Abstract
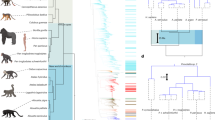
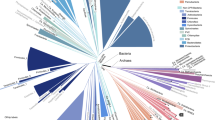
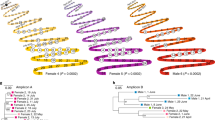
The CD209 gene family that encodes C-type lectins in primates includes CD209 (DC-SIGN), CD209L (L-SIGN) and CD209L2. Understanding the evolution of these genes can help understand the duplication events generating this family, the process leading to the repeated neck region and identify protein domains under selective pressure. We compiled sequences from 14 primates representing 40 million years of evolution and from three non-primate mammal species. Phylogenetic analyses used Bayesian inference, and nucleotide substitutional patterns were assessed by codon-based maximum likelihood. Analyses suggest that CD209 genes emerged from a first duplication event in the common ancestor of anthropoids, yielding CD209L2 and an ancestral CD209 gene, which, in turn, duplicated in the common Old World primate ancestor, giving rise to CD209L and CD209. KA/KS values averaged over the entire tree were 0.43 (CD209), 0.52 (CD209L) and 0.35 (CD209L2), consistent with overall signatures of purifying selection. We also assessed the Toll-like receptor (TLR) gene family, which shares with CD209 genes a common profile of evolutionary constraint. The general feature of purifying selection of CD209 genes, despite an apparent redundancy (gene absence and gene loss), may reflect the need to faithfully recognize a multiplicity of pathogen motifs, commensals and a number of self-antigens.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Bashirova AA, Wu L, Cheng J, Martin TD, Martin MP, Benveniste RE et al. Novel member of the CD209 (DC-SIGN) gene family in primates. J Virol 2003; 77: 217–227.
Koppel EA, van Gisbergen KP, Geijtenbeek TB, van Kooyk Y . Distinct functions of DC-SIGN and its homologues L-SIGN (DC-SIGNR) and mSIGNR1 in pathogen recognition and immune regulation. Cell Microbiol 2005; 7: 157–165.
Wu L, Kewalramani VN . Dendritic-cell interactions with HIV: infection and viral dissemination. Nat Rev Immunol 2006; 6: 859–868.
Figdor CG, van KY, Adema GJ . C-type lectin receptors on dendritic cells and Langerhans cells. Nat Rev Immunol 2002; 2: 77–84.
Feinberg H, Guo Y, Mitchell DA, Drickamer K, Weis WI . Extended neck regions stabilize tetramers of the receptors DC-SIGN and DC-SIGNR. J Biol Chem 2005; 280: 1327–1335.
Yang Z . The power of phylogenetic comparison in revealing protein function. Proc Natl Acad Sci USA 2005; 102: 3179–3180.
Nielsen R, Hellmann I, Hubisz M, Bustamante C, Clark AG . Recent and ongoing selection in the human genome. Nat Rev Genet 2007; 8: 857–868.
Goodman M . The genomic record of Humankind's evolutionary roots. Am J Hum Genet 1999; 64: 31–39.
Yang Z, Wong WS, Nielsen R . Bayes empirical bayes inference of amino acid sites under positive selection. Mol Biol Evol 2005; 22: 1107–1118.
Snyder GA, Colonna M, Sun PD . The structure of DC-SIGNR with a portion of its repeat domain lends insights to modeling of the receptor tetramer. J Mol Biol 2005; 347: 979–989.
Barreiro LB, Patin E, Neyrolles O, Cann HM, Gicquel B, Quintana-Murci L . The heritage of pathogen pressures and ancient demography in the human innate-immunity CD209/CD209L region. Am J Hum Genet 2005; 77: 869–886.
Wagner A . Rapid detection of positive selection in genes and genomes through variation clusters. Genetics 2007; 176: 2451–2463.
Ortiz M, Bleiber G, Martinez R, Kaessmann H, Telenti A . Patterns of evolution of host proteins involved in retroviral pathogenesis. Retrovirology 2006; 3: 11.
Sawyer SL, Emerman M, Malik HS . Ancient adaptive evolution of the primate antiviral DNA-editing enzyme APOBEC3G. PLoS Biol 2004; 2: E275.
Sawyer SL, Wu LI, Emerman M, Malik HS . Positive selection of primate TRIM5{alpha} identifies a critical species-specific retroviral restriction domain. Proc Natl Acad Sci USA 2005; 102: 2832–2837.
Goldschmidt V, Ciuffi A, Ortiz M, Brawand D, Munoz M, Kaessmann H et al. Antiretroviral activity of ancestral TRIM5alpha. J Virol 2008; 82: 2089–2096.
Barreiro LB, Quintana-Murci L . DC-SIGNR neck-region polymorphisms and HIV-1 susceptibility: From population stratification to a possible advantage of the 7/5 heterozygous genotype. J Infect Dis 2006; 194: 1184–1185.
