Abstract
When a ribosome complex is stalled during the translation elongation process in eukaryotes, the mono-ubiquitination of Rps3 has recently been shown to be critical to ribosome quality control. We have discovered that the regulatory role of Rps3 mono-ubiquitination is controlled by a deubiquitinase. We also showed that an autophagic signal appears to be coupled to the mono-ubiquitination of Rps3p through the entrance of Ubp3p into the autophagosome in yeasts. The mono-ubiquitination of the Rps3 protein is tightly modulated by reciprocal action between the Hel2p E3 ligase and the Ubp3p deubiquitinase in yeasts and the reciprocal action between the RNF123 E3 ligase and the USP10 deubiquitinase in mammalian cells. We also found that the Ubp3p/USP10 deubiquitinases critically modulate Hel2p/RNF123-mediated Rps3p mono-ubiquitination. In addition, we found that Hel2p/RNF123 and Ubp3p/USP10 appeared to be differently localized in the ribosome complex after ultraviolet irradiation. Together, our results support a model in which coordinated ubiquitination and deubiquitination activities can finely balance the level of regulatory Rps3p mono-ubiquitination in ribosome-associated quality control and autophagy processes.
Similar content being viewed by others
Introduction
The ribosome is macromolecular machinery that is responsible for protein synthesis.1 However, several ribosomal proteins have been found to have extra-ribosomal functions, including stress response,2, 3 the induction of apoptosis4 and repairing both mitochondrial and nuclear DNA damage.5, 6 For example, it has been reported that RPS3 has endonuclease activity toward several damaged sites that are involved in apoptosis.7, 8, 9, 10 Drosophila rpS3 also has DNA deoxyribophosphodiesterase activity11 and DNA glycosylase activity,12 in addition to processing repair activities on 8-oxoguanine and AP sites.13 Both bacterial and human RPS3 are part of the basic downstream tunnel that leads to mRNA.14 The structure formed by proteins RPS3, RPS4, and RPS5 has a role in helicase processivity.15 It has been reported that human RPS3 can act as a special component in NF-κB complexes and regulates selective gene expression.16 Therefore, the RPS3 protein seems to be capable of residing outside the ribosome and shuttles between the cytoplasm and nucleus, functioning in both the ribosome complex and the nucleus.
Post-translational modification by ubiquitin and ubiquitin-like proteins has been revealed to be a means to modulate the stability, localization, or function of target proteins.17, 18 Previous studies have shown that ribosome function is regulated by the post-translational modification (such as phosphorylation, sumoylation, neddylation, and ubiquitination) of several ribosomal proteins.19, 20 For example, upon viral infection, ribosomal proteins such as RPS6, RPS2, and RPL30 are phosphorylated.21 The RPS3 is also a target of post-translational modification in the ribosome. The yeast Rps3p is phosphorylated by Hrr25p protein kinase, and this modification appears to play an essential role in ribosome maturation.22, 23, 24 Moreover, the ubiquitination of several 40S ribosomal proteins (RPS2, RPS3, and RPS20) regulates ribosome-associated quality control (RQC) that induces and degrades the poly-ubiquitination of partially synthesized polypeptides.25
RQC is responsible for the maintenance of cellular protein homeostasis.26, 27 The RQC pathway has been known to be implemented when the interruption of translational elongation stalls ribosomes. Three stages have been unraveled in the process of RQC. First, the GTP-Hbs1p-Dom34p/PELOTA complex must use an uncharacterized mechanism to discriminate between a stalling ribosome and a pausing ribosome. Second, Hbs1p hydrolyzes GTP, and Dom34p/PELOTA is consequently accommodated in the ribosomal A site, resulting in the recruitment of Rli1p/ABCE1. Finally, a stalled ribosome complex will split into two subunits: the 40S small subunit and the 60S large subunit. While mRNA surveillance mechanisms such as no-go decay degrades the corresponding mRNA on the 40S subunit, Ltn1p/LISTERIN and Rqc2p/NEMF will facilitate the ubiquitination of partially synthesized polypeptides that are tethered on the 60S large subunit.28, 29 Subsequently, the poly-ubiquitinated polypeptide is transferred to the proteasome by the Rqc1p/TCF25 and Cdc48p complex. In addition to the abovementioned RQC factors, Asc1p/RACK1 and Hel2p/RNF123 have been recently reported to be involved in the early stage of RQC pathway.30
Materials and methods
Yeast strains and culture condition
All Saccharomyces cerevisiae strains and plasmid constructs used in this study are listed in Table 1. The yeast cells were grown in yeast extract-peptone-dextrose (YPD) medium (1% yeast extract, 2% peptone and 2% dextrose) and a synthetic complement medium (2% glucose, 0.67% yeast nitrogen base w/o amino acids (BD Pharmingen, Franklin Lakes, NJ, USA)), and with 15 or fewer proper amino acids). The cells were grown to early log phase in each media and were treated with each drug for 1 h. For ultraviolet (UV) irradiation, BY4741 cells were grown at 30 °C in YPD medium to early log phase, washed once with sterile distilled water, and resuspended in 2 ml of cold 2% glucose on a plate. The UV radiation was delivered using a CL-1000 UV Crosslinker set at 254 nm. The cells were irradiated with 200 J m−2 and then added to one-tenth volume of a stock solution containing 10% yeast extract and 20% peptone, followed by incubation in the dark at 30 °C for 1 h.
Antibodies and reagents
Antibodies against Pgk1p, ubiquitin, and HA were obtained from Santa Cruz Biotechnology (Dallas, TX, USA). The antibody against FLAG was purchased from Cell Signaling Technology (Danvers, MA, USA). The antibody against eIF2α~p was purchased from Abcam (Cambridge, MA, USA). Anti-Rps3p, anti-RPS3, anti-monoubiquitinated Rps3p antibodies and Rps3 siRNAs were obtained from HAEL Lab (seoul, Korea). Horseradish peroxidase (HRP) -conjugated goat anti-rabbit and goat anti-mouse secondary antibodies were purchased from Jackson ImmunoResearch (West Grove, PA, USA). Rapamycin, emetine, cycloheximide, and 3-AT were purchased from Sigma-Aldrich (St Louis, MO, USA). Blasticidin S, anisomycin, and hygromycin B were purchased from Merck (Darmstadt, Germany).
