Abstract
Mutations occur during cell division in all somatic lineages. Because neurogenesis persists throughout human life, somatic mutations in the brain arise during development and accumulate with the aging process. The human brain consists of 100 billion neurons that form an extraordinarily intricate network of connections to achieve higher level cognitive functions. Due to this network architecture, perturbed neuronal functions are rarely restricted to a focal area; instead, they are often spread via the neuronal network to affect other connected areas. Although somatic diversity is an evident feature of the brain, the extent to which somatic mutations affect the neuronal structure and function and their contribution to neurological disorders associated with disrupted brain connectivity remain largely unexplored. Notably, recent reports indicate that brain somatic mutations can indeed play a critical role that leads to the structural and functional abnormalities of the brain observed in several neurodevelopmental disorders. Here, I review the extent and significance of brain somatic mutations and provide my perspective regarding these mutations as potential molecular lesions underlying relatively common conditions with disrupted brain connectivity. Moreover, I discuss emerging technical platforms that will facilitate the detection of low-frequency somatic mutations and validate the biological functions of the identified mutations in the context of brain connectivity.
Similar content being viewed by others
Introduction
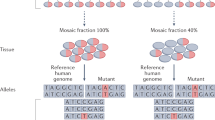
Somatic mutations are genetic variations in somatic cells that are not inherited from a parent and are also not passed on to offspring. Somatic cells in humans are known to acquire two to three mutations per DNA replication and one to nine base-substitution mutations per year in a replication-independent manner.1 Thus, somatic mutations exist in all somatic lineages and comprise any tissue or organ, and thus, the cells of the human body do not all contain identical genome sequences.2 Such somatic mutations can be single nucleotide variations (SNVs), copy number variations (CNVs), loss of heterozygosity or large structural variations (SVs). Based on when or where the mutations arise, somatic mutations occur as a mosaic pattern that affects the whole body or they are restricted to a certain tissue or even to a focal area within an organ. For example, somatic mosaic variations caused by early-embryonic mutation events can be distributed across many organs and tissues and are not limited to a given organ or tissue. Genome mosaicism and its implications in human disease have been carefully reviewed in several other papers.3, 4 In this review, I focus on somatic mutations in the brain and the potentially related neurological disorders.
Somatic mutations in the human brain can arise during development or aging. During the 4–24-week gestational interval, the neuronal precursor cells in the developing human brain undergo more than 105 divisions per minute.5 During the neonatal period, the subventricular zone shows a robust neurogenesis and new neurons migrate to the prefrontal cortex.6 In the adult brain, new neurons are generated in the dentate gyrus of the hippocampus as well as in the subventricular zone adjacent to the lateral ventricle7, 8, 9, 10 (Figure 1a). A recent carbon dating study showed that 700 new neurons are added in the hippocampus per day, and one-third of the hippocampal neurons are subject to exchange throughout life.9 Neurogenesis in the adult human brain is also observed in the striatum, which is adjacent to the lateral ventricle with a neurogenic niche.8 In addition to lifelong neurogenesis, increasing evidence demonstrates that the brain exhibits somatic mutations during development or aging.11, 12, 13, 14 For example, single-cell whole-genome sequencing in neurons revealed clonal somatic CNVs in the normal human brain.11, 12 A single-cell analysis of the somatic mutations in multiple brain regions also suggested that focally distributed mutations are prevalent in the human brain.14 Somatic mutations in mitochondrial DNA have been reported in the aging brain.13 Therefore, it is not surprising that somatic mutations in the brain occur during the post-fertilization or post-zygote period and accumulate with aging (Figure 1b).
Regions of neural stem cells and the occurrence of somatic mutations in the human brain. (a) The figure shows the subventricular zones (red) along with the wall of the lateral ventricle and the dentate gyrus (blue) in the hippocampus, from which neural stem cells have been isolated in the human brain. (b) The schematic figure shows the concept of a mosaic brain with somatic mutations that can arise during development or over aging. Depending on the timing and location of the mutations, somatic mutations may be present in the regional brain structures of various sizes or even in the entire brain.
Regarding neurological disorders, it is well known that somatic mutations cause most brain tumors, which is similar to the occurrence of other human cancers.15 Other than brain tumors, neurological disease-causing or -associated mutations have been considered inherited or de novo mutations in germ cells, and therefore, they could be detected in any tissue of the affected patient, such as readily accessible blood or saliva. However, damaging mutations in biologically important genes may be incompatible with normal development or the survival of an organism and may not be detected as inherited or de novo mutations in germ cells but might be detected as somatic mutations in a small fraction of the somatic cells. In addition, brain function is more vulnerable to the focal functional or structural damage caused by somatic mutations in comparison to other organs because of the nature of brain connectivity.16 Nevertheless, the extent and significance of somatic mutations in most brain disorders, including neurodevelopmental, neuropsychiatric and neurodegenerative disorders, has remained unresolved.
