Abstract
Purpose: To determine the carrier frequency of familial Mediterranean fever (FMF) mutations of individuals in three different US testing populations: Cystic fibrosis, Factor V Leiden, and Ashkenazi Jews.
Methods: DNA samples from 1234 anonymous samples were screened for 12 FMF mutations using a laboratory-developed test.
Results: Genotyping revealed carrier frequencies of 1:16, 1:46, and 1:8, respectively.
Conclusion: MEFV mutation frequency seems to correlate positively with Mediterranean influence of the tested population and the high overall carrier rate for MEFV mutations in the Factor V Leiden testing population (1:46) suggests that the disease may be under-diagnosed in the US population or that the mutant alleles have a low penetrance.
Similar content being viewed by others
Main
Familial Mediterranean fever (FMF) is an autosomal recessive disorder (MIM 249100) mainly affecting Mediterranean populations such as Turks, Arabs, Jews, and Armenians. Carrier rates in these populations can be as high as 1 in 6.1,2 FMF is an autoinflammatory disease characterized by seemingly unprovoked attacks of fever, abdominal or chest pain, and arthritis. Prophylactic colchicine therapy is generally prescribed to prevent amyloidosis, which can lead to kidney failure and other organ damage.3 The disease is caused by mutations in the MEFV gene, which encodes pyrin (or marenostrin). Pyrin is presumed to act as a down-regulator of inflammation in polymorphonuclear cells.4
MEFV is located on chromosome 16 (16p13) and contains 10 exons. The Infevers database (http://fmf.igh.cnrs.fr/ISSAID/infevers/) currently reports 51 FMF causing mutations in MEFV. Five mutations account for 70% of the deleterious alleles in susceptible populations: M694V (2080A>G), V726A (2177T>C), M680I (2040G>C or A), M694I (2082G>A), and E148Q (442G>C).5 Four other mutations are more frequent in certain populations: A744S (2230G>T) and I692del (2074–2076del) in Arabs, R761H (2282G>A) in Armenians, and F479L (1437C>G) in Greeks. Our assay was designed to detect the mutations mentioned above as well as P369S (1105C>T) and K695R (2084A>G). All sequence variants correspond to National Center for Biotechnology Information reference sequence NM_000243.
There is a paucity of data regarding the frequency of FMF mutations in the US population. Quest Diagnostics receives samples for cystic fibrosis (CF), Factor V Leiden (FV), and Ashkenazi Jewish (AJ)-associated mutation screening from throughout the United States. The AJ panel includes tests for Gaucher Disease, Tay-Sachs disease, Nieman Pick Disease, Bloom syndrome, Canavan disease, Familial dysautonomia, Fanconi anemia, and Mucolipidosis Type IV mutations. To establish the carrier rate of FMF-associated mutations in these screening populations, we examined the frequencies of the 12 mutations in deidentified samples previously submitted for CF, FV, and AJ screening tests.
MATERIALS AND METHODS
DNA samples
A total of 1234 clinical specimens previously submitted to our laboratory for CF (n = 445), FV (n = 459), and AJ (n = 330) screening tests were analyzed for the E148Q, P369S, M694I, M694V, I692del, R761H, M680I (G>A and G>C), A744S, K695R, F479L, and V726A mutations of the MEFV gene. All samples were deidentified before analysis.
Specimens and DNA extraction
Genomic DNA was extracted from ethylenediaminetetraacetic acid- or acid-citrate-dextrose-anticoagulated whole-blood specimens using the Genovision GenoMTM-96 platform (Qiagen, Foster City, CA) according to the manufacturer's instructions.
