Abstract
Purpose: The polymerase chain reaction is generally used for mutational analysis of the galactose-1-phosphate uridyl transferase (GALT) gene in the diagnosis of galactosemia. This method is problematic when used in families of Ashkenazi Jewish descent.
Methods: We amplified the GALT gene from leukocyte DNA followed by allele specific oligonucleotide hybridization, DNA sequencing and Southern Blot analysis to determine the mutant alleles causing galactosemia in a representative Jewish family.
Results: The proband's diagnosis of galactosemia was confirmed by high levels of erythrocyte galactose-1-phosphate, absence of erythrocyte GALT activity and impaired total body oxidation of galactose to expired CO2. Initial molecular analysis of GALT alleles in the family showed homozygosity for a K285N missense mutation in the proband, homozygosity for N314D in the mother and heterozygosity for N314D and K285N in the father. These results contradicted Mendelian logic. Southern blot hybridization with GALT cDNA proved the presence of a complex 5 kb GALT deletion in the proband and her mother's DNA enabling a corrected genotype.
Conclusions: Since a deletion of the GALT gene is a common mutation causing galactosemia among Ashkenazim Jewish families, this deletion should be suspected and tested for by genomic hybridization or by using primers specific for the 5 kb deletion.
Similar content being viewed by others
Main
Classic galactosemia is a well-described autosomal recessive disorder characterized by symptoms and signs presenting shortly after birth that include jaundice, poor feeding, hepatic dysfunction and E coli sepsis.1,2 This disease is caused by mutations in the galactose-1-phosphate uridyl transferase (GALT) gene.3,4 Mutations in GALT result in loss of enzymatic activity and in the accumulation of galactose-1-phosphate (Gal-1-P), galactose and galactitol.1,2 High concentrations of Gal-1-P impair the UTP dependent glucose-1-phosphate pyrophosphorylase and initiate environmental stress response in yeast and human diploid cells. These toxic effects produce cell death and are probably related to organ dysfunctions.5,6
Gene variants
Over 180 mutations in the GALT gene have been identified and many occur with increased frequency in specific ethnic groups.7,8 Three common mutations, Q188R, K285N and N314D account for 70% of the galactosemia mutations in the white population while S135L accounts for 62% of the alleles causing galactosemia in the African-American population.7,8 A large 5 kb deletion was first described in 1996 in Jewish Ashkenazim.4,10 Lymphoblastoid cell lines from patients homozygous for this 5kb deletion have provided evidence for alternate pathways for galactose oxidation.11
Clinical context
In the United States all newborns are screened for galactosemia by the Beutler Method, which measures enzyme linked NADPH production from glucose-1-P by fluorescence.12 Some States also quantitate total galactose and galactose-1-phosphate. If there is no fluorescence or partial fluorescence, diagnosis is then accomplished by measuring Gal-1-P and GALT activity in erythrocytes, and identifying mutations in the GALT gene.9 Management of galactosemia includes a galactose restricted diet.2 Long-term sequelae of galactosemia may include ataxia, verbal apraxia, growth and developmental restriction, and premature ovarian failure.13–15 The best predictors of outcome are total body galactose oxidation to CO2 and the specific mutations of the GALT gene.14,15
Here we report an exemplary family whose proband presented with classical galactosemia, high levels of erythrocyte Gal-1-P, absent erythrocyte GALT and markedly impaired oxidation of 13C-Galactose to 13CO2 by breath test. Initial, DNA, polymerase-based methods for mutational analysis of the family's GALT alleles indicated that the proband was homozygous for the K285N, the mother was homozygous for the N314D and the father heterozygous for K285N and N314D mutations. Uniparental disomy was considered because the father, but not the mother had the K285N allele. These results were considered unlikely because they contradicted Mendelian logic and there are no publications suggesting uniparental disomy in the GALT region of chromosome 9. Solutions to this problem are presented below.
MATERIALS AND METHODS
Patient's clinical history
This female infant was the 6 pounds, 10 ounces product of a full term gestation born by cesarean section. Both parents were of Jewish Ashkenazi descent. The newborn had worsening of neonatal jaundice, which progressed by age 11 days to bloody stools, thrombocytopenia, elevated liver enzymes and deranged partial prothrombin time (PTT). The report of her first newborn screen indicated low GALT activity (1.4 units/g of hemoglobin per hour), with a second screen at 10 days of age being 0.7 units/g of hemoglobin per hour. She was referred to the Center for Medical Genetics at the University of Miami where a lactose-free formula was initiated at age 12 days with rapid improvement in liver function. Diagnostic, biochemical and molecular analyses of her GALT gene and enzyme as well as total body oxidation of 1 13C-D-Galactose to 13CO2 in breath were completed. The latter study followed informed parental consent and approval by the Internal Review Committees of both the Florida State Department of Health and The University of Miami.
