case with epilepsy onset at 18 years10 and another with breathing abnormality onset at 17 years,4 indicating it is possible for teenagers to develop these symptoms. The hyperventilation episodes, unlike seizures, are seen in the oldest patients, although there is an 18-year-old patient reported without breathing abnormalities.4 Despite the younger average age of onset for epilepsy than for breathing abnormalities, one series suggested that hyperventilation episodes were more common in epileptic patients, and hyperventilation preceded epilepsy.10
With aCGH detecting chromosome abnormalities in patients with nonspecific symptoms, such as developmental delay, earlier diagnoses may be reached for syndromes such as PTHS. This allows for better patient care, medical management, and appropriate surveillance for the development or presence of other syndromic features.
Haploinsufficiency of TCF4
Animal studies support a role for TCF4 in neurologic development. Class I bHLH genes have been shown to be widely expressed in the developing vertebrate embryo, including the central and peripheral nervous systems.23 TCF4 is highly expressed throughout the developing human central nervous system.10 Studies of embryologic expression of tcf4 in zebrafish showed expression in the telencephalon early, increasing with time in the hindbrain, diencephalon, midbrain, branchial arches, and retina.7 Although Tcf4± mice do not have any obvious phenotypic abnormalities,24 and Tcf4−/− mice do not show any gross structural brain defects, murine brains completely lacking Tcf4 have a substantial reduction in the neurons of the pontine nucleus. This defect is attributed to abnormal migration because of the absence of the specific activity of the heterodimer formed by Tcf4 and Math1, another bHLH transcription factor essential in neuronal differentiation. Thus, humans with TCF4 haploinsufficiency could be missing subsets of neurons because of impaired differentiation.25
Decreased interaction of TCF4 with ASCL1, another bHLH transcription factor that plays a role in the central nervous system development and differentiation of neural crest cells,3,5 has been proposed to cause the breathing abnormalities in PTHS. ASCL1 is part of a developmental pathway that also includes PHOX2A, PHOX2B, and RET, which controls the development of noradrenergic neurons important in the autonomic nervous system.26,27 Animal and in vitro models lacking Ascl1 have abnormal respiratory networks, and mutations in ASCL1 have also been found in individuals with congenital central hypoventilation syndrome (OMIM 209880).27,28 TCF4 functionally interacts with ASCL1,29 and if TCF4 plays an important role in this PHOX pathway, then haploinsufficiency may lead to the breathing abnormalities because of impaired noradrenergic neuronal development. Although Tcf4−/− mice have a normal locus ceruleus (a noradrenergic center),25 mice may not be a good model of this condition because heterozygote mice have a different phenotype than humans with PTHS. Thus, impaired interaction of TCF4 with ASCL1 may still have an etiological role in the PTHS phenotype.
Other genetic factors may impact the phenotype in individuals with whole-gene TCF4 deletions. In particular, the phenotype of individuals with deletions may be impacted by haploinsufficiency of neighboring genes, and this may be of clinical significance. For example, the deletion of gene in Patient 1 includes SMAD4, and haploinsufficiency of this gene can cause juvenile polyposis syndrome and hereditary hemorrhagic telangiectasia.30–33 Furthermore, juvenile polyposis syndrome can cause constipation, which overlaps with the PTHS phenotype. In addition, absent flexion creases on the thumbs have mostly been reported in deletion patients, although this has also been seen in a patient with a nonsense mutation.4,10 However, analysis of the phenotype in reported individuals with deletions shows no significant differences from those with other types of TCF4 mutations. Thus, the phenotype in the patients with larger deletions may be mostly explained by TCF4 haploinsufficiency.
