Abstract
Purpose: In children there is frequently a reason to exclude cystic fibrosis. Sweat testing is used for this. Because sweat testing has some disadvantages we investigated whether analyzing DNA for the local most common CFTR mutations, harvested from buccal cells, is reliable as a method to exclude cystic fibrosis.
Methods: In patients in whom a sweat test had been ordered during the period January 1, 2002 to December 31, 2004, we harvested buccal cell DNA for analysis. When blood was available, DNA from leukocytes was also analyzed.
Results: A total of 73 sweat tests were ordered during the two-year study period, mostly because of recurrent pulmonary infections (36; 49%), failure to thrive (20; 27%) and chronic diarrhea (10, 14%). In 70, children the results of the sweat test were normal, in three patients the results were borderline. Sixty buccal smears were analyzed and no patient was homozygous for cystic fibrosis, two were heterozygous for cystic fibrosis. In none of the children the diagnosis of cystic fibrosis was established.
Conclusion: Analyzing DNA in cells, harvested from the buccal cells, is a reliable alternative to exclude cystic fibrosis. It is safe, simple, and child-friendly.
Similar content being viewed by others
Main
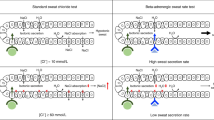
In children, there is quite often a reason to exclude Cystic Fibrosis (CF), e.g., when presenting with chronic respiratory infections or failure to thrive. Nowadays the sweat test is the gold standard. This test has the advantages of noninvasiveness and quick availability of the results. However, sweat testing has some disadvantages. The test cannot be performed in prematurely born infants; it is time consuming for children, their parents as well as for laboratory personnel. Moreover, the sweat test has been reported to have an unacceptably high false positive (up to 15%) and false-negative (up to 12%) rate, attributable to inaccurate methodology, technical errors, and patient physiology.1
An attractive alternative for diagnosing CF is DNA genotyping. It has been established that absence of 6 common mutations (including delta F508) excludes in most (>97%) cases the diagnosis of CF.2 The American College of Medical Genetics has proposed a panel of 25 mutations (each with an allele frequency > 0.1%) for the same purpose.3 Traditionally, DNA has been obtained from leukocytes. However, harvesting DNA from the buccal cells is noninvasive, and generates sufficient DNA for mutation analysis. Therefore, analysis of buccal cell DNA seems to be a reliable replacement for analysis of leukocyte DNA. We tested whether analysis of CF mutations in DNA obtained from buccal cells could serve as a reliable alternative to exclude CF as compared to sweat testing.
METHODS AND MATERIALS
This pilot study was undertaken in a large, general teaching hospital in the northern part of the Netherlands. All children, in whom the pediatrician felt it was necessary to order a sweat test, were eligible. When they were referred for sweat testing and their parents gave informed consent, buccal cells were harvested by brushing the inner side of the mouth with a swab (Epicenter, Madison, WI). When blood was available, DNA in leukocytes was also analyzed.
Sweat testing
Sweat tests were performed as described elsewhere.4 A chloride concentration > 40 mmol/l on two separate occasions is considered a positive test.
DNA isolation
DNA was isolated from buccal swabs using BuccalAmp™ DNA Extraction Kit (Epicenter) or from 200 μl whole blood using High Pure Template Kit (Roche) according to the manufacturer's instructions. DNA isolates were stored at −20°C until analysis.
Polymerase Chain Reaction (PCR)
PCR followed by restriction-fragment length polymorphism (RFLP) was performed to analyze delta F508, G542X, G551D, R553X, N1303K, and A455E according to previously described methods on a Perkin Elmer PE 2400 thermocycler.5 As an alternative, the delta F508 mutation was also analyzed by means of amplification refraction mutation system (ARMS) using the following primer combination: common reverse primer 5′GGGTAGTGTGAAGGGTTCATATGCATAATC3′, Wildtype Forward primer 5′GCCTGGCACCATTAAAGAAAATATCATCTT3′, and Mutant Forward primer 5′GCCTGGCACCATTAAAGAAAATATCATTGG3′.6
After PCR was performed, amplicons were digested (in the case of RFLP) using appropriate restriction enzymes as previously described. Finally, DNA fragments were separated on 3% Proranose LM-3 agarose gel (Sphaero Q) and visualized using ethidium bromide.
Demographics & statistical analysis
Data such as age and indication for ordering the test were collected. For analysis we used SPSS for Windows (version 11.0). All parents gave written informed consent. The local ethical committee approved this study.
RESULTS
In the study period of January 1, 2003 to December 31, 2004 a sweat test was ordered in 73 patients. The mean age was 5 years and 3 months and the median age was 3 years and 3 months (range 1 month to 34 years).
The reasons for ordering a sweat test were recurrent respiratory infections in 36 (49%); failure to thrive in 20 (27%), chronic diarrhea in 10 (14%), and evaluation of liver disease in 3 (4%). Other reasons were recurrent pancreatitis, delayed passing of first stool, nasal polyps, and severe iron deficiency (Table 1).
In 60 patients, buccal cells were collected and analyzed. Of 13 individuals no buccal cell DNA was obtained, due to parental refusal or violation of the study protocol. In 44 patients, leukocytes were obtained for DNA analysis.