The implication of somatic mutations in disorders with disrupted brain connectivity
The human brain is an extraordinarily intricate network comprising ~1011 neurons with ~1015 connections. The connection matrix of the human brain is the so-called ‘connectome’.17 An important property of an interdependent network, such as the connectome, is that the dysfunction of a node in the network can spread throughout the connections and disrupt the function of the entire network.16, 18 In the brain, neurons act as nodes, and synaptic and axonal contacts function as connections and conduits for the pathological spread.16 Therefore, neuronal dysfunctions or abnormalities are not limited to a restricted area but spread to or affect other regions, thereby altering neural connectivity.16 Indeed, various studies in the human brain using MRI, functional MRI (fMRI), diffusion tensor imaging (DTI) or electroencephalography (EEG) have shown that brain disorders, including neurodevelopmental (for example, epilepsy or autism), neuropsychiatric (for example, schizophrenia (SZ) or depression) and neurodegenerative (for example, Alzheimer’s disease) disorders are associated with the altered structure or function of the focal regions and the disrupted neural connectivity in the brain.19, 20, 21, 22, 23 Consistent with these observations, such conditions are often understood as disorders with disrupted brain connectivity.24, 25, 26
What is the genetic etiology of such brain disorders? These conditions are often familial, and twin studies have indicated that their concordance rates range from 20 to 90% (20–75% for epilepsy;27 70–90% for autism;28 ~80% for SZ;29 ~40% for major depression30 and 74% for Alzheimer’s disease31). These findings suggest that genetic factors play a major role in the development of these disorders. Inherited or de novo genetic variations, including SNVs and CNVs, are considered the most likely genetic etiology of disorders with disrupted brain connectivity. Although such concordance rates imply that heritable variations substantially contribute to these conditions, these diseases frequently occur as isolated cases. As a possible explanation for this, de novo mutations in germ cells have also been considered to account for the genetic etiologies of most of sporadic cases that are not sufficiently explained by inherited mutations.32, 33 Indeed, recent large-scale genomic studies using whole-exome or whole-genome sequencing in peripheral tissue samples (for example, blood) from patients and their family members showed that the candidate genes harboring recurrent de novo mutations are implicated in epilepsy,34 autism35 and SZ.36 However, the identified de novo mutations explain only a limited portion of the sporadic cases. For example, a recent large-scale exome sequencing of peripheral samples from 264 patients–parent trios with epileptic encephalopathy identified genetic causes in ~7% of the patients in the cohort.34 In autism, recent whole exome sequencing (WES) studies suggest that de novo SNVs and CNVs contribute to the risk of autism in ~30% of the sporadic cases in simplex families35 and that no single gene accounts for >1% of autism cases.37 In SZ, de novo mutations appear to contribute to ~25% of all sporadic cases.38, 39 However, the genetic contribution to most of the sporadic cases remains unanswered. Regarding this point, cancer genomics may offer some instructive insight to explain the genetic contribution to the sporadic form of brain network disorders. Although cancers have a substantial heritability with several defined mutations at the level of the germ cells, most of the mutations underlying cancer arise somatically in affected tissues or cell types, and these somatic mutations can explain the genetic contribution to sporadic cancers.15 Among the >20 000 protein-coding genes, cancer genomic studies, to date, have identified ~140 core genes that are recurrently mutated and drive tumorigenesis in affected but not in unaffected tissues.15 The occurrence of somatic mutations demonstrated in cancer genomics may be applicable to and testable in sporadic cases with disrupted brain connectivity. However, due to the difficulty associated with accessing brain tissues and the limitations of the sequencing technology for detecting low-level somatic mutations without cell-proliferative effects (as discussed later in this review), the extent of the brain somatic mutations in these diseases has remained largely unexplored. Recently, the notion of brain somatic mutations has received great attention as the underlying pathogenesis in focal epilepsy. Herein, I first review the current evidence for the pathogenic roles of brain somatic mutations in epilepsy. Subsequently, I provide my perspective regarding brain somatic mutations as a potential molecular lesion underlying other conditions of disrupted brain connectivity, such as autism spectrum disorder (ASD), SZ, major depression disorder and Alzheimer’s disease.
Epilepsy
Epilepsy is a typical brain disorder with neural network dysfunction, including excessive and synchronous electrical activity throughout the brain.19 Approximately 60% of the people with epilepsy have focal seizures that originate in just one part of the brain. These seizures often spread to the other side of brain or the entire brain via the neural connection and contribute largely to the development of chronic epilepsy. Approximately 20–30% of epilepsy cases are medically refractory and such intractable epilepsies often require epilepsy surgery to resect the suspected origin of the seizures as an alternative treatment, which allows access to the affected brain tissue for the testing of brain somatic mutations.40 In particular, for a long time it was thought that somatic mutations might play a role in intractable epilepsy combined with focal cortical malformations because most of these diseases occur sporadically without a family history and sometimes show discordance in monozygotic twins.41, 42 For a rare condition, called ‘hemimegalencephaly (HME),’ which presents an enlargement of one side of the brain and is associated with intractable focal epilepsy, we and others provide evidence for somatic mutations in the PI3K–AKT–mTOR pathway genes and thereby suggest that brain somatic mutations might be important for other neurodevelopmental disorders.43, 44, 45 Somatic mutations accounted for up to 30% of the studied HME patients, and the frequency of the mutated allele ranged from 8 to 39% in the enlarged brain tissues.43 Recently, brain somatic mutations in MTOR and its related genes were identified in up to 25% of the patients studied with focal cortical dysplasia, which is more common and accounts for up to 50% of the children undergoing epilepsy surgery46, 47, 48, 49 (Figure 2a). Notably, brain MRIs from patients carrying somatic mutations were frequently reported to be normal, which thereby implies an extremely low level of mutational burden in the affected areas.48 Indeed, the frequency of the mutated allele was reported to be as low as ~1% in the resected brain tissues, but the mutated allele was not detected in the peripheral samples, such as the blood or saliva.48, 49 More surprisingly, such a small fraction of single mTOR missense mutation-carrying neurons were sufficient to cause intractable epilepsy via the neural network in an in utero electroporated mouse model (Figures 2b and c).48 This finding provides direct evidence that the low-level brain somatic mutations found in patients indeed cause the brain network disorder by disrupting the entire brain function as a whole in vivo (Figure 2d). Although these recent studies revealed the extent of the brain somatic mutations in only a few examples of intractable epilepsies that permitted access to the surgically resected brain tissues, these findings imply that somatic mutations that alter the excitability in the affected brain regions might be responsible for a substantial proportion of the epilepsy cases that remain unexplained by genomic studies in the peripheral samples.34
Low-level somatic mutations in a focal brain region cause the synchronized electrical discharge of the entire brain network. (a) Post-op MRI and pathology from focal cortical dysplasia II (FCDII) with intractable epilepsy show the resected brain region (white arrow) and the scattered dysplastic neurons (black arrow) with the mTOR-activating somatic mutation (p.Leu2427Pro). Within the resected brain tissue, deep WES detected 6.9% mutated alleles.48 Scale bar, 50 μm (b) In utero electroporation of the mTOR p.Leu2427Pro mutant plasmid into the focal area of the embryonic brain (E14) followed by monitoring of the mTOR mutant expression using a GFP reporter. Imaging of the GFP signal in the coronal sections of an electroporated mouse brain at ~3 months of age shows a small fraction of GFP-positive cells expressing the mutated mTOR, accounting for 1.30% of all the DAPI-positive cells in the left cerebral cortex. Scale bar, 500 μm. (c) The in utero electroporated mouse exhibits a synchronized epileptic discharge with behavioral seizures by video-electroencephalograph (EEG) monitoring. LF, left frontal electrode; LT, left temporal electrode; RF, right frontal electrode; RT, right temporal electrode. (d) The schematic figure shows how somatic mutations in a focal region can affect the function of the entire brain via the brain connectivity. (a, c) are adopted from ref. 48. GFP, green fluorescent protein; MRI, magnetic resonance imaging.