Western blot and immunoprecipitation
The harvested yeast cells were washed with PBS and lysed on ice in TAP lysis buffer (10 mM Tris-Cl pH 8.0, 150 mM NaCl, and 0.1% NP-40) that contained protease inhibitors (1 mM PMSF, 1 mM Aprotinin, 5 mM Leupeptin, and 1 mM Pepstatin A). Glass beads were then added, and the mixture was vortexed four times with a bead beater (MP Biomedicals, Santa Ana, CA, USA) for 1 min. Cell lysates were obtained after centrifugation at 10,000 g for 5 min. Equal amounts of lysates were subjected to western blotting with each antibody. For co-immunoprecipitation, 1.2 mg of protein lysate was mixed with 2.5 μg of Rps3p antibodies on a rotator at 4 °C overnight. Subsequently, 20 μl of protein A-agarose (Roche, Basel, Switzerland) was added and the solution was incubated on a rotator at 4 °C for 4 h. The immunoprecipitants were resolved on 10% SDS–polyacrylamide gel electrophoresis for western blotting.
Yeast transformation
Overnight-cultured yeast cells that were grown at 30 °C in YPD were reseeded into 20 ml of fresh YPD medium at OD600=0.2 and incubated at 30 °C for 4 h (until the OD600 reached 0.8–1.0). The cells were harvested by centrifugation at 13,000 r.p.m. for 5 min and then washed twice. The first washing buffer was 1 × TE (10 mM Tris-Cl and 1 mM EDTA). The second washing buffer was 0.1 M LiAc in 1 × TE. The cells were resuspended in 400 μl 0.1 M LiAc/1 × TE, and 100 μl of the resuspended cells were used for each transformation. Then, 350 μl 40% PEG/0.1 M LiAc/1 × TE was added to each transformation, followed by the addition of 5 μl of 10 mg ml−1 carrier DNA (boiled for 15 min and cooled directly on ice) and the DNA to be transformed. The solution was well mixed by vortexing. The mixture was then incubated at 30 °C for 30 min and mixed with 45 μl dimethylsulphoxide. This mixture was then heat-shocked for 15 min at 42 °C and cooled on ice for 3 min. The cells were pelleted by spin-down, and the supernatant was discarded. These cells were then resuspended in 100 μl 1 × TE, spread onto appropriate SC drop-out medium plates and incubated at 30 °C for at least 2 days.
Ribosome pelleting
Cultured yeast cells were harvested by centrifugation at 13,000 r.p.m. for 5 min. The harvested yeast cells were washed with PBS and lysed on ice in 1 ml of modified TAP lysis buffer (10 mM Tris-Cl pH 8.0, 150 mM NaCl, 0.1% NP-40, 30 mM MgCl2, 50 μg ml−1 cycloheximide, 40 units per ml rRNasin and 1.5 mM dithiothreitol) that contained protease inhibitors (1 mM PMSF, 1 mM Aprotinin, 5 mM Leupeptin, and 1 mM Pepstatin A). The cells were added to glass beads, and the mixture was vortexed four times with a bead beater for 1 min. The cell lysates were obtained after centrifugation at 10,000 g for 10 min. Lysate (2 mg) was added to 11 ml of 30% sucrose that contained 50 mM Tris-acetate, pH 7.0, 50 mM NH4Cl, and 12 mM MgCl2, followed by centrifugation (SW41Ti of Beckman) at 32,000 r.p.m. for 210 min at 4 °C. The resulting pellet was resuspended and subjected to 10% SDS–polyacrylamide gel electrophoresis for western blotting.
In vitro ubiquitination and deubiquitination assays
Five hundred nanograms of human ubiquitin-activating enzyme E1 (Enzo), 250 ng of UBCH5b (Enzo), 3 μg of His6-Hel2p, 2 μg of His6-Rps3p, 3 μg of His6-Ubp3p, and 1.5 μg of His6-ubiquitin were added to a 50 μl reaction containing 50 mM Tris-HCl (pH 7.5), 150 mM NaCl, 2 mM MgCl2, 1 mM dithiothreitol, and 5 mM ATP. The reaction mixture was incubated at 30 °C overnight. The reaction was stopped by adding 50 μl of 2 × SB. The proteins were subjected to immunoblotting using anti-Rps3p.
Spotting assay
Approximately 2 × 107 of cultured yeast cells (OD600=1.00) were harvested by centrifugation at 13,000 r.p.m. for 5 min. The harvested cells were spotted onto YPD plates after a series of 10-fold dilutions, followed by their incubation at 30 °C for 2 days.
Results
Mono-ubiquitination of the K212 residue on Rps3p is increased when protein synthesis is stopped by either translation inhibition or Tor kinase inhibition
Higgins et al.25 have previously demonstrated that the mono-ubiquitination of mammalian RPS3 is induced by translational inhibitors. Therefore, we further explored whether some stresses could induce the mono-ubiquitination of yeast Rps3p. In line with the previous study, treatments with translation-inhibiting drugs such as cycloheximide, emetine, blasticidin S, and anisomycin induced mono-ubiquitination of Rps3p (Figure 1a). The high doses of UVC irradiation (200 J m−2) that could evoke ribotoxic stress and inhibit peptidyltransferase activity31 also induced the mono-ubiquitination of Rps3p. Interestingly, rapamycin treatment and amino acid depletion conditions (such as tryptophan depletion, histidine depletion, amino acid N-source starvation, and treatment with 3-aminotriazole (3-AT)) also resulted in the mono-ubiquitination of Rps3p (Figure 1b). 3-AT is a competitive inhibitor of HIS3 that catalyzes histidine biosynthesis, while rapamycin is an inhibitor of Tor kinase. In contrast, nutrient-rich conditions (10-fold amino acid) did not induce the mono-ubiquitination of Rps3p. Thus, we concluded that Tor kinase inhibition and elongation inhibition could induce the mono-ubiquitination of Rps3p.
Ubiquitination of K212 on Rps3p is increased by starvation and translation inhibition. Each sample was harvested and lysed for western blotting. The proteins were confirmed by immunoblotting using an anti-Rps3p antibody. The anti-pgk1p antibody was used as a loading control. (a) The BY4741 cells that were grown to the stationary phase in YPD media were reseeded and grown to the early log phase. After cells were treated with each translation inhibition drug such as Cycloheximide (100 μg ml−1), Emetine (10 μM), Blasticidin S (25 μg ml−1), Anisomycin (100 μM) or exposed to UV (200 J m−2), they were incubated for 1 h. (b) The JS143-7D and BY4741 strains were grown to stationary phase in SC media or YPD media. The cells were reseeded into each appropriate media and grown to the early log phase. The media was changed to SC media that lacked Trp and His (tryptophan, histidine depletion), SC media that contained a lower (1/10) or higher concentration of amino acids (10 ×), nitrogen starvation media, SC-His media containing 80 mM 3-AT, or YPD media that contained 100 ng ml−1 rapamycin (Tor kinase inhibition). The incubation time was 1 h, after which the cells were harvested by centrifugation. (c) The lysates from the Rps3 or Rps3K212R cells that were exposed to UV stress were analyzed by western blotting with indicated antibodies. The anti-eIF2α~p antibody was used as a positive control for UV stress. (d) The total extracts from the Rps3 and Rps3K212R mutant cells that were exposed to UV stress (+) or not (−) were subjected to centrifugation in a 5~45% sucrose gradient and analyzed by UV absorbance at 254 nm. The fractions were collected and analyzed by immunoblotting with anti-Rps3p, anti-ubiquitinated Rps3p, and anti-Pgk1p antibodies.
