Abstract
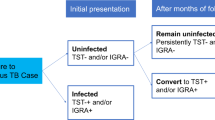
Human genetic susceptibility for tuberculosis (TB) has been demonstrated by several studies, but few have examined the multiple innate and adaptive immunity genes comprehensively, age-specific effects and/or resistance to Mycobacterium tuberculosis (Mtb) infection (resistors (RSTRs)). We hypothesized that RSTRs, defined by a persistently negative tuberculin skin test, may have different genetic influences than Mtb disease. We examined 29 candidate genes in pathways that mediate immune responses to Mtb in subjects in a household contact study in Kampala, Uganda. We genotyped 546 haplotype-tagging single-nucleotide polymorphisms (SNPs) in 835 individuals from 481 families; 28.7% had TB, 10.5% were RSTRs, and the remaining 60.8% had latent Mtb infection. Among our most significant findings were SNPs in TICAM2 (P=3.6 × 10−6) and IL1B (P=4.3 × 10−5) associated with TB. Multiple SNPs in IL4 and TOLLIP were associated with TB (P<0.05). Age–genotype interaction analysis revealed SNPs in IL18 and TLR6 that were suggestively associated with TB in children aged ⩽10 years (P=2.9 × 10−3). By contrast, RSTR was associated with SNPs in NOD2, SLC6A3 and TLR4 (nominal P<0.05); these genes were not associated with TB, suggesting distinct genetic influences. We report the first association between TICAM2 polymorphisms and TB and between IL18 and pediatric TB.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Mahan C, Zalwango S, Thiel B, Malone LL, Chervenak K, Baseke J et al. Innate and adaptive immune responses during acute M. tuberculosis infection in adult household contacts in Kampala, Uganda. Am J Trop Med Hyg 2012; 86: 690–697.
Stein CM, Zalwango S, Malone LL, Won S, Mayanja-Kizza H, Mugerwa RD et al. Genome scan of M. tuberculosis infection and disease in Ugandans. PLoS ONE 2008; 3: e4094.
Moller M, Hoal EG . Current findings, challenges and novel approaches in human genetic susceptibility to tuberculosis. Tuberculosis (Edinb) 2010; 90: 71–83.
Stein CM . Genetics of Susceptibility to Tuberculosis. Encyclopedia of Life Sciences. John Wiley & Sons, Ltd; Chichester, UK, 2012.
Ma N, Zalwango S, Malone LL, Nsereko M, Wampande EM, Thiel BA et al. Clinical and epidemiological characteristics of individuals resistant to M. tuberculosis infection in a longitudinal TB household contact study in Kampala, Uganda. BMC Infect Dis 2014; 14: 352.
Azad AK, Sadee W, Schlesinger LS . Innate immune gene polymorphisms in tuberculosis. Infect Immun 2012; 80: 3343–3359.
Berrington WR, Hawn TR . Mycobacterium tuberculosis, macrophages, and the innate immune response: does common variation matter? Immunol Rev 2007; 219: 167–186.
Stein CM . Genetic epidemiology of tuberculosis susceptibility: impact of study design. PLoS Pathog 2011; 7: e1001189.
Grant AV, El Baghdadi J, Sabri A, El Azbaoui S, Alaoui-Tahiri K, Abderrahmani RI et al. Age-dependent association between pulmonary tuberculosis and common TOX variants in the 8q12-13 linkage region. Am J Hum Genet 2013; 92: 407–414.
Leung KH, Yip SP, Wong WS, Yiu LS, Chan KK, Lai WM et al. Sex- and age-dependent association of SLC11A1 polymorphisms with tuberculosis in Chinese: a case control study. BMC Infect Dis 2007; 7: 19.
Alcais A, Fieschi C, Abel L, Casanova JL . Tuberculosis in children and adults: two distinct genetic diseases. J Exp Med 2005; 202: 1617–1621.
