Abstract
MicroRNAs (miRNAs) participate in diverse biological functions and carcinogenesis by inhibiting specific gene expression. We previously reported that suppression of adenine nucleotide translocase 2 (ANT2) by using the short hairpin RNA (shRNA) approach has an antitumor effect in several cancer cells. We here examined the influence of ANT2 on expression of miRNAs in hepatocellular carcinoma (HCC) to further elucidate the tumor-suppressive mechanism of ANT2 shRNA. We first carried out screening for miRNAs, whose expression is regulated by ANT2 suppression in the Hep3B HCC cell line using miRNA microarrays. Validation of candidate miRNAs was done by incorporating clinical samples, and their effects on the tumorigenesis of HCC were studied in vitro and in vivo. miR-636 was one of the miRNAs whose expression was highly upregulated by ANT2 suppression in miRNA microarray analysis, as confirmed by real-time reverse transcription-polymerase chain reaction. Notably, miR-636 was markedly downregulated in HCC tissues compared with matched non-neoplastic liver in clinical samples. Restoration of miR-636 in Hep3B cells led to significant reduction of cell proliferation and colony formation. miR-636 restoration resulted in a decreased level of Ras, one of the putative targets of miR-636, and inactivation of its signaling pathway. Moreover, tumorigenesis was efficiently suppressed by miR-636 in an in vivo tumor xenograft model of HCC. The data suggest that miR-636 might function as a tumor suppressor miRNA affecting HCC tumorigenesis via downregulation of Ras, and that ANT2 suppression by shRNA could exert an anticancer effect by restoring miR-636 expression in HCC.
Similar content being viewed by others
Introduction
Cancer is affected by many molecular pathways involving both canonical protein-coding genes and recently discovered noncoding genes. MicroRNAs (miRNAs), a class of small RNAs 17–27 nucleotides in length, are representative molecules transcribed from noncoding genes, and control gene expression at the level of mRNA translation.1 An aberrant miRNA expression signature is a hallmark of several diseases, including cancer,2 and miRNAs can function as oncogenes or tumor suppressors.3 Moreover, these tumor-associated miRNAs can serve as biomarkers for tumor diagnosis and prognosis or as therapeutic targets.4 Several recent studies have reported that miRNA expression is deregulated in human hepatocellular carcinoma (HCC) in comparison with matched non-neoplastic tissue.5, 6 The downregulated miRNAs in HCC include several members of the large let-7 family and the miR-143/145.7 Although for members of the let-7 family the results have been contentious, these miRNAs were found to be consistently downregulated in several human cancers.8 Among the downregulated miRNAs in HCC is the hepato-specific miR-122; downregulation of miR-122 may have a more direct role in tumorigenesis through the activation of cyclin G1.9
The Ras signaling pathway controls cell proliferation, differentiation and survival in all multicellular organisms.10, 11, 12, 13 Consequently, proper regulation of Ras signaling is critically important for normal development, and dysregulation of the Ras pathway leads to pathologic diseases.14, 15, 16 Indeed, constitutively active Ras promotes cell proliferation, neoplastic transformation and tumorigenesis.14, 17, 18 Oncogenic Ras proteins are commonly detected in human cancers, including ∼90% of pancreatic cancers, ∼70% of malignant neoplasias and ∼30% of all human cancers.19 Despite the central importance of Ras signaling in human cancer, direct inhibition of hyperactivated Ras has not achieved sufficient clinical efficacy.15, 18, 20 We reasoned that downregulation of Ras by the specific miRNA might provide a more effective strategy for inhibiting oncogenic Ras signaling and Ras-mediated tumorigenesis in human cancer.