Wichukchinda N, Kitamura Y, Rojanawiwat A, Nakayama EE, Song H, Pathipvanich P et al. The polymorphisms in DC-SIGNR affect susceptibility to HIV type 1 infection. AIDS Res Hum Retroviruses 2007; 23: 686–692.
Martin MP, Lederman MM, Hutcheson HB, Goedert JJ, Nelson GW, van KY et al. Association of DC-SIGN promoter polymorphism with increased risk for parenteral, but not mucosal, acquisition of human immunodeficiency virus type 1 infection. J Virol 2004; 78: 14053–14056.
Liu H, Hwangbo Y, Holte S, Lee J, Wang C, Kaupp N et al. Analysis of genetic polymorphisms in CCR5, CCR2, stromal cell-derived factor-1, RANTES, and dendritic cell-specific intercellular adhesion molecule-3-grabbing nonintegrin in seronegative individuals repeatedly exposed to HIV-1. J Infect Dis 2004; 190: 1055–1058.
Gramberg T, Zhu T, Chaipan C, Marzi A, Liu H, Wegele A et al. Impact of polymorphisms in the DC-SIGNR neck domain on the interaction with pathogens. Virology 2006; 347: 354–363.
Olesen R, Wejse C, Velez DR, Bisseye C, Sodemann M, Aaby P et al. DC-SIGN (CD209), pentraxin 3 and vitamin D receptor gene variants associate with pulmonary tuberculosis risk in West Africans. Genes Immun 2007; 8: 456–467.
Vannberg FO, Chapman SJ, Khor CC, Tosh K, Floyd S, Jackson-Sillah D et al. CD209 genetic polymorphism and tuberculosis disease. PLoS ONE 2008; 3: e1388.
Lynch M, Conery JS . The evolutionary fate and consequences of duplicate genes. Science 2000; 290: 1151–1155.
Medzhitov R . Toll-like receptors and innate immunity. Nat Rev Immunol 2001; 1: 135–145.
Edgar RC . MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 2004; 32: 1792–1797.
Rice P, Longden I, Bleasby A . EMBOSS: the European Molecular Biology Open Software Suite. Trends Genet 2000; 16: 276–277.
Ronquist F, Huelsenbeck JP . MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 2003; 19: 1572–1574.
Yang Z . PAML: a program package for phylogenetic analysis by maximum likelihood. Comput Appl Biosci 1997; 13: 555–556.
Yang Z, Bielawski JP . Statistical methods for detecting molecular adaptation. Trends Ecol Evol 2000; 15: 496–503.
Yang Z, Nielsen R, Goldman N, Pedersen AM . Codon-substitution models for heterogeneous selection pressure at amino acid sites. Genetics 2000; 155: 431–449.
Yang Z . Likelihood ratio tests for detecting positive selection and application to primate lysozyme evolution. Mol Biol Evol 1998; 15: 568–573.
Acknowledgements
We thank Keith Mansfield and Kuei-Chin Lin from the New England Primate Center, and Charles Buillard and Eugène Chabloz from the Zoo of Servion for materials. This work was funded by the Swiss National Science Foundation and a grant for interdisciplinary research from the Faculty of Biology and Medicine of the University of Lausanne. This project has been funded in whole or in part with federal funds from the National Cancer Institute, National Institutes of Health, under contract N01-CO-12400. The content of this publication does not necessarily reflect the views or policies of the Department of Health and Human Services, nor does mention of trade names, commercial products or organizations imply endorsement by the US Government. This research was supported in part by the Intramural Research Program of the NIH, National Cancer Institute, Center for Cancer Research.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on Genes and Immunity website (http://www.nature.com/gene)
Supplementary information
Rights and permissions
About this article
Cite this article
Ortiz, M., Kaessmann, H., Zhang, K. et al. The evolutionary history of the CD209 (DC-SIGN) family in humans and non-human primates. Genes Immun 9, 483–492 (2008). https://doi.org/10.1038/gene.2008.40
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gene.2008.40
Keywords
This article is cited by
-
Evolutionary history of type II transmembrane serine proteases involved in viral priming
Human Genetics (2022)
-
A monoclonal antibody to Siglec-8 suppresses non-allergic airway inflammation and inhibits IgE-independent mast cell activation
Mucosal Immunology (2021)
-
Human gene polymorphisms and their possible impact on the clinical outcome of SARS-CoV-2 infection
Archives of Virology (2021)
-
Akt isoform-specific effects on thyroid cancer development and progression in a murine thyroid cancer model
Scientific Reports (2020)
-
The presence of CD209 expressing dendritic cells correlates with biofilm positivity in chronic rhinosinusitis with nasal polyposis
European Archives of Oto-Rhino-Laryngology (2013)