Autism spectrum disorder
ASDs are associated with disrupted brain connectivity.50 However, the genetic etiology that results in the disruption of brain connectivity in ASD remains poorly understood. Recent ASD family-based genomic studies show that de novo genetic variations occurring at the germ cell level are significantly associated with ASDs, but specific causes, including mutations in the synapse proteins, were identified in only a modest fraction of the individuals with ASDs.51, 52 Moreover, a recent study showed that most of the observed de novo mutation events in the germline have limited effects on ASD and that those that do confer a risk are not necessarily sufficient for disease,52 which thereby implies the possibility of a genetic cause of ASD that may not be detected at the germline level. Many functional and structural MRIs studies in ASD show altered activities and increased cortical thickness in specific brain regions, including the prefrontal cortex that mediates autistic symptoms.53 In addition, neuropathological findings from the postmortem brains of autism patients indicated the presence of focal abnormalities in the prefrontal cortex, including a patched cortical dyslamination and dysplastic changes in the neurons.54, 55, 56 Interestingly, the prefrontal cortex in humans is known to be the major target area of the new neurons generated from robust postnatal neurogenesis in the subventricular zone.6 The burden of the somatic mutations can accumulate during neuronal cell proliferation. Therefore, it is worth testing the hypothesis that somatic mutations in the affected brain regions, such as the prefrontal cortex, participate in the pathogenesis of ASD.57
SZ and major depressive disorder (MDD)
Can brain somatic mutations be a pathogenic factor in neuropsychiatric disorders such as SZ and MDD? Although the contribution of brain region-specific somatic mutations in mitochondrial DNA has been explored in neuropsychiatric disorders,58 investigators have recently increasingly recognized the potential role of brain somatic mutations in the genomic DNA.5 The high discordance rates (~40–60%) in monozygotic twins have raised the possibility that de novo mutations might be implicated in these mental disorders.59 Consistent with these findings, recent large-scale genomic studies identified the association of SZ and MDD with several common or rare de novo genetic variations.36, 60, 61 However, these findings are insufficient to explain most of the cases of sporadic SZ and MDD. Although numerous efforts, such as the identification of disrupted synaptic networks caused by multiple de novo mutated genes or epigenetic changes, have been made to find the another level of genetic contribution,36, 62 this unexplained genetic contribution gives rise to the possibility that the genetic variations implicated in SZ and MDD may not be easily detected in peripheral samples such as blood and saliva but rather may be localized to the affected brain areas. Dysfunction in the prefrontal cortex, which is critical for converting the external stimuli into a response, is frequently implicated in SZ and MDD.63, 64 Interestingly, it was recently reported that the somatic copy number of L1 retrotransposition in the prefrontal cortex increases in SZ and MDD patients, and the brain-specific L1 insertion is preferentially localized to synapse-related genes in SZ.65 In addition, somatic deletions are reported in the prefrontal cortex of SZ patients, as well as unaffected controls.66 To date, only a few have investigated brain somatic mutations in neuropsychiatric disorders, and no studies have found SZ- and MDD-associated brain-specific SNVs or the pathogenic causality of the brain somatic mutations observed in SZ or MDD yet. Therefore, additional systematic studies are needed to test the hypothesis regarding the pathogenic role of brain somatic mutations in neuropsychiatric disorders.
Alzheimer’s disease
Investigators have focused on identifying the molecular genetic causes of Alzheimer’s disease (AD) for more than a century. Indeed, these efforts enabled the identification of the disease-causing mutations in the APP or Presenilin 1 or 2 genes in early-onset familial AD and the disease-associated mutations in the APOE4 or TREM2 gene in late-onset AD.67 However, these genetic variations were absent in the peripheral genomic DNA collected from most cases of sporadic AD (sAD).67 Most sAD has the characteristics of a sporadic occurrence, a focal onset of pathology and a progressive spread of the protein aggregates. These features have raised the hypothesis that brain somatic mutations contribute to neurodegenerative disorders including sAD.68 Neurofibrillary tangles parallel the neuronal loss and progression of cognitive decline.69 Interestingly, neurofibrillary tangles first appear in the hippocampal area (for example, entorhinal cortex) and spread to the other brain regions, including the neocortex, and thereby, correlate with the symptomatic progression.69, 70 Somatic mutations accumulate with aging, and their occurrence increases in proliferating cells. Considering that hippocampal neurogenesis continues during the aging process and that one-third of the hippocampal neurons are subject to change over a life-time,9 it is plausible to hypothesize that the accumulation of somatic mutations in the hippocampal regions may contribute to the production or progression of neurofibrillary tangles, which might play a role in the potential pathogenesis of AD. Indeed, it was recently reported that brain somatic mutations with a low allele frequency are present in the entorhinal cortex or hippocampal genomic DNA from patients with sAD.71, 72 However, these studies still lack the direct evidence that brain somatic mutations contribute to or are associated with the pathogenesis of sAD. Therefore, further studies with larger samples or more systematic deep sequencing at the genome-wide level will be necessary to test the hypothesis that brain somatic mutations are a genetic cause of sAD.