Adenine nucleotide translocase (ANT) is a nuclear-encoded protein abundantly located in the inner mitochondrial membrane, and the role of this protein is to catalyze the exchange of mitochondrial adenosine triphosphate with cytosolic adenosine diphosphate. ANT therefore plays an important role in cellular energy metabolism by influencing mitochondrial oxidative phosphorylation. In addition, ANT is the major component of the mitochondrial permeability–transition pore complex that interacts with Bcl2 family proteins, thereby contributing to mitochondria-mediated apoptosis.21, 22 ANT-deficient mice are able to form mitochondrial permeability–transition pore complex, which has prompted discussion regarding the roles of ANT in mitochondrial permeability–transition pore complex.23 Human ANT has four isoforms (ANT1, ANT2, ANT3 and ANT4) and the relative expressions of these isoforms are dependent on developmental stages, proliferation status, as well as tissue types or cell types.14, 15 ANT2 is specifically expressed in undifferentiated cells or tissues that are able to proliferate and regenerate; for example, lymphocytes, kidney and liver. The expression of ANT2 was recently found to be upregulated in several hormone-dependent cancers, and the induction of ANT2 expression in cancer cells was shown to be directly associated with glycolytic metabolisms, raising a question regarding the role of ANT2 during carcinogenesis.24, 25, 26, 27 Therefore, ANT2 repression effectively leads to cell growth arrest and increases mitochondrial membrane potential from human cells as well as chemosensitized cancer cells, implying that ANT2 acts as an antiapoptotic oncoprotein.17, 28, 29 Thus, ANT2 could be a promising candidate for cancer therapy based on specific miRNA regulation.
We hypothesized that upregulation of a specifc miRNA by ANT2 short hairpin RNA (shRNA) may suppress HCC tumorigenesis. Presently, miR-636 miRNA expression was highly upregulated by ANT2 suppression. miR-636 restoration resulted in a decrease in level of Ras, one of the putative targets of miR-636, and inactivation of its signaling pathway. The data suggest that miR-636 might function as a tumor suppressor miRNA affecting HCC tumorigenesis via downregulation of Ras, and that ANT2 suppression by shRNA could exert an anticancer effect by restoring miR-636 expression in HCC.
Therefore, the purpose of our study was to determine whether or not ANT2 shRNA-mediated miR-636 restoration suppressed HCC cell proliferation, and to investigate the potential of miR-636 for future use as a therapeutic agent targeting HCC cells.
Materials and methods
Cell lines and culture
The human hepatoma cell line, Hep3B, HepG2 and SNU449 were purchased from the American Type Culture Collection (ATCC, Manassas, VA, USA) or Korean Cell Line Bank (KCLB, Seoul, Korea) and cultured in Dulbecco’s Modified Eagle Medium supplemented with 10% fetal bovine serum, 100 U ml−1 penicillin, and 100 μg ml−1 streptomycin (Gibco, Grand Island, NY, USA) in a humidified 5% CO2/95% air atmosphere at 37 °C.
Antibodies and reagents
Anti-phospho-Akt and anti-Akt antibodies were purchased from Cell Signaling Technology (Beverly, MA, USA).
Construction of ANT2 shRNA expression vector
The ANT2 shRNA expression vector used to achieve specific downregulation of ANT2 was described previously.30 ANT2 small-interfering RNA (5′-GCAGAUCACUGCAGAUAAGTT-3′) that was designed to be complementary to exon 2 of ANT2 (GenBank accession number NM001152) was synthesized, and DNA vectors expressing the shRNA forms of the small-interfering RNAs were generated using pSilencer 3.1-H1 puro plasmids with a TTCAAGAGA linker sequence that forms looped structures (Ambion, Austin, TX, USA). The vector expressing ANT2 shRNA was used throughout this study. A scramble shRNA (Ambion) with no significant homology to human gene sequences was used as a control to detect nonspecific effects.