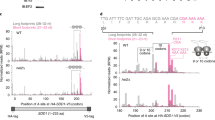
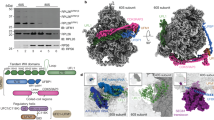
In mammalian cells, mass spectrometry analysis identified that lysine 214 at RPS3 was a major mono-ubiquitination site under UV irradiation (Supplementary Fig. S1A). Immunoprecipitation using the anti-RPS3 antibody, followed by western blotting with an anti-ubiquitin antibody, confirmed this modification (Supplementary Fig. S1B). To test whether the mono-ubiquitination residue of Rps3p was at lysine 212 (its corresponding residue is lysine 214 in human RPS3), we generated a non-ubiquitable Rps3p mutant whose lysine 212 residue was substituted with an arginine from its endogenous locus. Different from the wild-type Rps3p, the Rps3pK212R mutant failed to become mono-ubiquitinated under UV irradiation (Figure 1c). In the absence of UV irradiation, the Rps3pK212R mutant was efficiently assembled in the ribosome complex, similar to the wild type (Figure 1d, upper panel). Wild-type Rps3p mono-ubiquitination was detected mostly in monosomes, not in the free subunit (Figure 1d, lower panel). No significant difference in the protein expression level between the wild type and the Rps3pK212R mutant after treatment with cycloheximide was found, indicating protein stability (Supplementary Fig. S2). Collectively, these results suggest that the mono-ubiquitination of the lysine 212 residue in yeast Rps3p can be increased by inhibiting Tor kinase or inhibiting translation elongation.
Hel2p is an E3 ligase of ribosomal protein S3
In yeasts, ubiquitin conjugation is catalyzed by an E1 activating enzyme, 11 E2 conjugating enzymes, and 60~100 substrate-specific E3 ligases.32 To identify the specific E3 ubiquitin ligase that is involved in the mono-ubiquitination of Rps3p, we performed western blotting analyses for non-essential E3 ligase gene deletion strains under the conditions of rapamycin treatment (Supplementary Fig. S3). The E3 ligases depicted in Supplementary Fig. S3 are localized at the ribosome and the ER. They are involved in protein quality control and amino acid sensing. The level of Rps3p mono-ubiquitination significantly decreased only in the hel2Δ strain (Supplementary Fig. S3F and Figure 2a). In addition, a decrement in the Rps3p mono-ubiquitination was not observed, even in strains in which the ribosome-associated E3 ligases such as Rkr1p and Not4p (Supplementary Figures S3F and H) were deleted. Moreover, we expressed a catalytically inactive Hel2p mutant (Hel2C79A, H81A, C87A) by mutating residues in its RING domain in the hel2Δ strain33 to further corroborate its function as an E3 ligase in Rps3p mono-ubiquitination. We found that the knockout of Hel2p was directly responsible for Rps3p mono-ubiquitination in vivo because the expression of FLAG-Hel2p was restored using a GAL promoter (Figure 2b, lanes 3 and 4). We also found that the Hel2p overexpression was enough to induce Rps3p mono-ubiquitination without UV irradiation. Moreover, the Hel2p mutant could not efficiently ubiquitinate Rps3p, unlike the Hel2p wild type (Figure 2b, lane 5 and 6). To determine whether the Hel2p was associated with the ribosome complex that contained Rps3p or Rps3pK212R, we generated strains by reintroducing FLAG-Hel2p and its promoter into the hel2Δ strains that express Rps3p or Rps3pK212R. After subsequent ribosome pelleting, we found that the association of Hel2p in the ribosome complex was increased under UV irradiation with Rps3p, but not with Rps3pK212R (Figure 2c). These findings suggest that the mono-ubiquitination of Rps3p is catalyzed by Hel2p, which is a ribosome-associated E3 ligase.
Hel2p is an E3 ligase of ribosomal protein s3. (a) The wild-type (wt) and hel2Δ strains were grown to early log phase in YPD media, exposed to UV stress (200 J m−2) and incubated for 1 h. The whole cell lysates were analyzed by immunoblot with anti-Rps3p, ubiquitinated Rps3p, eIF2α~p, and Pgk1 antibodies. (b) The BY4741 hel2Δ strain was transformed with plasmid encoding FLAG (control), FLAG-Hel2p (WT-Hel2), or FLAG-Hel2pC79A, H81A, C87A (mutation of the RING domain). The cells expressing FLAG, FLAG-Hel2p, or FLAG-Hel2pC79A, H81A, C87A were grown to the stationary phase in SD (U−) media. The cells were reseeded in SRG (U−) media and then grown to the early log phase. The whole cells were analyzed by immunoblotting with the indicated antibodies. (c) The BY4741 hel2Δ strains harboring the vectors expressing FLAG or Hel2p-FLAG were grown to the early log phase in SC (H−) media. The cells grown to early log phase were exposed to UV stress (+) or not (−) and incubated for 3 min. The cell lysates (left) or the ribosomes that were purified by ribosome pelleting (right) were confirmed by western blotting with the indicated antibodies. The incubation time after UV stress was 1 h, except in the case of c.
Ubp3p is a deubiquitination enzyme of ribosomal protein S3
Previous studies have shown that Ubp3p, an ubiquitin protease, can deubiquitinate the ubiquitination of Rpl25p, resulting in the selective degradation of the 60S ribosomal large subunit.34 If Ubp3p behaves as a specific ubiquitin protease for Rps3p mono-ubiquitination, the deletion of Ubp3p might affect the ubiquitination of Rps3p. To test this possibility, we performed an immunoblot analysis for the ubp3Δ strain. As shown in Figure 3a, with or without UV irradiation, the level of Rps3p mono-ubiquitination in the ubp3Δ strain was strikingly elevated compared to that of the wild type. On the other hand, Rps3p mono-ubiquitination was not increased under UV stress in the His6-Ubp3p-overexpressing BY4741 strain compared to that in the His6-overexpressing BY4741 strain (Figure 3b, lane 4). To confirm whether the function of Ubp3p required its catalytic activity under UV stress, we created a point mutant whose Cys469 and His861 were substituted with Ala to abolish its catalytic activity.35 Compared to the His6-Ubp3p-overexpressing strain, the level of Rps3p mono-ubiquitination induced by UV irradiation was restored in the Ubp3p mutant (Ubp3C469A, H861A) (Figure 3b, lane 6). Next, we attempted to clarify whether the Rps3p or Rps3pK212R in the ribosome complex might affect the association of Ubp3p. After reintroducing Ubp3p-HA and its promoter into the ubp3Δ strains that express Rps3p or Rps3pK212R, ribosome pelleting was performed. We found that the association of Ubp3p in the ribosome complex was not changed in the two strains, regardless of the presence of UV irradiation (Figure 3c, right panel). Additionally, we showed that the ubiquitination of Rps3pK212R was not caused by UV stress in the ubp3Δ strain (Figure 3d). Therefore, the association of Ubp3p with the ribosome complex appears to maintain a steady-state level irrespective of UV irradiation or the presence of a non-ubiquitinable mutation in Rps3p. Collectively, these results indicate that the mono-ubiquitination of Rps3p can be deubiquitinated by Ubp3p, a ribosome-associated deubiquitination enzyme.