Mutations in TCF4
Previous reports of individuals with TCF4 mutations and deletions suggested the phenotype was independent of the mutation type, which supported a model of haploinsufficiency as the cause of PTHS.4,5,8 A notable exception to the typical PTHS phenotype seen in association with a TCF4 mutation is a female with mild to moderate mental retardation who had a disruption of TCF4 caused by a balanced translocation.6 This individual was shown to produce a fusion transcript of the noncoding Exon 1 of CHD6 with the 5′ end of TCF4, starting at Exon 4.6 Residual function of a TCF4 protein truncated at the amino terminus may have prevented the emergence of a PTHS phenotype in this patient. Our analysis of a larger number of patients in this article confirms that, for most PTHS features, phenotype is independent of mutation type, with the exception of seizures, which are significantly more prevalent in individuals with missense mutations. In addition, there is a nonstatistically significant trend toward increased breathing and brain abnormalities in this group. Supporting this observation is a report of somatic mosaicism for a TCF4 missense mutation in a mother of a patient with PTHS who developed seizures at the age of 20 years.10 Studies by Zweier et al.3 and de Pontual et al.10 in a human cell line have shown lower, although not significant, TCF4 transcription activity with transfection of genes containing a missense mutation in the bHLH domain compared with those with nonsense mutations. Cells lacking transfected TCF4 also had significantly more transcription activity than cells with any type of mutant TCF4 transfected.10 In addition, three of the four patients ascertained by Amiel et al.5 using strict phenotypic criteria had missense mutations, a higher proportion compared with subsequent studies that screened more individuals for TCF4 mutations following less-strict phenotypic criteria. Therefore, a missense mutation may be more likely to result in a clinically recognizable PTHS diagnosis. Although the only significant difference in individuals with missense mutations in our literature review is the increased prevalence of seizures, this population is also enriched for hyperventilation episodes and brain abnormalities. There may be an ascertainment bias in the earlier published PTHS cohorts because clinical suspicion of PTHS needed to exist for inclusion in the study, whereas our cohort only required a suspicion of a chromosome abnormality for inclusion. Therefore, there may be more mildly affected individuals with TCF4 point mutations who have not yet been identified. However, these phenotypic differences may exist because of a dominant-negative activity of mutant TCF4 with an amino acid substitution.
Nine of the 12 patients reported with missense mutations, and seven of the eight patients with missense mutations and sufficient phenotypic information to be used in the phenotypic analysis in this article, have an amino acid change in the highly conserved basic portion of the bHLH domain (Fig. 3), which is predicted to affect the ability of the protein to bind DNA while leaving the HLH, or the protein–protein interaction domain, intact.3–5,8 Therefore, these missense mutations could have a dominant negative effect, sequestering other HLH partners of TCF4 but not correctly binding to DNA,5 an effect seen with similar engineered mutations in other bHLH genes.34,35 Thus, missense mutations in the bHLH domain may have a greater impact on TCF4 and other TCF4 partners' functions compared with haploinsufficiency caused by deletion or other nonsense mutations. Similar dominant-negative activities caused by missense mutations in DNA-binding domains of transcription factors have been shown to result in disease. Such mutations have been described in PITX1, associated with autosomal dominant club foot,36 in STAT3, associated with hyper-IgE syndrome,37 in AIRE, associated with autosomal dominant autoimmune thyroiditis,38 and in somatic mutations of IRF5 in chronic lymphocytic leukemia and adult T-cell leukemia/lymphoma.39 There is also precedent for different mutations within the same gene causing variable disease phenotypes; in connective tissue disorders, such as osteogenesis imperfecta, missense mutations are associated with a more severe phenotype than nonsense mutations,40 although the effect seen here is not as substantive. Similarly, in Townes-Brocks syndrome, truncations of the SALL1 transcription factor gene are thought to have a dominant-negative effect, preventing the normal nuclear import of the wild-type protein,41 whereas deletions of the gene show a milder phenotype.42,43 Only missense mutations in the bHLH domain of TCF4 would be expected to have a dominant-negative effect by this mechanism. The effect of missense mutations in other regions is unclear, and further studies of all TCF4 mutations and patients who have them are required to understand better how this affects the PTHS phenotype.
References
Pitt D, Hopkins I . A syndrome of mental retardation, wide mouth and intermittent overbreathing. Aust Paediatr J 1978; 14: 182–184.
Peippo MM, Simola KO, Valanne LK, et al. Pitt-Hopkins syndrome in two patients and further definition of the phenotype. Clin Dysmorphol 2006; 15: 47–54.
Zweier C, Peippo MM, Hoyer J, et al. Haploinsufficiency of TCF4 causes syndromal mental retardation with intermittent hyperventilation (Pitt-Hopkins syndrome). Am J Hum Genet 2007; 80: 994–1001.
Zweier C, Sticht H, Bijlsma EK, et al. Further delineation of Pitt-Hopkins syndrome: phenotypic and genotypic description of 16 novel patients. J Med Genet 2008; 45: 738–744.
Amiel J, Rio M, de Pontual L, et al. Mutations in TCF4, encoding a class I basic helix-loop-helix transcription factor, are responsible for Pitt-Hopkins syndrome, a severe epileptic encephalopathy associated with autonomic dysfunction. Am J Hum Genet 2007; 80: 988–993.
Kalscheuer VM, Feenstra I, Van Ravenswaaij-Arts CM, et al. Disruption of the TCF4 gene in a girl with mental retardation but without the classical Pitt-Hopkins syndrome. Am J Med Genet A 2008; 146: 2053–2059.
Brockschmidt A, Todt U, Ryu S, et al. Severe mental retardation with breathing abnormalities (Pitt-Hopkins syndrome) is caused by haploinsufficiency of the neuronal bHLH transcription factor TCF4. Hum Mol Genet 2007; 16: 1488–1494.