Detecting brain somatic mutations with a low allele frequency
Unlike somatic mutations, which have highly proliferative effects in human cancers, brain somatic mutations in non-cancerous neurological disorders are likely to have no or fewer proliferative effects. Consequently, mutation-carrying cells can be surrounded by a large number of normal cells, and the allele frequency of the somatic mutations in the affected brain regions can be much lower than the typical mutational burden found in tumor samples. For example, recent studies using ultra-deep sequencing showed that somatic mutations in the affected brain of intractable focal epilepsy or sAD patients were detected in as low as 1% of the mutated allele.48, 49, 71 In particular, the low level of the brain somatic mutations found in intractable focal epilepsy was sufficient to cause a spontaneous seizure (Figures 2b and c).48, 49 Therefore, it is important to sensitively and accurately detect the brain somatic mutations that occur at a low allele frequency. To achieve this goal, several criteria should be considered: (1) the use of different sequencing platforms with a high read depth; (2) single-cell sequencing or an enrichment of the mutated cells; and (3) the selection of the suitable variant callers.
The use of different sequencing platforms with a high read depth
Using traditional Sanger sequencing, which is generally considered the gold standard to confirm SNVs, it is difficult to detect <10–20% of the low level mutations.73 To detect such low-level mutations, high-coverage sequencing that quantifies the level of the mutations is necessary. A read depth of >1000 × is known to detect an allele frequency of 1% with >90% probability.74 Even with such a high read depth, unexpected sequencing artifacts or erroneous calls can arise from the sample, and library preparation or sequencing and imaging specific to the sequencing platforms can often mimic true somatic mutations with a low frequency. For example, hybrid capture and PCR-based amplicon sequencing may have unique artifacts arising during the library preparation.75, 76 In addition, according to the type of sequencing platform used, including Illumina, Ion Torrent, PacBio and digital PCR, the sequencing error rate ranges from almost 0 to ~4% with different types of errors, such as substitution, indel and deletion.77 Therefore, an increased sequencing read depth and various experimental replications, such as technical, biological and cross-platform replicates have been proposed as tools to minimize such false positive or erroneous variants.78 Indeed, technical and biological replications, which are commonly performed in molecular biology experiments, as well as cross-platform replications with a high read depth, reduce the false variants and detect the true variants when the mutated alleles are present at 1%.48, 49, 78 Nevertheless, further studies that optimize the read depth and experimental replications with different platforms will be necessary to provide an accurate and cost-effective method for detecting somatic mutations with a low allele frequency.
Single cell sequencing or enrichment of the mutated cells
The sequencing technologies with a high read depth described above utilize pooled genomic DNA isolated from brain tissues containing many distinct cell types. By contrast, single cell sequencing reads the genome of individual cells, which thereby pinpoints specific cells or cell types that contain somatic mutations in a given brain tissue. To do this, single cells should first be isolated from brain tissues using various techniques, such as fluorescence-activated cell sorting, laser capture microdissection (LCM) and microfluidic devices, which have unique technical advantages and disadvantages (for example, unbiased vs biased sampling or high- vs low-throughput).79 The single-cell genome (for example, ~6 pg DNA per a cell) is then amplified to obtain sufficient amounts of DNA for next-generation sequencing (for example, 250–500 ng DNA for whole genome or exome sequencing).80 The use of such single-cell sequencing technologies has revealed that rare somatic CNVs and L1 retrotransposition events occur in the normal and diseased human brain.11, 12, 14 However, single-cell genome sequencing methods, especially for the detection of SNVs, are challenged by extensive technical errors and poor physical coverage data arising due to the nature of the whole genome amplification process.80, 81 Many ongoing investigations are currently focused on overcoming these technical challenges.81 Alternatively, if researchers are able to specify or label the neuronal cells that potentially carry the mutations, the enrichment of these cells using single-cell isolation methods could be followed by sequencing without whole genome amplification, which might lead to the identification of brain somatic mutations with a low allele frequency. For example, a recent study of brain somatic mutations in epilepsy showed that even Sanger sequencing in a pool of 20–100 dysplastic neurons enriched by LCM was able to validate the low-level somatic mutations that were originally presented as ~5% of the mutation burden in the affected brain tissues.48
The selection of suitable variant callers
Most of the variant callers used to detect somatic mutations from whole genome or exome data were developed mainly in the field of cancer genomics. In general, these callers are designed to detect somatic mutations with a high frequency by jointly analyzing the paired diseased-normal samples from each patient. Recently, due to the challenges associated with tumor heterogeneity, several callers, such as MuTect, Strelka and Virmid, have been developed to detect low-level somatic mutations in impure tissue samples with a high sensitivity and specificity.82, 83, 84 By adopting these variant callers in recent studies of intractable focal epilepsy, brain somatic mutations present as 1–10% of the mutation burden could be identified from deep whole exome sequencing data in matched brain–blood (or saliva) tissues.48, 49 However, the sensitivity of these callers significantly declines in samples with <8% of the mutation burden,85 and they only detect SNVs or small indels in paired samples and not low-level CNVs and SVs, which might be implicated in disorders with disrupted brain connectivity. In addition, considering the difficulty related to gaining access to matched brain-peripheral samples from patients with the somatic mosaicism that is present at a low level in the brain and other peripheral tissues,73 the ideal somatic variant callers should be able to detect low-level SNVs, as well as CNVs and SVs in unmatched or matched tissue samples. Although a few recently developed algorithms address some of these issues, such as the low-frequency SNVs or deletions in the unmatched samples,86, 87 further studies are needed to develop more ideal variant callers.