Rps3p is deubiquitinated by Ubp3p. (a) The wild-type (WT) and ubp3Δ strains were grown to the early log phase in YPD media, exposed to UV stress (200 J m−2) and incubated for 1 h. (b) The BY4741 ubp3Δ strain was transformed with a plasmid expressing His6 (control), His6-Ubp3p (WT-Ubp3), or His6-Ubp3pC469A, H861A (catalytically inactive allele). The transformed cells were grown to the early log phase in SRG (U−) media for induction. Each cell was exposed to UV irradiation (+) or not (−). The whole cells were analyzed by immunoblotting with the indicated antibodies. (c) The indicated strains expressing Ubp3p-HA from its endogenous locus were grown to the early log phase in YPD media. The cells grown to the early log phase were exposed to UV stress (+) or not (−) and incubated for 3 min. The cell lysates (left) or the ribosomes that were purified by ribosome pelleting (right) were confirmed by western blotting with the indicated antibodies. (d) The indicated strain containing the UBP3 deletion was grown to the early log phase in YPD media. The proteins in the total extracts were analyzed by immunoblotting with the indicated antibodies. The incubation time after UV stress was 1 h, except in the case of c.
Ubiquitination of Rps3p is regulated by Hel2p and Ubp3p in vitro
Having determined that Hel2p and Ubp3p regulated the status of Rps3p mono-ubiquitination in vivo, we next tested whether they could also modulate the mono-ubiquitination of Rps3p in vitro. First, to find an adequate E2 conjugating enzyme, we performed an in vitro Rps3p ubiquitination assay with various recombinant human E2 conjugating enzymes. Among these E2 enzymes, UBCH5a, UBCH5b, and UBCH5c facilitated the mono-ubiquitination of Rps3p in vitro (Figure 4a). With another ubiquitin-conjugating enzyme, UBCH6, the mono-ubiquitination of Rps3p appeared to result in a different molecular weight. It is well-known that E2 conjugating enzymes (including UBCH5a, UBCH5b, UBCH5c, and UBCH6) are homologous to yeast Ubc4/5p.36 With the recent finding that Hel2p interacts with Ubc4p but not Ubc5p,33 we next examined the role of Ubc4p in the mono-ubiquitination of Rps3p. As shown in Figure 4b, the immunoblot analysis of the ubc4Δ strains showed that the mono-ubiquitination of Rsp3p disappeared after UV irradiation, suggesting that Ubc4p is a bona fide E2 conjugating enzyme for the mono-ubiquitination of Rps3p. In contrast, the decrease of Rps3p mono-ubiquitination did not occur in the UV-irradiated ubc5Δ strain (Supplementary Fig. S3E, lane 10). We also performed an in vitro deubiquitination assay on the mono-ubiquitinated Rps3p using recombinant E1, E2, ubiquitin, and Rps3p in combination with Hel2p and Ubp3p. As expected, the level of Rps3p mono-ubiquitination noticeably increased with the addition of the recombinant Hel2p (lane 2, Figure 4c). However, this level decreased with the addition of recombinant Ubp3p (lane 3, Figure 4c). These data collectively indicate that the mono-ubiquitination of Rps3p can be modulated by Ubc4p, Hel2p, and Ubp3p both in vivo and in vitro.
Ubiquitinated Rps3p is regulated byUbc4p, Hel2p and Ubp3p in vitro. (a) An in vitro ubiquitination assay was performed on purified His6-tagged E1 (human), Hel2p, Rps3p, Ubp3p, and ubiquitin. The indicated E2 conjugation enzyme was added into the mixtures. The mixtures were then analyzed by immunoblotting with anti-Rps3p and anti-ubiquitinated Rps3p antibodies. (b) The wild-type (wt) and ubc4Δ strains were grown to the early log phase in YPD media. The cells were exposed to UV stress (+) or not (−) and incubated for 1 h. The whole cell lysates were analyzed by immunoblotting with anti-Rps3p, anti-ubiquitinated Rps3p, anti-eIF2α~p, and anti-Pgk1p antibodies. (c) An in vitro ubiquitination assay was performed on purified His6-tagged E1 (human), UBCH5b, Hel2p, Rps3p, Ubp3p, and ubiquitin. The Hel2p was added (+) or not (−) into a solution that contained E1, UBCH5b, ubiquitin, and Rps3p (lanes 1 and 2). Ubp3p was added (+) or not (−) into a solution that contained E1, UBCH5b, ubiquitin, Hel2p, and Rps3p. The solutions were analyzed by immunoblotting with anti-Rps3p and anti-ubiquitinated Rps3p antibodies. The Rps3p and ubiquitinated Rps3p are indicated on the right.