Awomoyi AA, Charurat M, Marchant A, Miller EN, Blackwell JM, McAdam KP et al. Polymorphism in IL1B: IL1B-511 association with tuberculosis and decreased lipopolysaccharide-induced IL-1β in IFN-γ primed ex-vivo whole blood assay. J Endotoxin Res 2005; 11: 281–286.
Gomez LM, Camargo JF, Castiblanco J, Ruiz-Narvaez EA, Cadena J, Anaya JM . Analysis of IL1B, TAP1, TAP2 and IKBL polymorphisms on susceptibility to tuberculosis. Tissue Antigens 2006; 67: 290–296.
Thye T, Vannberg FO, Wong SH, Owusu-Dabo E, Osei I, Gyapong J et al. Genome-wide association analyses identifies a susceptibility locus for tuberculosis on chromosome 18q11.2. Nat Genet 2010; 42: 739–741.
Chanock SJ, Manolio T, Boehnke M, Boerwinkle E, Hunter DJ, Thomas G et al. Replicating genotype-phenotype associations. Nature 2007; 447: 655–660.
Shah JA, Vary JC, Chau TT, Bang ND, Yen NT, Farrar JJ et al. Human TOLLIP regulates TLR2 and TLR4 signaling and its polymorphisms are associated with susceptibility to tuberculosis. J Immunol 2012; 189: 1737–1746.
Seya T, Oshiumi H, Sasai M, Akazawa T, Matsumoto M . TICAM-1 and TICAM-2: toll-like receptor adapters that participate in induction of type 1 interferons. Int J Biochem Cell Biol 2005; 37: 524–529.
Matsumiya M, Stylianou E, Griffiths K, Lang Z, Meyer J, Harris SA et al. Roles for Treg expansion and HMGB1 signaling through the TLR1-2-6 axis in determining the magnitude of the antigen-specific immune response to MVA85A. PLoS ONE 2013; 8: e67922.
Naslednikova IO, Urazova OI, Voronkova OV, Strelis AK, Novitsky VV, Nikulina EL et al. Allelic polymorphism of cytokine genes during pulmonary tuberculosis. Bull Exp Biol Med 2009; 148: 175–180.
Abhimanyu, Mangangcha IR, Jha P, Arora K, Mukerji M, Banavaliker JN et al. Differential serum cytokine levels are associated with cytokine gene polymorphisms in north Indians with active pulmonary tuberculosis. Infect Genet Evol 2011; 11: 1015–1022.
Li B, Leal SM . Methods for detecting associations with rare variants for common diseases: application to analysis of sequence data. Am J Hum Genet 2008; 83: 311–321.
Lewinsohn DA, Zalwango S, Stein CM, Mayanja-Kizza H, Okwera A, Boom WH et al. Whole blood interferon-gamma responses to mycobacterium tuberculosis antigens in young household contacts of persons with tuberculosis in Uganda. PLoS ONE 2008; 3: e3407.
Gross O, Thomas CJ, Guarda G, Tschopp J . The inflammasome: an integrated view. Immunol Rev 2011; 243: 136–151.
Novick D, Kim S, Kaplanski G, Dinarello CA . Interleukin-18, more than a Th1 cytokine. Semin Immunol 2013; 25: 439–448.
Schneider BE, Korbel D, Hagens K, Koch M, Raupach B, Enders J et al. A role for IL-18 in protective immunity against Mycobacterium tuberculosis. Eur J Immunol 2010; 40: 396–405.
Robinson CM, Jung JY, Interferon-gamma Nau GJ . tumor necrosis factor, and interleukin-18 cooperate to control growth of Mycobacterium tuberculosis in human macrophages. Cytokine 2012; 60: 233–241.
Li DD, Jia LQ, Guo SJ, Shen YC, Wen FQ . Interleukin-18 promoter gene -607C/A polymorphism and tuberculosis risk: a meta-analysis. Chin Med J (Engl) 2013; 126: 3360–3363.
Zhang Y, Jiang T, Yang X, Xue Y, Wang C, Liu J et al. Toll-like receptor -1, -2, and -6 polymorphisms and pulmonary tuberculosis susceptibility: a systematic review and meta-analysis. PLoS ONE 2013; 8: e63357.