Verifying the biological functions of the somatic mutations in the context of brain connectivity
Recent advances in sequencing technologies and bioinformatics analysis tools have greatly improved the comprehensive catalogue of low-frequency somatic mutations that are present in genetically heterogeneous tissues. However, to verify the biological and pathological effects of such variants, laboratory experiments at the level of the cell or organism are essential. Although it has been difficult to design experiments that recapitulate the somatic alterations of the eukaryotic genome, sequence-specific programmable nucleases provide a new opportunity for the targeted modification of the DNA sequence itself.88 In particular, the RNA-guided CRISPR-associated Cas9 nuclease enables a convenient system for achieving targeted somatic mutagenesis in vitro, as well as in vivo.89, 90 The CRISPR–Cas9 system can be customized for the specific purpose of somatic genomic editing. In this system, double-strand break generation by the introduction of Cas9 and single-guide (sg) RNAs targeting protein-coding regions induce loss-of-function frameshift indel mutations via non-homologous end-joining. The addition of donor DNA or single-strand deoxyoligonucleotides that contain mutated sequence introduces precise mutations, including gain-of-function one via homology-directed repair.90 For example, the biological consequences of somatic loss-of-function alterations at the cellular level can be screened via high-throughput functional genomics using the CRISPR–Cas9 systems with lentiviral single-guide RNA libraries.91, 92 The somatic loss-of-function or gain-of-function mutations found in patients with lung and liver cancers were successfully recapitulated by CRISPR–Cas9-mediated genome editing in mouse models.93, 94 In particular, to understand the biological effects of somatic mutations at the level of brain connectivity, in vivo genome editing in the brain may be critical. In agreement with this notion, recent studies show that the delivery of Cas9/sgRNA vectors using in utero electroporation or adeno-associated virus (AAV) was able to edit the genome of neurons and phenocopy brain tumors or Rett-syndrome with learning and memory deficits in the mouse brain.95, 96 To date, none of the brain somatic mutations identified in patients with disrupted neural connectivity, such as intractable focal epilepsy, have been modeled by somatic genome editing, but modeling of the brain somatic mutations found in patients via CRISPR–Cas9-mediated genome editing appears to be achievable in the near future.
In addition to advances in genome editing technology, a wide range of experimental tools in neuroimaging and electrophysiology have been developed to investigate the structure and function of brain connectivity at many levels, from the whole brain to neurons and to individual synapses. DTI delineates white matter tracts to investigate structural connectivity; whereas, fMRI and EEG examine the functional connectivity or network activity at the whole brain level.97, 98 In vivo two-photon imaging using viral tracers or calcium/voltage-sensitive indicators, multi-electrode recordings of local field potentials and optogenetic approaches permit the examination of structural and functional changes in the neuronal interconnections at the local circuit level.99, 100, 101 Electron-microscopy with imaging reconstruction allows the visualization of neuronal connections at the level of the synapse.102 Together with in vivo somatic genome editing, these experimental tools will aid in verifying the biological or pathological effects of somatic mutations on the structure and function of brain connectivity. Furthermore, the successful generation of disease models with brain somatic mutations will facilitate the validation of therapeutic targets genetically identified in each patient,48 which will thereby pave the way toward the realization of precision medicine in brain disorders in the near future.
Conclusions and future directions
Our understanding of the extent and roles of somatic mutations in brain disorders is limited due to the difficulties associated with detecting low-level somatic mutations in surgical or postmortem brain tissues and verifying their functional consequences in the context of the neural network. However, recent advances in technology, for example, single-cell sequencing or deep sequencing with proper bioinformatics analysis, somatic genome editing and experimental tools for investigating brain connectivity, will enable a better understanding of (1) the prevalence of somatic mutations in disorders with disrupted brain connectivity; (2) the biological and pathological effects of somatic mutations on the structure and function of brain connectivity; and (3) the physiological roles of somatic mutations during brain development or aging, as well as provide improved guidance for (4) the development of precision medicine for brain disorders by discovering the genetically identified targets similar to those identified in cancer medicine. We are confident that studies of brain somatic mutations will substantially advance our knowledge of the genetics, pathogenesis and therapeutics of common but unexplained disorders with disrupted brain connectivity.
References
Lynch M . Rate, molecular spectrum, and consequences of human mutation. Proc Natl Acad Sci USA 2010; 107: 961–968.
Lupski JR . Genome mosaicism—one human, multiple genomes. Science 2013; 341: 358–359.
Biesecker LG, Spinner NB . A genomic view of mosaicism and human disease. Nat Rev Genet 2013; 14: 307–320.
Campbell IM, Shaw CA, Stankiewicz P, Lupski JR . Somatic mosaicism: implications for disease and transmission genetics. Trends Genet 2015; 31: 382–392.
Insel TR . Brain somatic mutations: the dark matter of psychiatric genetics? Mol Psychiatry 2014; 19: 156–158.
Sanai N, Nguyen T, Ihrie RA, Mirzadeh Z, Tsai H-H, Wong M et al. Corridors of migrating neurons in the human brain and their decline during infancy. Nature 2011; 478: 382–386.
Eriksson PS, Perfilieva E, Björk-Eriksson T, Alborn A-M, Nordborg C, Peterson DA et al. Neurogenesis in the adult human hippocampus. Nat Med 1998; 4: 1313–1317.
Ernst A, Alkass K, Bernard S, Salehpour M, Perl S, Tisdale J et al. Neurogenesis in the striatum of the adult human brain. Cell 2014; 156: 1072–1083.
Spalding KL, Bergmann O, Alkass K, Bernard S, Salehpour M, Huttner HB et al. Dynamics of hippocampal neurogenesis in adult humans. Cell 2013; 153: 1219–1227.
Sanai N, Tramontin AD, Quiñones-Hinojosa A, Barbaro NM, Gupta N, Kunwar S et al. Unique astrocyte ribbon in adult human brain contains neural stem cells but lacks chain migration. Nat Cell Biol 2004; 427: 740–744.
Cai X, Evrony GD, Lehmann HS, Elhosary PC, Mehta BK, Poduri A et al. Single-cell, genome-wide sequencing identifies clonal somatic copy-number variation in the human brain. Cell Rep 2014; 8: 1280–1289.
McConnell MJ, Lindberg MR, Brennand KJ, Piper JC, Voet T, Cowing-Zitron C et al. Mosaic copy number variation in human neurons. Science 2013; 342: 632–637.
Williams SL, Mash DC, Züchner S, Moraes CT . Somatic mtDNA mutation spectra in the aging human putamen. PLoS Genet 2013; 9: e1003990.
Evrony GD, Lee E, Mehta BK, Benjamini Y . Cell lineage analysis in human brain using endogenous retroelements. Neuron 2015; 85: 49–59.
Vogelstein B, Papadopoulos N, Velculescu VE, Zhou S, Diaz LA, Kinzler KW . Cancer genome landscapes. Science 2013; 339: 1546–1558.
Fornito A, Zalesky A, Breakspear M . The connectomics of brain disorders. Nat Rev Neurosci 2015; 16: 159–172.