Giurgea I, Missirian C, Cacciagli P, et al. TCF4 deletions in Pitt-Hopkins syndrome. Hum Mutat 2008; 29: E242–E251.
Andrieux J, Lepretre F, Cuisset JM, et al. Deletion 18q21.2q21.32 involving TCF4 in a boy diagnosed by CGH-array. Eur J Med Genet 2008; 51: 172–177.
de Pontual L, Mathieu Y, Golzio C, et al. Mutational, functional, and expression studies of the TCF4 gene in Pitt-Hopkins syndrome. Hum Mutat 2009; 30: 669–676.
Bergqvist I, Eriksson M, Saarikettu J, et al. The basic helix-loop-helix transcription factor E2–2 is involved in T lymphocyte development. Eur J Immunol 2000; 30: 2857–2863.
Massari ME, Murre C . Helix-loop-helix proteins: regulators of transcription in eukaryotic organisms. Mol Cell Biol 2000; 20: 429–440.
Ballif BC, Theisen A, Coppinger J, et al. Expanding the clinical phenotype of the 3q29 microdeletion syndrome and characterization of the reciprocal microduplication. Mol Cytogenet 2008; 1: 8.
Shaffer LG, McCaskill C, Han JY, et al. Molecular characterization of de novo secondary trisomy 13. Am J Hum Genet 1994; 55: 968–974.
Gustavsson P, Kimber E, Wahlstrom J, Anneren G . Monosomy 18q syndrome and atypical Rett syndrome in a girl with an interstitial deletion (18)(q21.1q22.3). Am J Med Genet 1999; 82: 348–351.
Engelen JJ, Moog U, Weber J, Haagen AA, van Uum CM, Hamers AJ . Deletion of chromosome region 18q21.1 –> 18q21.3 in a patient without clinical features of the 18q- phenotype. Am J Med Genet A 2003; 119: 356–359.
Feenstra I, Vissers LE, Orsel M, et al. Genotype-phenotype mapping of chromosome 18q deletions by high-resolution array CGH: an update of the phenotypic map. Am J Med Genet A 2007; 143: 1858–1867.
Linnankivi T, Tienari P, Somer M, et al. 18q deletions: clinical, molecular, and brain MRI findings of 14 individuals. Am J Med Genet A 2006; 140: 331–339.
Wilson MG, Towner JW, Forsman I, Siris E . Syndromes associated with deletion of the long arm of chromosome 18[del(18q)]. Am J Med Genet 1979; 3: 155–174.
Petty RE, Malleson P, Kalousek DK . Chronic arthritis in two children with partial deletion of chromosome 18. J Rheumatol 1987; 14: 586–587.
Stromme P, Bjornstad PG, Ramstad K . Prevalence estimation of Williams syndrome. J Child Neurol 2002; 17: 269–271.
Elsea SH, Girirajan S . Smith-Magenis syndrome. Eur J Hum Genet 2008; 16: 412–421.
Ik Tsen Heng J, Tan SS . The role of class I HLH genes in neural development—have they been overlooked?. Bioessays 2003; 25: 709–716.
Zhuang Y, Cheng P, Weintraub H . B-lymphocyte development is regulated by the combined dosage of three basic helix-loop-helix genes, E2A, E2–2, and HEB. Mol Cell Biol 1996; 16: 2898–2905.
Flora A, Garcia JJ, Thaller C, Zoghbi HY . The E-protein Tcf4 interacts with Math1 to regulate differentiation of a specific subset of neuronal progenitors. Proc Natl Acad Sci USA 2007; 104: 15382–15387.
Pattyn A, Goridis C, Brunet JF . Specification of the central noradrenergic phenotype by the homeobox gene Phox2b. Mol Cell Neurosci 2000; 15: 235–243.
de Pontual L, Nepote V, Attie-Bitach T, et al. Noradrenergic neuronal development is impaired by mutation of the proneural HASH-1 gene in congenital central hypoventilation syndrome (Ondine's curse). Hum Mol Genet 2003; 12: 3173–3180.
Guillemot F, Lo LC, Johnson JE, Auerbach A, Anderson DJ, Joyner AL . Mammalian achaete-scute homolog 1 is required for the early development of olfactory and autonomic neurons. Cell 1993; 75: 463–476.
Persson P, Jogi A, Grynfeld A, Pahlman S, Axelson H . HASH-1 and E2–2 are expressed in human neuroblastoma cells and form a functional complex. Biochem Biophys Res Commun 2000; 274: 22–31.