Transfection
For transfection, cells were plated on either six-well plates (2 × 105 cells per well) or 100-mm dishes (2 × 106 cells), and allowed to adhere for 24 h. Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) was used for the transfection. Cells were transfected with either pSilencer 3.1-H1 puro ANT2 shRNA (Ambion) or pSilencer 3.1-H1 puro scramble shRNA vectors (Ambion). Transfected cells were cultured for 6 h, whereupon the medium was replaced with fresh medium supplemented with 10% FBS. The cells were harvested 24–48 h after transfection. Negative mimics, miR-636 mimics, negative inhibitor or miR-636 inhibitor was transfected into the cells using the same method.
miRNAs microarray statistical analysis
Agilent miRNA assays (Agilent Technologies Inc, Santa Clara, CA, USA) integrate eight individual microarrays on a single glass slide. Each microarray includes approximately 15 000 features containing probes sourced from the miRBASE public database. The probes are 60-mer oligonucleotides that are directly synthesized on the array. In this study we used the human microRNA microarray v2.0, which contains 723 human and 76 human viral miRNAs, each replicated 16 times. Three hundred sixty-two miRNAs are interrogated by two different oligonucleotides, 45 miRNAs by three and 390 miRNAs by four. Only two miRNAs are interrogated by a single oligonucleotide. The array also contains a set of positive and negative controls that are replicated a variety of times. Some of the positive control probes target non-miRNA human RNAs. Each of these targets was interrogated with four different probes, with five replications. The signals from these positive controls can be bright or dim depending on the sample and, according to Agilent, they do not behave consistently enough to be used for normalization.
Quantitative real-time RT–PCR (RT–qPCR)
For RT–qPCR, total RNAs from the same samples used in the miRNA microarrays were tested by using an iCyler IQ real-time PCR system (Bio-Rad, Hercules, CA, USA). Results are shown as fold-change. RT–qPCR was carried out with Mir-X miRNA First-Strand Synthesis and SYBR Green Real time PCR Master Mix (TaKaRa, Otsu, Japan) according to the manufacturer's instructions. The housekeeping gene U6 was used for standardization of the initial miRNA content of a sample. Relative changes of gene expression were calculated by the following formula, and the data were represented as fold upregulation/downregulation: fold change= , where ΔΔCt=(Ct of gene of interest, treated−Ct of HK gene, treated)−(Ct of gene of interest, control−Ct of HK gene, control), Ct is the threshold cycle number and HK is the housekeeping gene.
, where ΔΔCt=(Ct of gene of interest, treated−Ct of HK gene, treated)−(Ct of gene of interest, control−Ct of HK gene, control), Ct is the threshold cycle number and HK is the housekeeping gene.
Western blotting
For western blot analysis, cells were harvested 24 or 48 h after transfection and lysed with lysis buffer consisting of 5 mM l−1 ethylenediamine tetraacetic acid, 300 mM l−1 NaCl, 0.1% NP-40, 0.5 mM l−1 NaF, 0.5 mM l−1 Na3VO4, 0.5 mM l−1 phenylmethylsulfonyl fluoride, and 10 μg ml−1 each of aprotinin, pepstatin and leupeptin (Sigma-Aldrich, St Louis, MO, USA). Following centrifugation at 15 000 g for 30 min, the concentrations of supernatant proteins were analyzed using the Bradford reagent (Bio-Rad). For analysis of protein contents, 50 μg total protein was subjected to 10% SDS-PAGE and the resolved proteins were transferred to polyvinylidene difluoride membranes (Millipore, Bedford, MA, USA), which were incubated with the antibodies indicated above. Immunoblots were visualized using an enhanced chemiluminescence detection system (Amersham Pharmacia Biotech, Uppsala, Sweden).
Cell proliferation assay
Cell proliferation was measured using MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide) (Sigma-Aldrich). The reduction of tetrazolium salts is now widely accepted as a reliable way to examine cell proliferation. The yellow tetrazolium MTT is reduced by metabolically active cells, in part by the action of dehydrogenase enzymes, to generate reducing equivalents such as nicotinamide adenine dinucleotide and nicotinamide adenine dinucleotide phosphate. The resulting intracellular purple formazan can be solubilized and quantified spectrophotometrically. Results of cell proliferation assays are presented as the mean values of three replicate experiments performed in triplicate.
Luciferase assay
Hep3B cells were transfected with miR-636 or a negative control (miR control), each of concentration 100 nM, for 6 h in reduced serum and antibiotics-free Opti-MEM with Lipofectamine 2000. After 24 h, cells were transfected with Ras 3′ untranslated region (UTR)-luciferase expression vector. Firefly luciferase was measured in cell lysates using a Signosis Biosignal Capture Luc-Screen System on a Fusion plate reader (Perkin-Elmer, Waltham, MA, USA). Firefly luciferase activity was used for normalization and as an internal control for transfection efficiency.