Rsp3p mono-ubiquitination induced by rapamycin is mediated by the entrance of Ubp3p into the autophagosome
As mentioned above, we found that Tor kinase inhibition could also induce Rps3p mono-ubiquitination in yeasts. However, the inhibition of Tor kinase by rapamycin treatment and nutrient starvation only affects translation initiation, not translation elongation. We found that RPS3 mono-ubiquitination was not induced in mammalian cells by the inhibition of translation initiation such as that caused by treatment with rapamycin (Supplementary Fig. S4A) or NSC119889 (Supplementary Fig. S4B). Such discrepancies between yeasts and humans led us to question how the Rps3p mono-ubiquitination/de-ubiquitination is modulated under Tor inhibition conditions in yeasts. It has been previously reported that Ubp3p is responsible for the deubiquitination of the mono-ubiquitinated Rpl25p in the 60S large ribosomal subunit, resulting in the progression of ribophagy.34 Therefore, we speculated that the Rps3p mono-ubiquitination that was induced by rapamycin might be elicited by the entrance of Ubp3p into the autophagosome, resulting in the depletion of Ubp3p. To address this possibility, we first checked the levels of Rps3p mono-ubiquitination in the Ubp3p- and Bre5p-deleted strains. Ubp3p catalyzes deubiquitination, while Bre5p is a Ubp3-associated cofactor.37 In the ubp3Δ (Figure 5a) and bre5Δ (Figure 5b) strains, Rps3p mono-ubiquitination was significantly maintained until 24 h after rapamycin treatment, while it was significantly maintained for 3 h in BY4741 wild-type cells. To inhibit the formation of autophagic membranes, we generated two different deletion strains: atg7Δ and atg8Δ. Atg7p is an E1 enzyme that can activate ubiquitin in a similar manner to Atg8p to form autophagosome membranes.38 As expected, in atg8Δ (Figure 5c) and atg7Δ (Figure 5d) strains, the levels of Rps3p mono-ubiquitination after the rapamycin treatment were less than or similar to those of the BY4741 wild-type cells. Additionally, the ufd3Δ stains showed the disappearance of the Rps3p mono-ubiquitination that was induced by rapamycin (Supplementary Fig. S5). Because Ufd3p is a positive regulator of Ubp3p-Bre5p mediated ribophagy,39 the deletion of Ufd3p blocks the entrance of the Ubp3p-Bre5p complex into the autophagosome. To strengthen this hypothesis, we investigated the associations in the ribosome complex of Ubp3p by immunoprecipitating it with the Rps3p antibody under the conditions of rapamycin treatment. The Ubp3p association was dramatically decreased after rapamycin treatment (Figure 5e). More interestingly, the Ubp3p association with the ribosome was not changed by UV irradiation. Thus, we suggested that the Rps3p mono-ubiquitination induced by rapamycin is due to the entrance of Ubp3p into the autophagosome.
Ubp3p-dependent autophagy is indispensable for rapamycin-induced Rps3p mono-ubiquitination. The wild-type (WT), Δubp3 (a), Δbre5 (b), Δatg8 (c), and Δatg7 (d) deletion strains were grown to the early log phase in YPD media and then treated with rapamycin for the indicated time. The cells were analyzed by western blotting with anti-Rps3p, eIF2α~p, Pgk1p, and ubiquitinated Rps3p antibodies. (e) The strains with Ubp3p-HA that was reintroduced at its endogenous locus were grown to the early log phase in YPD media and exposed to UV stress or treated with rapamycin for the indicated time. The whole cell lysates were immunoprecipitated with the Rps3p antibody. The immunoprecipitates were analyzed by western blotting with the indicated antibodies. The arrow denotes mono-ubiquitinated Rps3p, while the asterisk denotes a non-specific band.
Human RPS3 mono-ubiquitination is regulated by both RNF123 and USP10
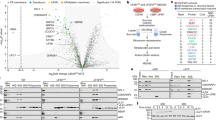
Next, we wondered whether the above observation might also be applicable to human cells. RPS3 mono-ubiquitination was induced by various translation elongation inhibitors, including UV irradiation (Figure 6a), but not genotoxic stress (Supplementary Fig. S6). UV irradiation is a ribotoxic stressor that can induce the cleavage of the 28S rRNA and inhibit peptidyltransferase activity, which results in the inhibition of translation elongation. The levels of RPS3 mono-ubiquitination peaked 1 to 4 h after UV-irradiation and then gradually decreased (Supplementary Fig. S7A). Since modified RPS3 was present in the cytoplasm (Supplementary Fig. S7B), we separated the free ribosomal proteins from the monosomes and polysomes in UV-irradiated cells by ultracentrifugation. The mono-ubiquitinated RPS3 was detected mostly in the monosomes, but not in the free subunit (Supplementary Fig. S7C). Furthermore, the mono-ubiquitination of RPS3 was independent of p53 (Supplementary Fig. S7D). In previous literature, although several specific lysine residues—K90, K214, K227, and K230—were reported to be mono-ubiquitination sites of human RPS3,25 we confirmed here that the K214 residue was a major mono-ubiquitination site of RPS3 (Figure 6b and Supplementary Fig. S8). Next, the knockdown of RNF123, a functional homolog of yeast Hel2p,33 repressed the RPS3 mono-ubiquitination that was induced by UV irradiation (Figure 6c). Even though the depletion of USP10, a homolog of yeast Ubp3p, did not result in increases in RPS3 mono-ubiquitination (Figure 6d), we observed increases in the di-ubiquitination of RPS3, suggesting multi-ubiquitination.25 Subsequently, to further extend our findings, we performed ribosomal pelleting experiments in UV-irradiated cells. As shown in Figure 6e, the association of RNF123 and USP10 with the ribosome complex was increased at early times after UV irradiation. Meanwhile, we could not show the increase in either RNF123 or USP10 in the non-ubiquitable RPS3K214R-expressing ribosome (Figure 6f). Therefore, we conclude that human RPS3 mono-ubiquitination is regulated by both RNF123 and USP10.
RNF123 and USP10 are implicated in the status of human RPS3 mono-ubiquitination. (a) The HT1080 cells were treated with various translational inhibitors, including 50 μg ml−1 cycloheximide, 20 μg ml−1 blasticidin S, 2 μg ml−1 DON, 2 μg ml−1 anisomycin, 150 J m−2 UV, 5 μM pactamycin, 50 μg ml−1 G418, 50 μg ml−1 puromycin, 50 μg ml−1 hygromycin B, and 20 μM emetine for 2 h. The cell lysates were subjected to immunoblot analysis using the indicated antibodies. (b) The HT1080 cells stably expressing the Flag-RPS3 wild-type or the K214R mutant were UV-irradiated (200 J m−2), harvested after 2 h and subjected to immunoblotting with anti-RPS3 or anti-Flag antibodies. (c and d) The HT1080 cells were transfected with siRNAs against RNF123 or USP10, treated with 150 J m−2 of UV irradiation and harvested after being incubated in fresh media for 2 h. The lysates were assayed by immunoblotting with the indicated antibodies. (e) The UV-irradiated HT1080 cells were incubated for the indicated times, followed by ultracentrifugation with a 20% sucrose cushion to isolate the ribosomal pellets. Each cell lysate and ribosomal pellet were subjected to immunoblot analysis using the indicated antibodies. (f) The HT1080 cells stably expressing the Flag-RPS3 wild-type or the K214R mutant were transfected with RPS3 5’-UTR siRNA. After 48 h, these cells were treated (or not) with 150 J m−2 UV and incubated for 15 min. Each cell lysate was ultracentrifuged with a 20% sucrose cushion to isolate the ribosomal fraction. Each cell lysate and ribosomal fraction were separated by SDS–PAGE and subjected to immunoblot analysis using the indicated antibodies. SDS–PAGE, SDS–polyacrylamide gel electrophoresis; UTR, untranslated region.