Sporns O, Tononi G, Kötter R . The human connectome: a structural description of the human brain. PLOS Comput Biol 2005; 1: e42.
Buldyrev SV, Parshani R, Paul G, Stanley HE, Havlin S . Catastrophic cascade of failures in interdependent networks. Nature 2010; 464: 1025–1028.
Engel J Jr, Thompson PM, Stern JM, Staba RJ, Bragin A, Mody I et al. Connectomics and epilepsy. Curr Opin Neurobiol 2013; 26: 186–194.
Fornito A, Bullmore ET . Connectomic intermediate phenotypes for psychiatric disorders. Front Psychiatry 2012; 3: 32.
Hahamy A, Behrmann M, Malach R . The idiosyncratic brain: distortion of spontaneous connectivity patterns in autism spectrum disorder. Nat Neurosci 2015; 18: 302–309.
Matthews PM, Filippini N, Douaud G . Brain structural and functional connectivity and the progression of neuropathology in Alzheimer's disease. J Alzheimers Dis 2013; 33 (Suppl 1): S163–S172.
Narr KL . Leaver AM. connectome and schizophrenia. Curr Opin Psychiatry 2015; 28: 229–235.
Peled A . Brain ‘Globalopathies’ cause mental disorders. Med Hypothese 2013; 81: 1046–1055.
Toga AW, Thompson PM . Connectopathy in ageing and dementia. Brain 2014; 137: 3104–3106.
Seung S . Coonectome: How the Brain's Wiring Makes Us Who We Are. Houghton Mifflin Harcourt: New York, USA, 2012.
Speed D, O'Brien TJ, Palotie A, Shkura K, Marson AG, Balding DJ et al. Describing the genetic architecture of epilepsy through heritability analysis. Brain 2014; 137: 2680–2689.
Abrahams BS, Geschwind DH . Advances in autism genetics: on the threshold of a new neurobiology. Nat Rev Genet 2008; 9: 341–355.
Cannon TD, Kaprio J, Lonnqvist J, Huttunen M, Koskenvuo M . The genetic epidemiology of schizophrenia in a Finnish twin cohort—A population-based modeling study. Archives Gen Psychiatry 1998; 55: 67–74.
Sullivan PF, Neale MC, Kendler KS . Genetic epidemiology of major depression: Review and meta-analysis. Am J Psychiatry 2000; 157: 1552–1562.
Gatz M, Pedersen NL, Berg S, Johansson B, Johansson K, Mortimer JA et al. Heritability for Alzheimer's disease: the study of dementia in Swedish twins. J Gerontol A Biol Sci Med Sci 1997; 52A: M117–M125.
Gauthier J, Rouleau GA . De novo mutations in neurological and psychiatric disorders: effects, diagnosis and prevention. Genome Med 2012; 4: 71.
Veltman JA, Brunner HG . De novo mutations in human genetic disease. Nat Rev Genet 2012; 13: 565–575.
Consortium E Project EPG. De novo mutations in epileptic encephalopathies. Nature 2013; 501: 1–7.
Iossifov I, O'Roak BJ, Sanders SJ, Ronemus M, Krumm N, Levy D et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature 2014; 515: 216–221.
Fromer M, Pocklington AJ, Kavanagh DH, Williams HJ, Dwyer S, Gormley P et al. De novo mutations in schizophrenia implicate synaptic networks. Nature 2014; 506: 179–184.
Krumm N, O'Roak BJ, Shendure J, Eichler EE . A de novo convergence of autism genetics and molecular neuroscience. Trends Neurosci 2014; 37: 95–105.
O'Roak BJ, Vives L, Girirajan S, Karakoc E, Krumm N, Coe BP et al. Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations. Nature 2012; 485: 246–U136.
Bin Xu, Roos JL, Levy S, Van Rensburg EJ, Gogos JA, Karayiorgou M . Strong association of de novo copy number mutations with sporadic schizophrenia. Nat Genet 2008; 40: 880–885.
Ryvlin P, Cross JH, Rheims S . Epilepsy surgery in children and adults. Lancet Neurol 2014; 13: 1114–1126.
Briellmann RS, Jackson GD, Torn Broers Y, Berkovic SF . Causes of epilepsies: insights from discordant monozygous twins. Ann Neurol 2001; 49: 45–52.
Crino PB . Focal brain malformations: seizures, signaling, sequencing. Epilepsia 2009; 50 (Suppl 9): 3–8.
Lee JH, Huynh M, Silhavy JL, Kim S, Dixon-Salazar T, Heiberg A et al. De novo somatic mutations in components of the PI3K-AKT3-mTOR pathway cause hemimegalencephaly. Nat Genet 2012; 44: 941–945.
Poduri A, Evrony GD, Cai X, Elhosary PC, Beroukhim R, Lehtinen MK et al. Somatic activation of AKT3 causes hemispheric developmental brain malformations. Neuron 2012; 74: 41–48.
Rivière J-B, Mirzaa GM, O'Roak BJ, Beddaoui M, Alcantara D, Conway RL et al. De novo germline and postzygotic mutations in AKT3, PIK3R2 and PIK3CA cause a spectrum of related megalencephaly syndromes. Nat Genet 2012; 44: 934–940.
D’Gama AM, Geng Y, Couto JA, Martin B, Boyle EA, LaCoursiere CM et al. Mammalian target of rapamycin pathway mutations cause hemimegalencephaly and focal cortical dysplasia. Ann Neurol 2015; 77: 720–725.
Jansen LA, Mirzaa GM, Ishak GE, O'Roak BJ, Hiatt JB, Roden WH et al. PI3K/AKT pathway mutations cause a spectrum of brain malformations from megalencephaly to focal cortical dysplasia. Brain 2015; 138: 1613–1628.
Lim JS, Kim W-I, Kang H-C, Kim SH, Park AH, Park EK et al. Brain somatic mutations in MTOR cause focal cortical dysplasia type II leading to intractable epilepsy. Nat Med 2015; 21: 395–400.