Soft agar colony formation assay
Cells transfected with miR-636 or a negative control (miR control) were plated in triplicate in the appropriate medium containing 0.3% Bacto-agar on top of 0.6% bottom agar and cultured for 2 weeks. Colonies were visualized by staining with 0.1% p-iodonitro tetrazolium violet (Sigma-Aldrich). Monolayer colonies were stained with crystal violet dye after methanol fixation.
Tumorigenicity assays in nude mice
For in vivo tumorigenicity assay, negative mimics (negative control) or miR-636 mimic-transfected Hep3B cells (2 × 105) were suspended in 100 μl phosphate-buffered saline and then injected subcutaneously into the right or left side of the posterior flank of six BALB/c nude mice, respectively. Tumor growth was examined daily and the tumor volumes were calculated every week using the formula for hemi-ellipsoids: V=length (cm) × width (cm) × height (cm) × 0.5236. After 5 weeks, the mice were killed and the tumors were dissected and photographed.
Statistical analysis
GraphPad Prism software (Brothersoft, Beijing, China) was used to conduct statistical analysis. Results are expressed as the mean±s.e.m. For P-value calculation, unpaired Student’s t-tests (two tailed) were utilized for all comparisons. P-values <0.05 were considered as significant.
Results
ANT2 shRNA induces upregulation of miR-636 in Hep3B HCC cells
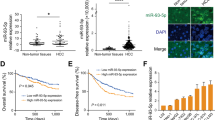
miRNA microarray screenings were done to identify ANT2 shRNA-regulated miRNAs in Hep3B HCC cells (Figure 1a). Among the 887 human miRNAs on the microarray, 8 miRNAs were upregulated, and 14 miRNAs were downregulated, with a greater than twofold difference between sc shRNA and ANT2 shRNA transfection in Hep3B. As a result, we obtained significant candidate miRNAs and reaffirmed by selecting a specific miRNA, miR-636, among the miRNAs. As shown in Figure 1b, miR-636 was upregulated by ANT2 shRNA. Furthermore, we confirmed this result in the other HCC cell lines, HepG2 and SNU449 (Supplementary Data 1). However, the functional attributes of miR-636 associated with liver tumor progression have not been experimentally established. Moreover, among candidate miRNAs, only miR-636 expression in HCC tissues was significantly lower than that in non-neoplastic liver tissue.
Knockdown of adenine nucleotide translocase 2 (ANT2) by short hairpin RNA (shRNA) induces specific restoration of miR-636 in Hep3B and expression of miR-636 was remarkably lower in hepatocellular carcinoma (HCC) tissue. (a) RNA extraction was performed 24 h after transfection of Hep3B cells with ANT2 shRNA for microRNA (miRNA) microarray analysis. The arrays were progressed using an Agilent scanner and microarray data extracted with Agilent Feature Extraction software. (b) Quantitative real-time RT–PCR (RT-qPCR) analysis of the effect of ANT2 shRNA-induced restoration on miR-636 expression. Cells were transfected with scramble or ANT2 shRNA. Total RNA was extracted 24 h after transfection and subjected to RT-qPCR using specific primers for human miR-636 or U6 (internal control). (c) RT–qPCR of 20 HCC tumor tissue pairs was conducted. RNAs extraction from the paraffin block was used by the Ambion mirVana miRNA isolation kit and subjected to RT–qPCR using specific primers for human miR-636 or U6 (internal control).