Mono-ubiquitination of Rps3p is responsible for cell viability and apoptosis
Thus far, we found that Rps3p mono-ubiquitination was modulated by a reciprocal action of the E3 ligase Hel2p/RNF123 and deubiquitinase Ubp3p/USP10. In yeasts, to determine whether the Rps3p mono-ubiquitination might have an important role in stress response, we generated a mutant strain whose non-ubiquitable RPS3 gene, which has an arginine substituted for lysine 212, was integrated into its own endogenous locus. When each transformed cell was plated on YPD plates after treatment with or without UV irradiation, the non-ubiquitable Rps3pK212R strain appeared to be more sensitive to UV irradiation than the Rps3p strains were (Figure 7a).
Mono-ubiquitination of Rps3p is responsible for cell viability and apoptosis. (a) Serial dilutions of the WT or Rps3pK212R mutant cells were spotted onto solid YPD media (lower). Serial dilutions of the WT or Rps3pK212Rmutant cells that were exposed to UV stress were spotted onto solid YPD media (upper). The plates were incubated for 2 days at 30 °C (b and c). The HT1080 cells stably expressing the Flag-RPS3 wild-type or the K214R mutant were transfected with RPS3 5’-UTR siRNA. After 48 h, these cells were irradiated (or not) with 150 J m−2 UV. (b) Subsequently, these cells were labeled with 35S-Met/35S-Cys (250 μCi per ml) for 15 min. Each cell lysate and ribosomal pellet were then subjected to immunoblot analysis using an anti-ubiquitin antibody (left). To detect the poly-ubiquitinated nascent polypeptides, immunoprecipitation with an anti-ubiquitin antibody was performed for ribosomal pellets, as described in ‘Materials and Methods,’ and subjected to autoradiography (right). (c) The UV-irradiated cells were incubated for 8 h, harvested, lysed, and analyzed by immunoblotting with the indicated antibodies. UTR, untranslated region.
According to a previous report, the mono-ubiquitination on the 40S small ribosomal subunit is responsible for the RQC in human cells.25 To confirm whether the non-ubiquitable RPS3K214R might affect the poly-ubiquitination of nascent polypeptides on the ribosome complex, we performed a series of experiments. First, to maximize the expression of exogenous FLAG-RPS3 and FLAG-RPS3K214R, an RPS3 3′-untranslated region targeting siRNA was transfected into each stable cell line. Then, each S35-Met-labeled cell lysate was subjected to ribosome pelleting, followed by immunoprecipitation with an anti-ubiquitin antibody. Consistent with a previous report,25 we confirmed that the poly-ubiquitination of the nascent polypeptide in RPS3K214R-expressing cells was increased more than in the wild-type RPS3-expressing cells (comparing lanes 9 and 11 (2.3-fold) and 10 and 12 (sixfold), Figure 7b). Because of PARP-1 cleavage, the RPS3K214R-expressing cells also showed more sensitivity to UV irradiation than the wild type did (Figure 7c). These results suggest that Rps3p mono-ubiquitination modulates cell viability and apoptosis through RQC.
Discussion
The ubiquitination of ribosomal proteins has been known to be able to regulate ribosome quality control. Many researchers are now trying to determine whether some E3 ligases and deubiquitinases are involved in the process, responding to translational stresses that cause ribosome stalling. The ribosome-associated E3 ligases have recently been demonstrated to have two different functions in the RQC pathway. Two ribosome-associated E3 ligases, Not4p and Ltn1p, have been identified to be involved in the ubiquitin-proteasome degradation pathway of partially synthesized polypeptides that are stuck on the stalled ribosome.40 Another ribosome-associated E3 ligase, Hel2p, seems to be recruited to the stalled ribosomes before the 80S monosome splits and contributes to ribosome stalling.30, 41 Although it has been recently reported that Hel2p is a no-go type of stalled translation surveillance,42 the detailed mechanism of Hel2p functioning in the RQC pathway has remained elusive. Surprisingly, we found that the mono-ubiquitination of Rps3p that is induced by translation elongation inhibitors was not increased in the hel2Δ strain. Additionally, in the HEL2 gene knockout experiment, UV-induced Rps3p mono-ubiquitination did not completely disappear in response to stress conditions. Thus, we cannot completely rule out the possibility of the involvement of additional E3 ligase(s) in the regulation of Rps3p mono-ubiquitination. Interestingly, the Rps3p mono-ubiquitination induced by rapamycin completely disappeared in an ASC1 deletion strain (Supplementary Fig. S9A). In human cells, the depletion of RACK1 also appeared to inhibit the UV-induced RPS3 mono-ubiquitination (Supplementary Fig. S9B). Asc1p, a scaffold protein in the ribosome complex, has been recently known to genetically and physically interact with many E3 ligases such as Mms22p,43 Prp19p,44 San1p,45 Cdc4p,46 Rad18p,46 and Hel2p. Among these ligases, Hel2p seems to play an early role in the RQC pathway with Asc1p.30, 41, 47 Hence, further studies are needed to test whether the Asc1p-associated E3 ligase has an additional role in Rps3p mono-ubiquitination.
Ubp3p, a yeast ortholog of mammalian USP10, has been demonstrated to be a deubiquitinase for the mono-ubiquitinated Rpl25p that regulates the selective autophagy of the 60S large subunit.37 In addition, Ubp3p forms a complex with the cofactor Bre5p to effectively deubiquitinate target proteins.39 Interestingly, Ubp3p has been reported to localize in both monosomes and polysomes.48 We wondered whether the mono-ubiquitinated Rps3p in the ribosome might act as a substrate for Ubp3p. Consistent with this expectation, the results of this study demonstrated that Ubp3p was a specific deubiquitinase for the mono-ubiquitinated Rps3p.
In this study, we found that rapamycin treatment had different effects on yeast and human cells in terms of the induction of Rps3p mono-ubiquitination. A recent study identified the ribophagy of the 60S large subunit in yeast under nitrogen starvation and suggested a model in which the Ubp3p-Bre5p complex has a key role in ribophagy by deubiquitinating the mono-ubiquitinated Rpl25p.39 On the basis of these findings, we propose that rapamycin treatment that promotes ribophagy might be able to sequester the Ubp3p-Bre5p complex in the 60S large ribosomal subunit far away from the 40S small ribosomal subunit. In other words, the detachment of Ubp3p from the ribosome complex might upset the balance of two proteins—the E3 ligase Hel2p and the deubiquitinase Ubp3p—that are responsible for the normal status of Rps3p mono-ubiquitination. Several lines of evidence presented in this study support this possibility. We first showed that rapamycin-induced Rps3p mono-ubiquitination was sustained for a long time in both ubp3Δ and bre5Δ strains compared to that in the wild-type strain. The deletion stain of UFD3, an interactor of Ubp3p and a positive regulator of ribophagy, could not induce Rps3p mono-ubiquitination under rapamycin treatment. In contrast, the rapamycin-induced Rps3p mono-ubiquitination was not increased in the atg7Δ or atg8Δ deletion strains, raising the possibility that the formation of autophagosome membranes could be negatively involved in rapamycin-induced Rps3p mono-ubiquitination. Therefore, it is suggested that the Rps3p mono-ubiquitination might cooperate with Ubp3p in the induction of ribophagy and the selective autophagy of 60S large ribosomal subunits.