Nakashima M, Saitsu H, Takei N, Tohyama J, Kato M, Kitaura H et al. Somatic mutations in the MTOR gene cause focal cortical dysplasia type IIb. Ann Neurol 2015; 78: 375–386.
Di Martino A, Yan CG, Li Q, Denio E, Castellanos FX, Alaerts K et al. The autism brain imaging data exchange: towards a large-scale evaluation of the intrinsic brain architecture in autism. Mol Psychiatry 2014; 19: 659–667.
O'Roak BJ, Vives L, Girirajan S, Karakoc E, Krumm N, Coe BP et al. Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations. Nature 2012; 485: 246–250.
Neale BM, Kou Y, Liu L, Ma'ayan A, Samocha KE, Sabo A et al. Patterns and rates of exonic de novo mutations in autism spectrum disorders. Nature 2012; 485: 242–U129.
Valk SL, Di Martino A, Milham MP, Bernhardt BC . Multicenter mapping of structural network alterations in autism. Hum Brain Mapp 2015; 36: 2364–2373.
Casanova MF, El-Baz AS, Kamat SS, Dombroski BA, Khalifa F, Elnakib A et al. Focal cortical dysplasias in autism spectrum disorders. Acta Neuropathol Commun 2013; 1: 67.
Stoner R, Chow ML, Boyle MP, Sunkin SM, Mouton PR, Roy S et al. Patches of disorganization in the neocortex of children with autism. N Engl J Med 2014; 370: 1209–1219.
Wegiel J, Kuchna I, Nowicki K, Imaki H, Wegiel J, Marchi E et al. The neuropathology of autism: defects of neurogenesis and neuronal migration, and dysplastic changes. Acta Neuropathol 2010; 119: 755–770.
D’Gama AM, Pochareddy S, Li M, Jamuar SS, Reiff RE, Lam A-TN et al. Targeted DNA sequencing from autism spectrum disorder brains implicates multiple genetic mechanisms. Neuron 2015; 88: 910–917.
Sequeira A, Martin MV, Rollins B, Moon EA, Bunney WE, Macciardi F et al. Mitochondrial mutations and polymorphisms in psychiatric disorders. Front Gene 2012; 3: 103.
Kato T, Iwamoto K, Kakiuchi C, Kuratomi G, Okazaki Y . Genetic or epigenetic difference causing discordance between monozygotic twins as a clue to molecular basis of mental disorders. Mol Psychiatry 2005; 10: 622–630.
Consortium C. Sparse whole-genome sequencing identifies two loci for major depressive disorder. Nature 2015; 523: 588–591.
Consortium SWGOTPG. Biological insights from 108 schizophrenia-associated genetic loci. Nature 2014; 511: 421–427.
Dempster EL, Pidsley R, Schalkwyk LC, Owens S, Georgiades A, Kane F et al. Disease-associated epigenetic changes in monozygotic twins discordant for schizophrenia and bipolar disorder. Hum Mol Gen 2011; 20: 4786–4796.
MacDonald AW III, Carter CS, Kerns JG . Specificity of prefrontal dysfunction and context processing deficits to schizophrenia in never-medicated patients with first-episode psychosis. Am J Psychiatry 2005; 162: 475–484.
Murray EA, Wise SP, Drevets WC . Localization of dysfunction in major depressive disorder: prefrontal cortex and amygdala. Biol Psychiatry 2011; 69: e43–e45.
Bundo M, Toyoshima M, Okada Y, Akamatsu W, Ueda J, Nemoto-Miyauchi T et al. Increased L1 retrotransposition in the neuronal genome in schizophrenia. Neuron 2014; 81: 306–313.
Kim J, Shin J-Y, Kim J-I, Seo J-S, Webster MJ, Lee D et al. Somatic deletions implicated in functional diversity of brain cells of individuals with schizophrenia and unaffected controls. Sci Rep 2014; 4: 3807.
Loy CT, Schofield PR, Turner AM, Kwok JB . Genetics of dementia. Lancet 2014; 383: 828–840.
Pamphlett R . The ‘somatic-spread’ hypothesis for sporadic neurodegenerative diseases. Med Hypothese 2011; 77: 544–547.
Serrano-Pozo A, Frosch MP, Masliah E, Hyman BT . Neuropathological alterations in alzheimer disease. Cold Spring Harb Perspect Med 2011; 1: a006189–a006189.
Brundin P, Melki R, Kopito R . Prion-like transmission of protein aggregates in neurodegenerative diseases. Nat Rev Mol Cell Biol 2010; 11: 301–307.
Frigerio CS, Lau P, Troakes C, Deramecourt V, Gele P, Van Loo P et al. On the identification of low allele frequency mosaic mutations in the brains of Alzheimer disease patients. Alzheimers Dement 2015; 11: 1265–1276.
Parcerisas A, Rubio SE, Muhaisen A, Gómez-Ramos A, Pujadas L, Puiggros M et al. Somatic signature of brain-specific single nucleotide variations in sporadic Alzheimer's disease. J Alzheimers Dis 2014; 42: 1357–1382.
Jamuar SS, Lam A-TN, Kircher M, D’Gama AM, Wang J, Barry BJ et al. Somatic mutations in cerebral cortical malformations. N Engl J Med 2014; 371: 733–743.
Shirley MD, Tang H, Gallione CJ, Baugher JD, Frelin LP, Cohen B et al. Sturge–Weber Syndrome and Port-Wine Stains caused by somatic mutation in GNAQ. N Engl J Med 2013; 368: 1971–1979.
Schirmer M, Ijaz UZ, D'Amore R, Hall N, Sloan WT, Quince C . Insight into biases and sequencing errors for amplicon sequencing with the Illumina MiSeq platform. Nucleic Acids Res 2015; 43: e37.
Costello M, Pugh TJ, Fennell TJ, Stewart C, Lichtenstein L, Meldrim JC et al. Discovery and characterization of artifactual mutations in deep coverage targeted capture sequencing data due to oxidative DNA damage during sample preparation. Nucleic Acids Res 2013; 41: e67–e67.
GLENN TC . Field guide to next‐generation DNA sequencers. Mol Ecol Resour 2011; 11: 759–769.