METHODS
Total body oxidation of galactose
The total body oxidation of galactose was quantitated following an oral bolus of 13C galactose (7 mg/kg) by 13CO2 enrichment of expired air using dual isotope gas mass spectrometry (Europa). The results were expressed as the cumulative percent dose (CUMPD) recovered over 120 minutes.16
GALT enzyme activity assay
GALT enzyme activity in red blood cells (RBC's) was assayed by measuring the conversion of [14C]-galactose-1-phosphate ([14C]-Gal-1-P) and UPD-glucose (UDPG) to glucose-1-phosphate (Glu-1-P) and 14C-UDP-galactose at 37°C for 1 hour by erythrocyte hemolysate normalized to hemoglobin content.9
Multiplex PCR and ASO hybridization
The GALT alleles were amplified using three sets of primers: IVS2-F, S135L-R, GALT 6-5, GSINT7R, GALT 9-5, and INTJR. Primer sequences are shown in Table 1. Genomic DNA was isolated from blood buffy coat and used as a template. The amplified fragments were labeled with digoxigenin during PCR by introducing Dig-dUTP (Roche, Indiana, USA). Allele specific oligonucleotide (ASO) hybridization was performed using Roche PCR ELISA (DIG) detection kit (Roche, cat #1965409). Denatured, digoxigenin-labeled-PCR fragments were hybridized to specific biotinylated oligonucleotides corresponding to either normal or mutated sequence of each altered allele. The hybridized products were trapped in streptavidin-coated microwells and detected by antidigoxigenin peroxidase conjugated antibodies. Hybridized products developed a green color after incubation with peroxidase substrate (Fig. 1). This method enabled us to detect the presence of the seven most common mutations of the GALT gene–Q188R, N314D, S135L, L195P, Y209C, IVS2 and K285N.2,4 Notably this method will not detect GALT deletions because PCR amplification is required.
Sequencing
Exon 9 was amplified by PCR using genomic DNA as a template and two primers: GALT 9-5 and GALT 10-3 (see Table 1). The reaction ran for 40 cycles: 30 seconds at 95°C, 30 seconds at 58°C, 1 minute, 30 seconds at 72°C. The PCR product was gel-purified and sequenced using GALT 9-5 as primer.
Southern blot analysis
Genomic DNA samples from the proband, mother and father were digested with EcoR1 restriction enzyme. Equal amounts of DNA were electrophoresed on 0.7% agarose gel and transferred to nylon membrane. Bands specific for the GALT gene were identified by hybridization with GALT cDNA labeled with [32P]dATP. For control, GALT-positive bands were normalized to actin for quantitation.
RESULTS
The proband presented in the neonatal period with classical galactosemia confirmed by high levels of Gal-1-P [25.1 mg/dL (normal <1.0 mg%)] and absent activity of erythrocyte GALT [normal = 36.3 ± 1.7 μmoles/gHbg/(Hr)] (Fig. 2, Table 2). Both parents had biochemical phenotypes of carriers for GALT deficiency (Table 2; Fig. 2). The GALT genotypes were compared among mother, father and proband following multiplex PCR of their GALT DNA (Fig. 1). Note that from this initial PCR-based analysis, the proband appeared homozygous for the K285N mutation, the mother homozygous for the N314D mutation and the father, compound heterozygous for both the K285N and the N314D mutations. The biochemical phenotypes and molecular genotypes for all three family members from these initial studies were tabulated in Table 2. The initial data did not follow Mendelian logic because no maternal GALT allele was found in the proband. Although never before reported, it was conceivable that the proband's GALT alleles were derived by uniparental, paternal disomy of his K285N allele. Further sequencing of PCR amplified GALT DNA found only the G to T transversion at nucleotide 855 in the proband. Since the family history indicated that both parents were of Ashkenazim Jewish ancestry, it was likely that the mother and her affected daughter carried a 5 kb GALT deletion that was not amplified by PCR.2,10 To test this latter hypothesis, we performed a Southern blot for GALT on genomic DNA from a control, the proband and the proband's mother (Fig. 3). Equal amounts of DNA were loaded on the gel but the signal from the proband and her mother's DNA were less than half that of control. The father's GALT DNA signal was similar to control, confirming that the mother had transmitted her deleted GALT allele to her daughter while the father had transmitted his K285N mutant allele. Both the initial and the corrected interpretations are shown in Table 2.
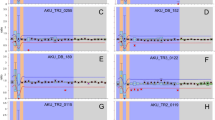
Southern blot hybridization. DNA isolated from proband her mother and a control were digested with EcoRI endonuclease and separated on 0.7% agarose gel. One band around 10 Kb was positive for GALT DNA. For loading control the same filter was probed with human actin cDNA. Quantitation results of GALT band normalized to loading control are shown in the table and pictured on the graph.