Shey MS, Randhawa AK, Bowmaker M, Smith E, Scriba TJ, de Kock M et al. Single nucleotide polymorphisms in toll-like receptor 6 are associated with altered lipopeptide- and mycobacteria-induced interleukin-6 secretion. Genes Immun 2010; 11: 561–572.
Randhawa AK, Shey MS, Keyser A, Peixoto B, Wells RD, de Kock M et al. Association of human TLR1 and TLR6 deficiency with altered immune responses to BCG vaccination in South African infants. PLoS Pathog 2011; 7: e1002174.
Pickrell JK, Coop G, Novembre J, Kudaravalli S, Li JZ, Absher D et al. Signals of recent positive selection in a worldwide sample of human populations. Genome Res 2009; 19: 826–837.
Baker AR, Qiu F, Randhawa AK, Horne DJ, Adams MD, Shey M et al. Genetic variation in TLR genes in Ugandan and South African populations and comparison with HapMap data. PLoS ONE 2012; 7: e47597.
Kleinnijenhuis J, Joosten LA, van de Veerdonk FL, Savage N, van Crevel R, Kullberg BJ et al. Transcriptional and inflammasome-mediated pathways for the induction of IL-1beta production by Mycobacterium tuberculosis. Eur J Immunol 2009; 39: 1914–1922.
Zhao M, Jiang F, Zhang W, Li F, Wei L, Liu J et al. A novel single nucleotide polymorphism within the NOD2 gene is associated with pulmonary tuberculosis in the Chinese Han, Uygur and Kazak populations. BMC Infect Dis 2012; 12: 91.
Brooks MN, Rajaram MV, Azad AK, Amer AO, Valdivia-Arenas MA, Park JH et al. NOD2 controls the nature of the inflammatory response and subsequent fate of Mycobacterium tuberculosis and M. bovis BCG in human macrophages. Cell Microbiol 2011; 13: 402–418.
Ferwerda G, Girardin SE, Kullberg BJ, Le BL, de Jong DJ, Langenberg DM et al. NOD2 and toll-like receptors are nonredundant recognition systems of Mycobacterium tuberculosis. PLoS Pathog 2005; 1: 279–285.
Cobat A, Gallant CJ, Simkin L, Black GF, Stanley K, Hughes J et al. Two loci control tuberculin skin test reactivity in an area hyperendemic for tuberculosis. J Exp Med 2009; 206: 2583–2591.
Stein CM, Hall NB, Malone LL, Mupere E . The household contact study design for genetic epidemiological studies of infectious diseases. Front Genet 2013; 4: 61.
Li HT, Zhang TT, Zhou YQ, Huang QH, Huang J . SLC11A1 (formerly NRAMP1) gene polymorphisms and tuberculosis susceptibility: a meta-analysis. Int J Tuberc Lung Dis 2006; 10: 3–12.
Li X, Yang Y, Zhou F, Zhang Y, Lu H, Jin Q et al. SLC11A1 (NRAMP1) polymorphisms and tuberculosis susceptibility: updated systematic review and meta-analysis. PLoS ONE 2011; 6: e15831.
Meilang Q, Zhang Y, Zhang J, Zhao Y, Tian C, Huang J et al. Polymorphisms in the SLC11A1 gene and tuberculosis risk: a meta-analysis update. Int J Tuberc Lung Dis 2012; 16: 437–446.
Velez DR, Hulme WF, Myers JL, Stryjewski ME, Abbate E, Estevan R et al. Association of SLC11A1 with tuberculosis and interactions with NOS2A and TLR2 in African-Americans and Caucasians. Int J Tuberc Lung Dis 2009; 13: 1068–1076.
Guwattude D, Nakakeeto M, Jones-Lopez E, Maganda A, Chiunda A, Mugerwa R et al. Tuberculosis in household contacts of infectious cases in Kampala, Uganda. Am J Epidemiol 2003; 158: 887–898.