Robasky K, Lewis NE, Church GM . The role of replicates for error mitigation in next-generation sequencing. Nat Rev Genet 2013; 15: 56–62.
Shapiro E, Biezuner T, Linnarsson S . Single-cell sequencing-based technologies will revolutionize whole-organism science. Nat Rev Genet 2013; 14: 618–630.
de Bourcy CFA, De Vlaminck I, Kanbar JN, Wang J, Gawad C, Quake SR . A quantitative comparison of single-cell whole genome amplification methods. PLoS ONE 2014; 9: e105585.
Voet T, Van Loo P . SNES makes sense? Single-cell exome sequencing evolves. Genome Biol 2015; 16: 86.
Cibulskis K, Lawrence MS, Carter SL, Sivachenko A, Jaffe D, Sougnez C et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat Biotechnol 2013; 31: 213–219.
Kim S, Jeong K, Bhutani K, Lee J, Patel A, Scott E et al. Virmid: accurate detection of somatic mutations with sample impurity inference. Genome Biol 2013; 14: R90.
Saunders CT, Wong WSW, Swamy S, Becq J, Murray LJ, Cheetham RK . Strelka: accurate somatic small-variant calling from sequenced tumor-normal sample pairs. Bioinformatics 2012; 28: 1811–1817.
Xu H, DiCarlo J, Satya RV, Peng Q, Wang Y . Comparison of somatic mutation calling methods in amplicon and whole exome sequence data. BMC Genomics 2014; 15: 1.
Kim J, Kim S, Nam H, Kim S, Lee D . SoloDel: a probabilistic model for detecting low-frequent somatic deletions from unmatched sequencing data. Bioinformatics 2015; 31: btv358–3113.
Wilm A, Aw PPK, Bertrand D, Yeo GHT, Ong SH, Wong CH et al. LoFreq: a sequence-quality aware, ultra-sensitive variant caller for uncovering cell-population heterogeneity from high-throughput sequencing datasets. Nucleic Acids Res 2012; 40: gks918–11201.
Kim H, Kim J-S . A guide to genome engineering with programmable nucleases. Nat Rev Genet 2014; 15: 321–334.
Sander JD, Joung JK . CRISPR-Cas systems for editing, regulating and targeting genomes. Nat Biotechnol 2014; 32: 347–355.
Hsu PD, Lander ES, Zhang F . Development and applications of CRISPR-Cas9 for genome engineering. Cell 2014; 157: 1262–1278.
Wang T, Wei JJ, Sabatini DM, Lander ES . Genetic screens in human cells using the CRISPR/Cas9 system. Science 2014; 343: 80–84.
Shalem O, Sanjana NE, Hartenian E, Shi X, Scott DA, Mikkelsen TS et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 2014; 343: 84–87.
Sánchez-Rivera FJ, Papagiannakopoulos T, Romero R, Tammela T, Bauer MR, Bhutkar A et al. Rapid modelling of cooperating genetic events in cancer through somatic genome editing. Nature 2014; 516: 428–431.
Xue W, Chen S, Yin H, Tammela T, Papagiannakopoulos T, Joshi NS et al. CRISPR-mediated direct mutation of cancer genes in the mouse liver. Nature 2000; 514: 380–384.
Swiech L, Heidenreich M, Banerjee A, Habib N, Li Y, Trombetta J et al. In vivo interrogation of gene function in the mammalian brain using CRISPR-Cas9. Nat Biotechnol 2015; 33: 102–106.
Zuckermann M, Hovestadt V, Knobbe-Thomsen CB, Zapatka M, Northcott PA, Schramm K et al. Somatic CRISPR/Cas9-mediated tumour suppressor disruption enables versatile brain tumour modelling. Nat Commun 2015; 6: 7391.
Babiloni F, Cincotti F, Babiloni C, Carducci F, Mattia D, Astolfi L et al. Estimation of the cortical functional connectivity with the multimodal integration of high-resolution EEG and fMRI data by directed transfer function. Neuroimage 2005; 24: 118–131.
Guye M, Bartolomei F, Ranjeva J-P . Imaging structural and functional connectivity: towards a unified definition of human brain organization? Curr Opin Neurobiol 2008; 21: 393–403.
Einevoll GT, Franke F, Hagen E, Pouzat C, Harris KD . Towards reliable spike-train recordings from thousands of neurons with multielectrodes. Curr Opin Neurobiol 2012; 22: 11–17.
Oh SW, Harris JA, Ng L, Winslow B, Cain N, Mihalas S et al. A mesoscale connectome of the mouse brain. Nature 2014; 508: 207–214.
Stosiek C, Garaschuk O, Holthoff K, Konnerth A . In vivo two-photon calcium imaging of neuronal networks. Proc Natl Acad Sci USA 2003; 100: 7319–7324.
Mikula S, Denk W . High-resolution whole-brain staining for electron microscopic circuit reconstruction. Nat Methods 2015; 12: 541–546.
Acknowledgements
I thank Prof. Eunjoon Kim, Prof Jae Myung Suh, Jang Keun Kim, Myeong-Heui Kim at KAIST, Prof Hyongbum Kim and Prof Sangwoo Kim at the Yonsei University College of Medicine for their helpful comments on this manuscript, as well as Jae Seok Lim at KAIST for sharing unpublished data and helping with the art work. This work was supported by grants from the Korean Health Technology Research and Development (R&D) Project, Ministry of Health & Welfare, Republic of Korea (A121070 & HI13C0208), the Brain Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Science, Information and Communication Technology (ICT) & Future Planning (2013M3C7A1056564), the KAIST Future Systems Healthcare Project from the Ministry of Science, ICT and Future Planning and the Korea Institute of Science and Technology Information (K-15-L03-C02-S01).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares no conflict of interest.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Lee, J. Somatic mutations in disorders with disrupted brain connectivity. Exp Mol Med 48, e239 (2016). https://doi.org/10.1038/emm.2016.53
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/emm.2016.53
This article is cited by
-
Human glioblastoma arises from subventricular zone cells with low-level driver mutations
Nature (2018)
-
Somatic mutations in neurons during aging and neurodegeneration
Acta Neuropathologica (2018)