Generally, temporary stalling of the ribosome complex often occurs during the translational elongation step under physiological conditions and may be caused by the presence of rare codons, mRNA secondary structures, and waiting for the appearance at the ribosomal exit channel of domains that are required for protein folding.27 There might exist a temporary and a quick rescue mechanism. In contrast, the severe ribosome stalling that is generated by translational inhibitors can activate the RQC pathway, mediating the poly-ubiquitination and degradation of ribosome-tethered nascent polypeptides. According to our results (Figure 7) and those of previous studies,25 the non-ubiquitable RPS3 mutant might induce an aberrant RQC, such as the non-removal of a defective nascent polypeptide, resulting in an increase in apoptosis. Considering the physiological significance of Rps3p mono-ubiquitination, we suggest here that Rps3p mono-ubiquitination, under normal conditions, maintains a basal level to monitor the reciprocal action between the E3 ligase Hel2p and the deubiquitinase Ubp3p. In line with this suggestion, we showed that both Hel2p (Figure 2c, right) and Ubp3p (Figure 3c, right) proteins existed at basal levels in the ribosome complex. In addition, we found that the deletion of Ubp3p induced Rps3p mono-ubiquitination without UV irradiation (Figure 3a, lane 3).
Therefore, we conclude that Rps3p mono-ubiquitination is involved in both proteasome-mediated degradation of aberrant nascent polypeptides in the RQC process and Ubp3p-dependent ribophagy that is regulated by the E3 ligase Hel2p/RNF123 and the deubiquitinase Ubp3p/USP10.
References
Ruggero D, Pandolfi PP Does the ribosome translate cancer?Nat Rev Cancer 2003; 3: 179–192.
Naora H . Involvement of ribosomal proteins in regulating cell growth and apoptosis: translational modulation or recruitment for extraribosomal activity? Immunol Cell Biol 1999; 77: 197–205.
Fumagalli S, Di Cara A, Neb-Gulati A, Natt F, Schwemberger S, Hall J et al. Absence of nucleolar disruption after impairment of 40S ribosome biogenesis reveals an rpL11-translation-dependent mechanism of p53 induction. Nat Cell Biol 2009; 11: 501–508.
Naora H, Takai I, Adachi M, Naora H . Altered cellular responses by varying expression of a ribosomal protein gene: sequential coordination of enhancement and suppression of ribosomal protein S3a gene expression induces apoptosis. J Cell Biol 1998; 141: 741–753.
Kim J, Chubatsu LS, Admon A, Stahl J, Fellous R, Linn S . Implication of mammalian ribosomal protein S3 in the processing of DNA damage. J Biol Chem 1995; 270: 13620–13629.
Kim YJ, Lee MS, Kim HD, Kim J . Ribosomal protein S3 (rpS3) secreted from various cancer cells is N-linked glycosylated. Oncotarget 2016; 7: 80350–80362.
Kim SH, Lee JY, Kim J . Characterization of a wide range base-damage-endonuclease activity of mammalian rpS3. Biochem Biophys Res Commun 2005; 328: 962–967.
Jang CY, Lee JY, Kim J . RpS3 a DNA repair endonuclease and ribosomal protein, is involved in apoptosis. FEBS Lett 2004; 560: 81–85.
Jung SO, Lee JY, Kim J . Yeast ribosomal protein S3 has an endonuclease activity on AP DNA. Mol Cells 2001; 12: 84–90.
Seong KM, Jung SO, Kim HD, Kim HJ, Jung YJ, Choi SY et al. Yeast ribosomal protein S3 possesses a beta-lyase activity on damaged DNA. FEBS Lett 2012; 586: 356–361.
Sandigursky M, Yacoub A, Kelley MR, Deutsch WA, Franklin WA . The Drosophila ribosomal protein S3 contains a DNA deoxyribophosphodiesterase (dRpase) activity. J Biol Chem 1997; 272: 17480–17484.
Deutsch WA, Yacoub A, Jaruga P, Zastawny TH, Dizdaroglu M . Characterization and mechanism of action of Drosophila ribosomal protein S3 DNA glycosylase activity for the removal of oxidatively damaged DNA bases. J Biol Chem 1997; 272: 32857–32860.
Yacoub A, Kelley MR, Deutsch WA . Drosophila ribosomal protein PO contains apurinic/apyrimidinic endonuclease activity. Nucleic Acids Res 1996; 24: 4298–4303.
Yusupova GZ, Yusupov MM, Cate JH, Noller HF . The path of messenger RNA through the ribosome. Cell 2001; 106: 233–241.
Takyar S, Hickerson RP, Noller HF . mRNA helicase activity of the ribosome. Cell 2005; 120: 49–58.
Wan F, Anderson DE, Barnitz RA, Snow A, Bidere N, Zheng L et al. Ribosomal protein S3: a KH domain subunit in NF-kappaB complexes that mediates selective gene regulation. Cell 2007; 131: 927–939.
Welchman RL, Gordon C, Mayer RJ . Ubiquitin and ubiquitin-like proteins as multifunctional signals. Nat Rev Mol Cell Biol 2005; 6: 599–609.
Denison C, Kirkpatrick DS, Gygi SP . Proteomic insights into ubiquitin and ubiquitin-like proteins. Curr Opin Chem Biol 2005; 9: 69–75.
Matafora V, D'Amato A, Mori S, Blasi F, Bachi A . Proteomics analysis of nucleolar SUMO-1 target proteins upon proteasome inhibition. Mol Cell Proteomics 2009; 8: 2243–2255.
Xirodimas DP, Sundqvist A, Nakamura A, Shen L, Botting C, Hay RT . Ribosomal proteins are targets for the NEDD8 pathway. EMBO Rep 2008; 9: 280–286.
Diaz JJ, Giraud S, Greco A . Alteration of ribosomal protein maps in herpes simplex virus type 1 infection. J Chromatogr B Analyt Technol Biomed Life Sci 2002; 771: 237–249.
Schafer T, Maco B, Petfalski E, Tollervey D, Bottcher B, Aebi U et al. Hrr25-dependent phosphorylation state regulates organization of the pre-40S subunit. Nature 2006; 441: 651–655.
Ghalei H, Schaub FX, Doherty JR, Noguchi Y, Roush WR, Cleveland JL et al. Hrr25/CK1delta-directed release of Ltv1 from pre-40S ribosomes is necessary for ribosome assembly and cell growth. J Cell Biol 2015; 208: 745–759.