Aretz S, Stienen D, Uhlhaas S, et al. High proportion of large genomic deletions and a genotype phenotype update in 80 unrelated families with juvenile polyposis syndrome. J Med Genet 2007; 44: 702–709.
van Hattem WA, Brosens LA, de Leng WW, et al. Large genomic deletions of SMAD4, BMPR1A and PTEN in juvenile polyposis. Gut 2008; 57: 623–627.
Calva-Cerqueira D, Chinnathambi S, Pechman B, Bair J, Larsen-Haidle J, Howe J . The rate of germline mutations and large deletions of SMAD4 and BMPR1A in juvenile polyposis. Clin Genet 2009; 75: 79–85.
Gallione CJ, Repetto GM, Legius E, et al. A combined syndrome of juvenile polyposis and hereditary haemorrhagic telangiectasia associated with mutations in MADH4 (SMAD4). Lancet 2004; 363: 852–859.
Zhuang Y, Barndt RJ, Pan L, Kelley R, Dai M . Functional replacement of the mouse E2A gene with a human HEB cDNA. Mol Cell Biol 1998; 18: 3340–3349.
Hosoda H, Motohashi J, Kato H, Masushige S, Kida S . A BMAL1 mutant with arginine 91 substituted with alanine acts as a dominant negative inhibitor. Gene 2004; 338: 235–241.
Gurnett CA, Alaee F, Kruse LM, et al. Asymmetric lower-limb malformations in individuals with homeobox PITX1 gene mutation. Am J Hum Genet 2008; 83: 616–622.
Jiao H, Toth B, Erdos M, et al. Novel and recurrent STAT3 mutations in hyper-IgE syndrome patients from different ethnic groups. Mol Immunol 2008; 46: 202–206.
Su MA, Giang K, Zumer K, et al. Mechanisms of an autoimmunity syndrome in mice caused by a dominant mutation in Aire. J Clin Invest 2008; 118: 1712–1726.
Yang L, Zhao T, Shi X, et al. Functional analysis of a dominant negative mutation of interferon regulatory factor 5. PLoS One 2009; 4: e5500.
Willing MC, Deschenes SP, Slayton RL, Roberts EJ . Premature chain termination is a unifying mechanism for COL1A1 null alleles in osteogenesis imperfecta type I cell strains. Am J Hum Genet 1996; 59: 799–809.
Sweetman D, Smith T, Farrell ER, Chantry A, Munsterberg A . The conserved glutamine-rich region of chick csal1 and csal3 mediates protein interactions with other spalt family members. Implications for Townes-Brocks syndrome. J Biol Chem 2003; 278: 6560–6566.
Borozdin W, Steinmann K, Albrecht B, et al. Detection of heterozygous SALL1 deletions by quantitative real time PCR proves the contribution of a SALL1 dosage effect in the pathogenesis of Townes-Brocks syndrome. Hum Mutat 2006; 27: 211–212.
Ballif BC, Theisen A, McDonald-McGinn DM, et al. Identification of a previously unrecognized microdeletion syndrome of 16q11.2q12.2. Clin Genet 2008; 74: 469–475.
Acknowledgements
We thank Sara Minier (Signature Genomic Laboratories, Spokane, WA) for her assistance with Figure 1 and Aaron Theisen (Signature Genomic Laboratories) for editing our manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
The opinions or assertions contained herein are the private views of the authors and are not to be construed as official or reflecting the views of the Department of the Army or the Department of Defense.
Shaffer and Bejjani sit on the Members' Board, are employees, and are part owners of Signature Genomic Laboratories, LLC. Rosenfeld, Ballif, and Torchia are employees of Signature Genomic Laboratories.
Rights and permissions
About this article
Cite this article
Rosenfeld, J., Leppig, K., Ballif, B. et al. Genotype–phenotype analysis of TCF4 mutations causing Pitt-Hopkins syndrome shows increased seizure activity with missense mutations. Genet Med 11, 797–805 (2009). https://doi.org/10.1097/GIM.0b013e3181bd38a9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1097/GIM.0b013e3181bd38a9
Keywords
This article is cited by
-
The role of GABAergic signalling in neurodevelopmental disorders
Nature Reviews Neuroscience (2021)
-
Transcription factor 4 and its association with psychiatric disorders
Translational Psychiatry (2021)
-
Disease-causing variants in TCF4 are a frequent cause of intellectual disability: lessons from large-scale sequencing approaches in diagnosis
European Journal of Human Genetics (2018)
-
Molecular Mechanisms of Transcription Factor 4 in Pitt-Hopkins Syndrome
Current Genetic Medicine Reports (2017)
-
Phenotype and natural history in 101 individuals with Pitt-Hopkins syndrome through an internet questionnaire system
Orphanet Journal of Rare Diseases (2016)