GALT activity assay. Activity of GALT enzyme of erythrocyte lysates was measured by the conversion of [14C]-Gal-1-P to 14C-UDP-gal for one hour. Reaction mixtures were loaded onto DEAE-TLC plates and separated in 0.05 M LiCl. The amount of radioactivity transferred to UDP-gal was quantitated in relation to a total amount of radioactivity added to the reaction and normalized to hemoglobin concentration. Lines 1, 4, and 7 - no UDP-glucose added. Lines 2, 3, 5, 6, 8, and 9 - both [14C]-Gal-1-P and UDP-glucose are added in duplicate reactions. Table on the right shows the calculated values for GALT activity for analyzed samples. N/D, not detected.
The proband's CUMPD at 1.5 years of age was 0.17% which was again consistent with classical (G/G) galactosemia. Normal values in children less than two years of age range from (14–28%) (unpublished data).16 This test confirmed the functional effects of her mutations, and helped in planning her continued care.
DISCUSSION
All states in the United States now have population-based newborn screening for galactosemia using dried blood on filter paper.2,17 Few states have established second-tier diagnostic analyses for the GALT biochemical phenotype and molecular genotype. The biochemical phenotype should include quantitative enzyme analysis and erythrocyte, Gal-1-P.2,9 Total body oxidation of 13C-Galactose to 13CO2 in expired air is a research tool at present and is a valid prediction of outcome in galactosemia.14–16 The molecular genotype could begin with simple hybridization methods for allele-specific oligonucleotides representing the most common mutations causing galactosemia as demonstrated in Figure 1. However, the 5 kb deletion of the GALT gene will be missed and this PCR-based approach will provide erroneous results as exemplified in this family and the initial interpretation outlined in Table 2.
Why is it necessary to fulfill a complete biochemical and molecular analysis on screen-positive newborns for galactosemia? First and foremost to confirm or deny the fluorescent screening assay. Many false positives occur in particular during hot, humid seasons that will result in unwarranted restriction of breast milk, stigmatization of the infant and parental anxiety.18 Second, the family and the primary care physician will want to know the prognosis of their affected child. Although severe, neonatal hepatocellular disease may be ameliorated in the newborn child with galactosemia, dyspraxic speech, ovarian failure, cognitive dysfunction and growth restriction may occur.13–15 The best predictors of these outcomes are the results of total body oxidation of galactose to expired CO2 and the GALT genotype.14,15 Given the results of these predictors, appropriate interventions can be offered for dyspraxia and ameliorate the systemic effects of ovarian failure. Genetic counseling in other family members at risk for being carriers of a GALT mutation and for prenatal diagnosis require these sensitive and specific molecular genetic discriminants.19 Finally providing screening and second tier biochemical and molecular diagnostics for galactosemia would be cost-effective. In Florida about 1/10,000 of the 220,000 newborns per year will screen-positive. Of these 22 newborns, 7 will have true galactosemia (G/G). These diagnostic tests cost about $500 per patient times 22 patients per year or about $11,000 per year. The costs of screening are approximately the same ($500 per patient) for a grand total annually of $22,000. If we then consider avoided life-time cost of care for one mentally retarded child ($1,014,000.00) and the avoided productivity losses for each “saved” patient (2 to 5 times lifetime costs or about $2–5 million) the benefit to cost ratio per detected and treated case of galactosemia in dollars today is about twenty to one.20
This study emphasizes a well-known problem faced in accomplishing a correct genotypic diagnosis when a major gene deletion is present and mutational analysis is based on polymerase chain reaction-based amplification of DNA. If one (or both) mutant alleles are caused by a deletion of the entire gene or at critical sites for primer hybridization, the one remaining allele will be amplified and give the false impression of homozygosity for the remaining allele. In our example both the proband and her mother were initially incorrectly thought to be homozygous for their K285N and N314D alleles respectively because their deleted allele did not amplify (Fig. 1, Table 2). Southern blot analysis detected the GALT deletion in both the mother and proband (Fig. 3, Table 3). Methods for mutational analysis other than Southern blotting might achieve the same objective now that the 5 kb deletion breakpoint sequence is known.21
Primer sets are now designed that will amplify at the breakpoints only if the complex deletion-rearrangement is present.21 We conclude that family history, biochemical phenotypes and molecular genotypes of parents and their proband will enable sensitive and specific diagnosis and appropriate genetic counseling for patients with galactosemia.
References
Holton JB, Walter JH, Tyfield LA . Galactosemia. In: Scriver CR, Beaudet AL, Sly WS and Vallee N (eds.) The Metabolic and Molecular Bases of Inherited Diseases. 8th Edition. McGraw Hill: New York 2001: 1533–1587.