Jaganath D, Zalwango S, Okware B, Nsereko M, Kisingo H, Malone L et al. Contact investigation for active tuberculosis among child contacts in Uganda. Clin Infect Dis 2013; 57: 1685–1692.
Targeted tuberculin testing and treatment of latent tuberculosis infection. This official statement of the American Thoracic Society was adopted by the ATS Board of Directors, July 1999. This is a Joint Statement of the American Thoracic Society (ATS) and the Centers for Disease Control and Prevention (CDC). This statement was endorsed by the Council of the Infectious Diseases Society of America. (IDSA), September 1999, and the sections of this statement. Am J Respir Crit Care Med 2000; 161: S221–S247.
Statistical analysis for genetic epidemiology [computer program]. Version 6.3. Case Western Reserve University: Cleveland, OH, USA, 2012.
Li J, Ji L . Adjusting multiple testing in multilocus analyses using the eigenvalues of a correlation matrix. Heredity (Edinb) 2005; 95: 221–227.
Stein CM, Zalwango S, Chiunda AB, Millard C, Leontiev DV, Horvath AL et al. Linkage and association analysis of candidate genes for TB and TNFalpha cytokine expression: evidence for association with IFNGR1, IL-10, and TNF receptor 1 genes. Hum Genet 2007; 121: 663–673.
Acknowledgements
We acknowledge the invaluable contributions made by Dr Christopher Whalen, Dr Sarah Zalwango, Dr Lorna Nshuti, Dr Roy Mugerwa, Dr Deo Mulindwa, Allan Chiunda, Bonnie Thiel, Mark Breda, Dennis Dobbs, Hussein Kisingo, Mary Rutaro, Albert Muganda, Richard Bamuhimbisa, Yusuf Mulumba, Deborah Nsamba, Barbara Kyeyune, Faith Kintu, Gladys Mpalanyi, Janet Mukose, Grace Tumusiime, Pierre Peters, Dr Alphonse Okwera, Keith Chervenak, Denise Johnson, Karen Morgan, Alfred Etwom, Micheal Angel Mugerwa, Lisa Kucharski and Dr Feiyou Qiu. We thank Dr Francis Adatu Engwau, former Head of the Uganda National Tuberculosis and Leprosy Program, for his support of this project. We also thank the staff at the National Tuberculosis Treatment Centre, Mulago Hospital, the Ugandan National Tuberculosis and Leprosy Program and the Uganda Tuberculosis Investigation Bacteriological Unit, Wandegeya, for their contributions. This study would not be possible without the generous participation of the Ugandan patients and families. Funding for this work was provided by the Tuberculosis Research Unit (grants N01-AI95383 and HHSN266200700022C/N01-AI70022 from the NIAID), National Institute of Allergy and Infectious Disease grant K08AI083739 and National Heart Lung and Blood Institutes (NHLBI) grants R01HL096811, R01HL10566113 and T32HL007567.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on Genes and Immunity website
Supplementary information
Rights and permissions
About this article
Cite this article
Hall, N., Igo, R., Malone, L. et al. Polymorphisms in TICAM2 and IL1B are associated with TB. Genes Immun 16, 127–133 (2015). https://doi.org/10.1038/gene.2014.77
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gene.2014.77
This article is cited by
-
Low transcriptomic of PTPRCv1 and CD3E is an independent predictor of mortality in HIV and tuberculosis co-infected patient
Scientific Reports (2022)
-
The influence of single nucleotide polymorphisms of NOD2 or CD14 on the risk of Mycobacterium tuberculosis diseases: a systematic review
Systematic Reviews (2021)
-
Inflammasome genetics and complex diseases: a comprehensive review
European Journal of Human Genetics (2020)
-
Towards new TB vaccines
Seminars in Immunopathology (2020)
-
Fine-mapping analysis of a chromosome 2 region linked to resistance to Mycobacterium tuberculosis infection in Uganda reveals potential regulatory variants
Genes & Immunity (2019)