Mitterer V, Murat G, Rety S, Blaud M, Delbos L, Stanborough T et al. Sequential domain assembly of ribosomal protein S3 drives 40S subunit maturation. Nat Commun 2016; 7: 10336.
Higgins R, Gendron JM, Rising L, Mak R, Webb K, Kaiser SE et al. The unfolded protein response triggers site-specific regulatory ubiquitylation of 40S ribosomal proteins. Mol Cell 2015; 59: 35–49.
Pechmann S, Willmund F, Frydman J . The ribosome as a hub for protein quality control. Mol Cell 2013; 49: 411–421.
Brandman O, Hegde RS . Ribosome-associated protein quality control. Nat Struct Mol Biol 2016; 23: 7–15.
Shao S, Brown A, Santhanam B, Hegde RS . Structure and assembly pathway of the ribosome quality control complex. Mol Cell 2015; 57: 433–444.
Bengtson MH, Joazeiro CA . Role of a ribosome-associated E3 ubiquitin ligase in protein quality control. Nature 2010; 467: 470–473.
Brandman O, Stewart-Ornstein J, Wong D, Larson A, Williams CC, Li GW et al. A ribosome-bound quality control complex triggers degradation of nascent peptides and signals translation stress. Cell 2012; 151: 1042–1054.
Iordanov MS, Pribnow D, Magun JL, Dinh TH, Pearson JA, Magun BE . Ultraviolet radiation triggers the ribotoxic stress response in mammalian cells. J Biol Chem 1998; 273: 15794–15803.
Finley D, Ulrich HD, Sommer T, Kaiser P . The ubiquitin-proteasome system of Saccharomyces cerevisiae. Genetics 2012; 192: 319–360.
Singh RK, Gonzalez M, Kabbaj MH, Gunjan A . Novel E3 ubiquitin ligases that regulate histone protein levels in the budding yeast Saccharomyces cerevisiae. PLoS ONE 2012; 7: e36295.
Ossareh-Nazari B, Nino CA, Bengtson MH, Lee JW, Joazeiro CA, Dargemont C . Ubiquitylation by the Ltn1 E3 ligase protects 60S ribosomes from starvation-induced selective autophagy. J Cell Biol 2014; 204: 909–917.
Sole C, Nadal-Ribelles M, Kraft C, Peter M, Posas F, de Nadal E . Control of Ubp3 ubiquitin protease activity by the Hog1 SAPK modulates transcription upon osmostress. EMBO J 2011; 30: 3274–3284.
Stewart MD, Ritterhoff T, Klevit RE, Brzovic PS . E2 enzymes: more than just middle men. Cell Res 2016; 26: 423–440.
Kraft C, Deplazes A, Sohrmann M, Peter M . Mature ribosomes are selectively degraded upon starvation by an autophagy pathway requiring the Ubp3p/Bre5p ubiquitin protease. Nat Cell Biol 2008; 10: 602–610.
Noda NN, Satoo K, Fujioka Y, Kumeta H, Ogura K, Nakatogawa H et al. Structural basis of Atg8 activation by a homodimeric E1, Atg7. Mol Cell 2011; 44: 462–475.
Ossareh-Nazari B, Bonizec M, Cohen M, Dokudovskaya S, Delalande F, Schaeffer C et al. Cdc48 and Ufd3, new partners of the ubiquitin protease Ubp3, are required for ribophagy. EMBO Rep 2010; 11: 548–554.
Panasenko OO . The role of the E3 ligase Not4 in cotranslational quality control. Front Genet 2014; 5: 141.
Kuroha K, Akamatsu M, Dimitrova L, Ito T, Kato Y, Shirahige K et al. Receptor for activated C kinase 1 stimulates nascent polypeptide-dependent translation arrest. EMBO Rep 2010; 11: 956–961.
Saito K, Horikawa W, Ito K . Inhibiting K63 polyubiquitination abolishes no-go type stalled translation surveillance in Saccharomyces cerevisiae. PLoS Genet 2015; 11: e1005197.
Buser R, Kellner V, Melnik A, Wilson-Zbinden C, Schellhaas R, Kastner L et al. The replisome-coupled E3 ubiquitin ligase Rtt101Mms22 counteracts Mrc1 function to tolerate genotoxic stress. PLoS Genet 2016; 12: e1005843.
Stevens SW, Ryan DE, Ge HY, Moore RE, Young MK, Lee TD et al. Composition and functional characterization of the yeast spliceosomal penta-snRNP. Mol Cell 2002; 9: 31–44.
Rosenbaum JC, Fredrickson EK, Oeser ML, Garrett-Engele CM, Locke MN, Richardson LA et al. Disorder targets misorder in nuclear quality control degradation: a disordered ubiquitin ligase directly recognizes its misfolded substrates. Mol Cell 2011; 41: 93–106.
Costanzo M, Baryshnikova A, Bellay J, Kim Y, Spear ED, Sevier CS et al. The genetic landscape of a cell. Science 2010; 327: 425–431.
Ikeuchi K, Inada T . Ribosome-associated Asc1/RACK1 is required for endonucleolytic cleavage induced by stalled ribosome at the 3' end of nonstop mRNA. Sci Rep 2016; 6: 28234.
Wang CY, Wen WL, Nilsson D, Sunnerhagen P, Chang TH, Wang SW . Analysis of stress granule assembly in Schizosaccharomyces pombe. RNA 2012; 18: 694–703.
Acknowledgements
This work was supported by NRF 2015R1A2A1A05001823 and 2017R1E1A1A01074101. The single deletion strains of yeast were gifted by Cheol-Won Yun (Division of Life Sciences, Korea University).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Experimental & Molecular Medicine website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Jung, Y., Kim, H., Yang, H. et al. Modulating cellular balance of Rps3 mono-ubiquitination by both Hel2 E3 ligase and Ubp3 deubiquitinase regulates protein quality control. Exp Mol Med 49, e390 (2017). https://doi.org/10.1038/emm.2017.128
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/emm.2017.128
This article is cited by
-
UBE2J1 inhibits colorectal cancer progression by promoting ubiquitination and degradation of RPS3
Oncogene (2023)
-
Remodeling of the ribosomal quality control and integrated stress response by viral ubiquitin deconjugases
Nature Communications (2023)
-
Loss of N-terminal acetyltransferase A activity induces thermally unstable ribosomal proteins and increases their turnover in Saccharomyces cerevisiae
Nature Communications (2023)
-
The deubiquitinase USP10 protects pancreatic cancer cells from endoplasmic reticulum stress
npj Precision Oncology (2022)
-
Ribosomal protein S3 associates with the TFIIH complex and positively regulates nucleotide excision repair
Cellular and Molecular Life Sciences (2021)