In 70 patients, the chloride content of the sweat was normal. In two children, the results were 48 and 52 mmol/l; in both cases collecting sweat was difficult. The sweat chloride concentration in one child was 41 mmol/l, which after repeat testing was within the normal reference range. No homozygotes for CF were found. Two patients were heterozygous for CF (both delta F508). They both had normal sweat chloride levels (20 and 28 mmol/l), but unfortunately no leukocyte DNA was available.
In all children in whom leukocyte DNA was analyzed, this was normal. The agreement between the results of sweat testing and of DNA analysis in buccal cells was 100% at a clinical level. The agreement between analysis of DNA from buccal cells and leukocytes was 100%.
DISCUSSION
In this study we demonstrated the reliability and feasibility of DNA-analysis of buccal cells for the exclusion of CF in an outpatient setting.
This study has some limitations. First, the study group is relatively small. This is partly due to the fact that the number of children in which a sweat test is ordered has fallen dramatically in the recent years because national guidelines do not recommend sweat testing in the work up for children with recurrent wheeze.7 A second limitation is that in the study period the diagnosis of CF was not established in any child. This reflects the relatively low incidence of CF in this region. We believe our analysis is correct because we found two heterozygous children and in our patients with established CF the DNA analysis was abnormal.
The main reason for ordering a sweat test is in most cases to exclude CF. The risk for a Dutch child to have CF is 1 in 2,500.2 Based on this incidence and the sensitivity of the sweat test, the theoretical risk for a child to have CF with a negative sweat test result is 1 in 25,000.8 The risk of a child to have CF when one mutation is found is 1 in 80, and when none of the six mutations are found, the risk is 1 in 119,000. Therefore we argue that in these populations the analysis of buccal cell DNA is as least as reliable as sweat testing.
Analyzing DNA has some major advantages compared to sweat analysis. It is child-friendly because taking a smear is rapid and not painful. Sweat testing is difficult in neonates, and not even possible in prematurely born children. In these small children sampling buccal smear is possible and reliable.9,10 The fact that in two children not enough sweat could be produced is another problem of sweat testing. We could harvest DNA from every swab. In a large study Richards et al. demonstrated that the success rate of PCR amplification from buccal cells was 99% and that there was a 100% agreement of DNA analysis between buccal cells and blood.11 The costs of buccal cell sampling and DNA analysis costs approximately USD 35, which is as expensive as sweat testing. A disadvantage is that it takes more time to get a result.
In summary, we demonstrated that analyzing DNA, harvested from buccal cells, is a safe, simple, child friendly, and a reliable technique for excluding cystic fibrosis in outpatients and is comparable with sweat testing and analyzing leukocyte DNA.
References
LeGrys VA, Barfitt MF, Gibson LE, Hammond RB, et al. Sweat testing: sample collection and quantitative analysis; Approved Guideline, 2nd Ed. NCCLS 2000; 20: 1–31.
Van den Bergh FAJTM, Martes A . Diagnosis of cystic fibrosis; simple genotyping to rule out the disease preferable to starting with the sweat test. Ned Tijdschr Geneeskd 2003; 147: 1001–1005.
Grody WW, Cutting GR, Klinger KW, Richards CS, et al. Laboratory standards and guidelines for population based cystic fibrosis carrier screening. Genet Med 2001; 3: 149–154.
Scott MG, Heusel JW, LeGrys VA, Siggaard-Andersen O . In: Burtis C, Ashwood ER, editors. Tietz textbook of clinical chemistry. 3rd ed. Philadelphia, PA: Saunders company; 1999: 1068–1072.
Friedman KJ, Highsmith WE Jr, Silverman LM . Detecting multiple Cystic Fibrosis mutations by polymerase chain reaction-mediated-site-directed mutagenesis. Clin Chem 1991; 37: 753–755.
Ferry RM, Schwarz MJ, Robertson NH, Vaudin S, et al. Development, multiplexing, and application of ARMS tests for common mutations in the CFTR gene. Am J Hum Genet 1992; 51: 251–262.
Duiverman EJ, Brackel HJL, Merkus PJFM, Rottier BL, et al. Guideline ‘treating asthma in children’ for pediatric pulmonologists. (second revised edition). Diagnosis and prevention. Ned Tijdschr Geneeskd 2003; 147: 1905–11.
Ogino S, Wilson RB, Grody WW . Bayesian risk assessment for autosomal recessive diseases. J Med Genet 2004; 41: e70.
Bennett LC, Kraemer R, Liechti-Gallati S . Buccal cell DNA analysis in premature and term neonates. Eur J Ped 2000; 159: 99–102.
Richards B, Skoletsky J, Shuber AP, Balfour R, et al. Multiplex PCR amplification from the CFTR gene using DNA prepared from buccal brushes. Hum Molec Gen 1993; 2: 159–163.
Parad RB . Buccal cell DNA mutation analysis for diagnosis of cystic fibrosis in newborns and infants inaccessible to sweat chloride measurement. Pediatrics 1998; 101: 851–855.
Acknowledgements
We thank Dr. J. Collins for reviewing the English.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
de Vries, T., Ajubi, N., Slomp, J. et al. Analyzing DNA from buccal cells is a reliable method for the exclusion of cystic fibrosis. Results of a pilot study. Genet Med 8, 175–177 (2006). https://doi.org/10.1097/01.gim.0000201066.23642.b1
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1097/01.gim.0000201066.23642.b1