Detection of FMF mutations
A laboratory-developed test using the ABI PRISM SNaPshot Multiplex Kit and Genotyper V3.7 by ABI (Foster City, CA) was used to detect the E148Q, P369S, M694I, M694V, I692del, R761H, M680I (G>A and G>C), A744S, K695R, F479L, and V726A mutations. Briefly, regions of exons 2, 3, 5, and 10 of the MEFV gene were amplified by multiplex polymerase chain reaction (PCR) and the amplicons were used as templates in single base-pair extension (SNaPshot) reactions. Extension products were extended by one fluorescent-labeled dideoxynucleotide, followed by fluorescent detection on an ABI 3100 Genetic Analyzer (Fig. 1).
SNaPshot results of a sample containing a novel variant (A744T, 2230G>A) in the MEFV gene predicted to be benign by PolyPhen (http://genetics.bwh.harvard.edu/pph/).
Exons 2, 3, 5, and 10 of the MEFV gene were amplified from genomic DNA using the following primers: 5′-GTGGGACAGCTTCATCATTTTG-3′ and 5′-CCTTCTCTCTGCGTTTGCTC-3′ (exon 2), 5′-CAGGACTCCCATGAAAGGAA-3′ and 5′-GACTGCAGATGAGGCAGATG-3′ (exon 3), 5′-TCTGCTCCCCAGAAACAAAC-3′ and 5′-GGGTGTCATATGCCTTCCTG-3′ (exon 5), 5′-TTACTGGGAGGTGGAGGTTG-3′ and 5′-GAGGAGCTGTGTTCTTCCC-TC-3′ (exon 10). A 2 μL aliquot of each sample DNA was added to 23 μL FMF PCR Master Mix (In-house preparation using Fast Start Taq, Roche Molecular Diagnostics, Pleasanton, CA) containing 0.40 μM each primer, 1× PCR buffer, 3 mM MgCl2, 5% dimethylsulfoxide, and 0.4 mM dinucleotide triphosphates. PCR amplification was performed under the following conditions: one cycle of 5 minutes at 95°C; 37 cycles of 30 seconds at 95°C, 30 seconds at 61°C, and 45 seconds at 72°C; and one cycle of 2 minutes at 72°C, followed by a 4°C hold. All PCR amplifications were performed in an ABI 9600 thermal cycler (ABI Foster City, CA).
Amplicon interrogation
A 2-μL aliquot of PCR product was digested with 1 U exonuclease and 1 U calf alkaline intestinal phosphatase at 37°C for 2 hours. A 3-μL aliquot of the inactivated digest was used as a template for a SNaPshot reaction. The following is a list of the extension primers used and the mutation interrogated: 5′-TGC GGT GCA GCC AGC CC-3′ E148Q; 5′-TTTTTTTTTTTT CACGTCCTCCAGTGAGGCCACAAA-3′ F479L; 5′-ACACTGTGGCAGGGGCTGGG-3′ P369S; 5′-ACTGACTGACTGACTGACTGACATCCATAAGCAGGAAAGGGAACA T-3′ M680I; 5′-TGACTGACTCAGAGAATGGCTACTGGGTGGTGAT-3′ I692del; 5′-TGACTTGGACGCCTGGTACTCATTTTCCTT-3′ M694I; 5′-CTGAGA ATGGCTACTGGGTGGTGATAATG-3′ M694V; 5′-CTGGACGCCTGGTACTCA TTTTCC-3′ K695R; 5′-TGACTGACTGACTGTGGGCATCTTCGTGGACTACAG AG-3′ V726A; 5′-TGACTGACTGACTGACTGACTTGACAGCCAGATCCCACA TCTATACATTC-3′ A744S; and 5′-TGACTGACTGACTGACTCAACCTATCTTCAGCC CTGGGACAC-3′ R761H. The products (fluorescently labeled oligonucleotides) were detected on an automated DNA sequencer (e.g., ABI PRISM 3100 Genetic Analyzer) both by size and by their fluorescent labels.
RESULTS
Six of the 12 FMF-associated mutations and one novel variant (A744T, 2230G>A) were detected (Tables 1 and 2; Figs. 1 and2). Not all mutations were detected, which is probably due to the limited representation of ethnic groups. E148Q was the most common mutation detected with in all three testing populations (Table 1). A total of 14 individuals with two MEFV mutations were found: two from the FV population, seven from the CF population, and five from the AJ population.