Downregulation of miR-636 in HCC compared with non-neoplastic liver tissue
To see if miR-636 might have some biological relevance in HCC, RT–qPCR of 20 HCC tumor tissue pairs was conducted. Expression of miR-636 was markedly lower in HCC tissue compared with matched non-neoplastic liver tissue (Figure 1c), which implied that the downregulation of miR-636 might be associated with tumorigenesis. We performed Mann–Whitney test and χ2 test for a two-sample test of miR-636 expression levels. For multiple-testing adjustment, there was a significant difference of mean miR expression level between the two groups only if the P-value was <0.005. As ANT2 is highly expressed in proliferating cells, and ANT2 induction in cancer cells is directly associated with carcinogenesis,25, 26, 27 ANT2 was enhanced in HCC tissue compared with non-neoplastic liver tissue (data not shown).
Restoration of miR-636 by ANT2 shRNA downregulates Ras gene transcripts, in which Ras is a critical downstream target of miR-636
K-Ras or H-Ras mRNAs were significantly decreased in cells with overexpression of miR-636 or cells in which ANT2 expression was knockdown (Figure 2a). Bioinformatic prediction recognized 1–7 miR-636 binding sites in the 3′UTRs of the Ras gene. To test whether the predicted binding sites in the target genes could mediate repression of translation by miR-636 (Figure 2b). Ras 3′UTRs were subcloned downstream of the luciferase gene. The reporter with the 3′UTR of Ras showed a significantly lower luciferase activity in overexpression of miR-636 or knockdown of ANT2 cells (Figure 2b). The result was confirmed in the HepG2 and SNU449 HCC cell lines (Supplementary Data 2). Taken together, the evidence demonstrates that Ras is a critical downstream target of miR-636.
miR-636 regulates Ras expression at the post-transcriptional level and restoration of miR-636 by adenine nucleotide translocase 2 (ANT2) short hairpin RNA (shRNA) suppresses Ras/phosphoinositide 3-kinase/Akt signaling. (a) After 24 h of transfection with negative/miR-636 mimics or scramble/ANT2 shRNA, total RNA was extracted and subjected to RT-qPCR using specific primers for human K-Ras, H-Ras or glyceraldehyde 3-phosphate dehydrogenase as the internal control. (b) Human embryonic kidney 293 cells were transiently co-transfected with Ras–3′ untranslated region-luciferase reporter gene vectors and then stimulated with negative/miR-636 mimics or scramble/ANT2 shRNA for 18 h. Luciferase activities in cell extracts were analyzed by dual-luciferase reporter assay system and normalized using pRL-TK-luciferase activity (Renilla luciferase activity) in each sample. Data present the mean (fold increase in luciferase activity compared with cells stimulated with negative mimics or scrambled shRNA) ±s.d. of triplicates. (c) After 24 h of transfection with negative or miR-636 mimics, cell extracts were prepared for western blotting with antiphospho-Akt/anti-Akt antibodies or antiphospho-Erk/anti-Erk antibodies. (d) Hep3B cells were transfected with scrambled/ANT2 shRNA and/or negative/miR-636 inhibitor, and cell extracts were prepared for western blotting with antiphospho-Akt/anti-Akt antibody or antiphospho-Erk/anti-Erk antibody.
Upregulation of miR-636 by ANT2 shRNA inactivates the Ras/phosphoinositide 3-kinase (PI3K)/Akt signaling pathway
Restoration of miR-636 by ANT2 shRNA downregulates Ras transcripts and inactivates the Ras/PI3K/Akt signaling pathway. Ras protein is among the best-characterized proto-oncogenes and Ras signaling regulates diverse cellular functions, including cell growth, survival and migration. With regard to the regulation of Ras/Mek/Erk or Ras/PI3K/Akt activity, we found that phosphorylated Akt levels were decreased by miR-636 mimics but not phosphorylated Erk levels (Figure 2c). Furthermore, we confirmed this result in the other HCC cell lines, HepG2 and SNU449 (Supplementary Data 3). Moreover, we confirmed that phosphorylated Akt levels were decreased by knockdown of ANT2 by shRNA and slightly recovered by miR-636 inhibitor in Figure 2d. Taken together, these findings suggested that ANT2 shRNA treatment led to restoration of miR-636 expression in Hep3B cells, accompanied by suppression of Ras/PI3K/Akt signaling. It is well known that Akt plays important roles as a mediator of Ras-mediated cell survival and proliferation; as a result, inhibition of Akt by Akt-specific inhibitor also inhibits cell proliferation of Hep3B (Supplementary Data 4).