Elsas LJ . Galactosemia in Gene Clinics Reviews : http://www.geneclinics.org, March, 2005.
Leslie ND, Immerman EB, Flach JE, Florez M, et al. The Human Galactose-1-Phosphate Uridyl Transferase Gene. Genomics 1992; 14: 474–480.
Elsas LJ, Lai K . The Molecular Biology of Galactosemia. Gen in Med 1998; 1: 40–48.
Lai K, Langley S, Kwaja F, Schmitt E, et al. Galactose-1-Phosphate Uridyl Transferase Deficiency causes UDP-Hexose Deficit in Human Galactosemic Cells. Glycobio 2003; 13: 285–294.
Slepak T, Tang M, Addo F, Lai K . Intracellular galactose-1-phosphate accumulation leads to environmental stress response in yeast model. Mol Gen and Met 2005; 86: 360–371.
Lai K, Elsas LJ . Structure-Function Analyses of a Common Mutation in Blacks with Transferase-Deficiency Galactosemia. Mol Gen and Met 2001; 74: 264–272.
Lai K, Langley SD, Singh R, Dembure PP, et al. A prevalent Mutation for Galactosemia Among Black Americans. J Peds 1996; 128: 89–95.
Elsas LJ, Langley S, Steele E, Evinger J, et al. Galactosemia: A Strategy to Identify New Biochemical Phenotypes and Molecular Genotypes. Am J Hum Gen 1995; 56: 630–639.
Muralidharan K, Zhang W, Dembure P, Langley SD, et al. A Naturally Occurring “Knockout” Mutation of Human Galactose-1-Phosphate Uridyl Transferase Gene Metabolism. Am J Hum Gen 1996; 59: A204.
Berry GT, Leslie N, Reynolds RA, Yager CT, et al. Evidence for Alternate Galactose Oxidation in a Patient with Deletion of Galactose-1-Phosphate Uridyl Transferase Gene. Mol Gen and Met 2001; 72: 316–321.
Beutler E, Baluda M . Biochemical Properties of Human Red blood Cell Galactose-1-Phosphate Uridyl Transferase (UDP Glucose: α-D-Galactose-1-Phosphate Uridyl Transferase) from Normal and Mutant Subjects. J Lab Clin Med 1966; 67: 947–954.
Waggoner DD, Buist NRM, Donnell GN . Long-Term Prognosis in Galactosemia: Results of a Survey of 350 Cases. J Inherit Metab. Dis 1990; 13: 802–818.
Guerrero NV, Singh RH, Manatunga A, Berry GT, et al. Risk Factors for Premature Ovarian Failure in Females with Galactosemia. J Peds 2000; 137: 833–841.
Webb AL, Singh RH, Kennedy MJ, Elsas LJ . Verbal Dyspraxia and Galactosemia. Ped Res 2003; 53: 396–402.
Berry GT, Singh RH, Mazur AT, Guerrero N, et al. Galactose Breath Testing Distinguishes Variant and Severe Galactose-1-Phosphate Uridyl Transferase Genotypes. Ped Res 2000; 48: 323–328.
Therrell B . National Newborn Screening and Genetics Resource Center, 2005. http://genes-r-us.uthscsa.edu.
Brown AL, Dembure PP, Drumheller J, Elsas LJ . Galactosemia Screening: Seasonal Incidence and Thermal Stability of Mutant Alleles. Am J Hum Genet 1987; 41: A1575.
Elsas LJ . Prenatal Diagnosis of Galactose-1-Phosphate Uridyl Transferase (GALT)-Deficient Galactosemia. Prenat Diagn 2001; 21: 302–303.
Economic costs associated with mental retardation, cerebral palsy, hearing loss and vision impairment — United States 2003. Center for Disease Control (CDC) Morb. Mortal. Wkly. Report 2004; 53: 57–59.
Muralidharan K, Coffee B, DeLorenzo A, Courtney EM, et al. Galactosemia in Ashkenazi Jewish due to Galactose-1-Phosphate Uridyl transferase Gene Deletion. Abstracts from the 55th Annual Meeting of the American Society of Hum Genet, Salt Lake City, Utah, 2005; p 265.
Acknowledgements
This study was supported in part by The Dr. John T. Macdonald Foundation and grants from the USPHS; NIH 2-R44-DK60054-02A1 and GCRC M01RR16587.
Author information
Authors and Affiliations
Additional information
Presented in part at the Annual Clinical Genetics Meeting, March, 2004.
Rights and permissions
About this article
Cite this article
Barbouth, D., Slepak, T., Klapper, H. et al. Prevention of a molecular misdiagnosis in galactosemia. Genet Med 8, 178–182 (2006). https://doi.org/10.1097/01.gim.0000204019.54509.40
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1097/01.gim.0000204019.54509.40