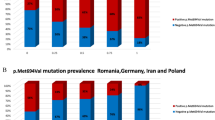
The frequency of the mutations detected in the three study groups, and all groups combined, is shown here. Each study group is represented by the shading indicated in the legend. FV, Factor V Leiden (solid black); CF, cystic fibrosis (wavy lines); AJ, Ashkenazi Jewish (solid gray); combined (solid white) represents all of the samples tested. The total frequency of all the tested alleles in each group is denoted as “All mut alleles.” “Exclude E148Q” and “Exclude P369S” represent the total frequency of all alleles tested except E148Q and P369S, respectively.
The overall carrier frequency of all mutations was higher than expected (1:15). The mutation frequency differed between screening groups, being lowest in the FV screening group (1:46) and highest in the AJ group (1:8) (Table 1; Fig. 2). The mutation frequency seemed to correlate positively with Ashkenazi genetic influence. That is, the frequency was lowest in the FV screening group (1:46), which is primarily Caucasian; intermediate in the CF group (1:16), comprising a mixture of Caucasian, AJ, and other populations; and highest in the AJ group (1:8), which is predominantly European. Excluding the mild E148Q mutation reduced the overall carrier frequency by more than half (1:32).
DISCUSSION
Minimal data are available on FMF mutation frequency in the American population. Here, we demonstrate that mutations in US AJ testing populations were detected at frequencies similar to those reported elsewhere: 1:8 overall versus 1:5 overall.6 Stoffman et al.6 looked at MEFV mutations in 400 healthy members of four different ethnic groups. The relative frequencies of the mutations found in our study are concordant with the results of Stoffman et al.6 where E148Q accounted for 53%, V726A accounted for 16%, and P369S accounted for 7.95% of the mutations found in Ashkenazi Jews. They observed the M694V mutation more frequently (29%) in healthy individuals with a mutation, whereas M694V only accounted for 1.25% of the mutations found in our AJ population. In another study, 100 unrelated patients referred to National Institutes of Health for possible FMF were genotyped to investigate ethnic distributions of mutations and to identify genotype–phenotype correlations.7 Along with identifying several new mutations (K695R, A744S, R761H, and E148Q), Samuels et al.7 found V726A, M694V, and E148Q to be the most common mutations. Also, a complex allele, E148Q/ V726A (in cis) was found in four individuals. See Table 3 for a comparison of the mutation frequencies for AJ populations in three studies.
Jews, Armenians, Turks, and Arabs are the four populations most commonly affected by FMF. Although 21% of the patients with mutations in the Samuels study were Ashkenazi Jews, it is significant to note that 19% were Italian, 2% were Greek, and 2% were Cuban. Some of these populations might account for the mutations found in our study of the US population. Samuels et al.7 found the V726A (31%) mutation more often than the E148Q (24%) mutation in his FMF AJ cohort, whereas we found E148Q (51%) more often than V726A (19%). Because the E148Q mutation is either not a disease causing mutation or has low penetrance, we expected to find E148Q in a higher proportion than Samuels et al.7 did because their population comprised people suspected to be affected by FMF. In fact, Stoffman et al.6 had healthy individuals in their study who were homozygous for E148Q. This is concordant with our data, as we found seven homozygotes.
MEFV mutation frequency seems to correlate positively with AJ influence of the tested population in our data in agreement with Stoffman et al.6 It is possible that the high frequency of MEFV mutations in the CF population is due to a modifier effect of MEFV on CF rather than a putative high rate of AJ individuals though most individuals screened for CF are wild-type. The high carrier rate for MEFV mutations in the FV population (1:46) seems to suggest that the disease may be under-diagnosed in the US population but in fact, mutations E148Q, K695R, and V726A are reported to have reduced penetrance.8–10 Nevertheless, the AJ population in the US can benefit from FMF mutation detection since early treatment improves prognosis for those with disease-causing mutations. In our series of presumably healthy individuals, we detected 14 with two FMF mutations. Because these samples were de-identified, it is impossible to determine if any of these individuals had symptoms.