Upregulation of miR-636 by ANT2 shRNA inhibits cell proliferation and suppresses colony formation
With regard to the biological functions of miR-636, we observed that restoration of miR-636 expression suppresses HCC cell proliferation. The knockdown of ANT2 and inhibition of miR-636 inhibited cell proliferation of Hep3B (Figure 3a). Overexpression of miR-636 by miR-636 mimics (Supplementary Data 5) or ANT2 knockdown by ANT2 shRNA also inhibited cell proliferation of Hep3B.30 Overexpression of miR-636 also impaired anchorage-independent growth in soft agar, significantly reducing the number and size of the colonies. Overexpression of miR-636 also reduced the proliferation of Hep3B cells (Figure 3b). Moreover, there was significant reduction in cell proliferation as well as lower anchorage-independent growth in soft agar (Figure 3c).
Restoration of miR-636 by adenine nucleotide translocase 2 (ANT2) short hairpin RNA (shRNA) inhibits cell proliferation and upregulation of miR-636 by ANT2 shRNA suppresses colony formation. (a) Hep3B cells were transfected with scramble, ANT2 shRNA, negative or miR-636 inhibitor. Cell viability (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT)) assays were performed 24 h after transfection. Three independent experiments were performed. Data were analyzed using the Student’s t-test. (b) Hep3B cells were transfected with negative or miR-636 mimics. Cell viability (MTT) assays were performed 24, 48 and 72 h after transfection. Three independent experiments were performed. Data were analyzed using the Student’s t-test. (c) Colony formation after transfection of Hep3B cells with negative or miR-636 mimics.
Overexpression of miR-636 suppresses in situ tumor formation
We determined whether miR-636 and its target gene are associated with tumorgenesis. miR-636 mimics transfected Hep3B cells implanted subcutaneously into the flanks of nude mice. Overexpression of miR-636 significantly suppressed overall tumor growth, as assessed by tumor volume (Figure 4). Thus, miR-636 would seem to regulate tumorigenesis, at least in part through its target Ras. Ras expression was significantly decreased in miR-636-overexpressed tumor tissue (data not shown). Consistent with the in vitro results, overexpression of miR-636 Hep3B cells significantly suppressed overall tumor growth in vivo. These in vitro and in vivo results imply that the gradual loss of miR-636 over a long period of time is likely to commit hepatocytes to tumorigenesis.
Overexpression of miR-636 suppresses in situ tumor formation. Balb/c nude mice were challenged with negative or miR-636 mimic-transfected 2 × 105 Hep3B cells by subcutaneous injection into the left or right flanks. Tumor sizes were measured by a caliper every week and volumes calculated using the formula: m12 × m2 × 0.5236, where m1 represents the short tumor axis and m2 the long axis, until day 73 following tumor challenge.
Discussion
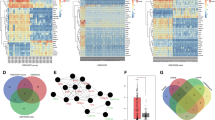
Microarrays have been used in several studies of miRNA expression profiling in HCC. One of the earliest works to detect the relationship between miRNAs and HCC31 reported that three miRNAs were expressed more exuberantly in HCC samples than in nontumorous tissues, while five miRNAs had lower expression in HCC tissue. The prediction accuracy of classifying HCC and nontumorous tissues with eight miRNAs was 97.8%. Another study reported that 18 miRNAs were overexpressed in HCC and 6 miRNAs were overexpressed in nontumorous samples.32 Aberrant expression of miR-21 contributes to progression and metastasis of HCC, and miR-21 can participate in the process of HCC development by targeting phosphatase and tensin homolog (PTEN33). Another study used a supervised algorithm to predict metastasis characterization in the examination of miRNA expression profiles of 482 cancerous and noncancerous specimens from radical resection of 241 patients with HCC.34 In comparative miRNA expression studies, several miRNAs that are important in HCC initiation, development and metastasis, such as miR-21, have been identified. Aberrant expression of miR-21 can contribute to HCC growth and spread by modulating PTEN expression and PTEN-dependent pathways involved in mediating the phenotypic characteristics of cancer cells, such as cell growth, migration and invasion.33
The function of miR-636 has yet not been studied in cancer cells. The only reports on miR-636 describe miR-636 as a potential candidate responsible for the post-transcriptional silencing of GR in GC-resistant cells.35 For the first time, our study suggests that miR-636 is a tumor suppressor miRNA affecting HCC tumorigenesis via suppression of the Ras signaling pathway. Restoration of miR-636 efficiently suppressed Hep3B cell proliferation in vitro and tumorigenesis in vivo at a level comparable to ANT2 shRNA treatment.