MEFV mutations were observed in a population (FV screening) not predicted to contain AJ heritage nor to derive from the Mediterranean region. This suggests that other European ethnicities might have contributed FMF genotypes to this population, perhaps Italians or other Mediterranean Europeans as suggested by Samuels et al.7 Based on the prevalence of mutations in a non-Mediterranean population, this data suggest that screening for FMF mutations in the United States may be warranted when there is sufficient clinical suspicion.
References
Sohar E, Gafni J, Pras M, Heller H . Familial Mediterranean fever. A survey of 470 cases and review of the literature. Am J Med 1967; 43: 227–253.
Livneh A, Langevitz P, Zemer D, Zaks N, et al. Criteria for the diagnosis of familial Mediterranean fever. Arthritis Rheum 1997; 40: 1879–1885.
Cazeneuve C, Sarkisian T, Pecheux C, Dervichian M, et al. MEFV-gene analysis in Armenian patients with familial Mediterranean fever: diagnostic value and unfavorable renal prognosis of the M694V homozygous genotype-genetic and therapeutic implications. Am J Hum Genet 1999; 65: 88–97.
Richards N, Schaner P, Diaz A, Stuckey J, et al. Interaction between pyrin and the apoptotic speck protein (ASC) modulates ASC-induced apoptosis. J Biol Chem 2001; 276: 39320–39329.
Bakkaloglu A . Familial Mediterranean fever. Pediatr Nephrol 2003; 18: 853–859.
Stoffman N, Magal N, Shohat T, Lotan R, et al. Higher than expected carrier rates for familial Mediterranean fever in various Jewish ethnic groups. Eur J Hum Genet 2000; 8: 307–310.
Samuels J, Aksentijevich I, Torosyan Y, Centola M, et al. Familial Mediterranean fever at the millennium: clinical spectrum, ancient mutations, and a survey of 100 American referrals to the National Institutes of Health. Medicine (Balt) 1998; 77: 268–297.
Ben-Chetrit E, Lerer I, Malamud E, Domingo C, et al. The E148Q mutation in the MEFV gene: is it a disease-causing mutation or a sequence variant?. Hum Mutat 2002; 15: 385–386.
Tchernitchko D, Legendre M, Cazeneuve C, Delahaye A, et al. The E148Q MEFV allele is not implicated in the development of familial Mediterranean fever. Hum Mutat 2000; 24: 339–340.
Aksentijevich I, Torosyan Y, Samuels J, Centola M, et al. Mutation and haplotype studies of familial Mediterranean fever reveal new ancestral relationships and evidence for a high carrier frequency with reduced penetrance in the Ashkenazi Jewish population. Am J Hum Genet 1999; 64: 949–962.
Acknowledgements
This work is supported by Quest Diagnostics.
Author information
Authors and Affiliations
Corresponding author
Additional information
Disclosure: All authors with the exception of Susan Olson declare ownership of stock in Quest Diagnostics.
Rights and permissions
About this article
Cite this article
Mikula, M., Buller, A., Sun, W. et al. Prevalence of known mutations in the familial Mediterranean fever gene (MEFV) in various carrier screening populations. Genet Med 10, 349–352 (2008). https://doi.org/10.1097/GIM.0b013e3181723cc8
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1097/GIM.0b013e3181723cc8
Keywords
This article is cited by
-
Clinical and molecular evaluation of MEFV gene variants in the Turkish population: a study by the National Genetics Consortium
Functional & Integrative Genomics (2022)
-
MEFV gene mutations and cardiac phenotype in children with familial Mediterranean fever: a cohort study
Pediatric Rheumatology (2014)