Physiological and oncogenic activation of Ras stimulates a wide range of downstream signaling pathways. The first Ras effector pathway to be identified was the Raf–Mek–Erk pathway.36, 37, 38, 39 This pathway is an essential shared element of mitogenic signaling involving tyrosine kinase receptors, leading to a wide range of cellular responses, including growth, differentiation, inflammation and apoptosis.40 The second best-characterized Ras effector family is PI3Ks, which play important roles as mediators of Ras-mediated cell survival and proliferation.41, 42 PI3Ks are important regulators of cellular growth, transformation, adhesion, apoptosis, survival and motility.43, 44, 45, 46 Ras-dependent PI3K activation is linked to prosurvival signaling due to the activation of Akt. Our findings that inactivated PI3K or Akt induces apoptosis and that dominant-negative Akt has the ability to enhance apoptosis highlight the importance of PI3K/Akt signaling in promoting survival.41, 47, 48, 49 Moreover, our results demonstrate the importance of Ras–PI3K interaction in Ras-driven tumorigenesis. Although multiple Ras effectors are essential to initiate tumor formation, only signaling through the PI3K/Akt pathway seems to be necessary to maintain tumor growth. The reduction of Ras oncogene by miR-636 induces inactivation of the PI3K/Akt pathway and inhibits tumor growth. Tumor maintenance in Ras-driven tumor growth is required for PI3K signaling.
In summary, the deduced expression profiles in HCC cell lines show that miR-636 can act as a tumor suppressor through the direct control of Ras expression. The restoration of miR-636 by ANT2 shRNA contributes to suppression of tumor formation. ANT2 suppression by shRNA might be able to exert an anticancer effect in HCC by regulation of tumor suppressor miRNAs.
Accession codes
References
Rosa V, Carlo CM . MiRNAs and cancer. Am J Pathol 2009; 174: 1131–1138.
Iorio MV, Croce CM . Micro RNA involvement in human cancer. Carcinogenesis 2012; 33: 1126–1133.
Baohong Z, Xiaoping P, George PC, Todd AA . Micro RNAs as oncogenes and tumor suppressors. Dev Biol 2007; 302: 1–12.
Scott AW, Andr T . MicroRNA signatures as diagnostic and therapeutic targets. Clin Chem 2008; 54: 943–944.
Laura G, Francesca F, Elisa C, Silvia S, Giovanni L, Carlo MC et al. MicroRNA involvement in hepatocellular carcinoma. Cell Mol Med 2008; 12: 2189–2204.
Manuela F, Sylwia EW, Ramiro G, Hansjuerg A, Ramana D, Chang-Gong L et al. A microRNA signature of Hypoxia. Mol Cell Biol 2007; 27: 1859–1867.
Panagiota C, Katerina V, Ioannis S, Christos P, Evangelos B . Alterations of microRNAs in solid cancers and their prognostic value. Cancers 2010; 2: 1328–1353.
Benjamin B, Park SM, Annika H, Andrea EM, Marcus EP, Endocr R . The role of let-7 in cell differentiation and cancer. Cancer 2010; 17: F19–F36.
Jun X, Yaxing X, Junli H, Saifeng W, Changfei L, Songdong M . MiR-122 in hepatic function and liver diseases. Protein Cell 2012; 3: 364–371.
Bollag G, McCormick F . Regulators and effectors of ras proteins. Annu Rev Cell Biol 1991; 7: 601–632.
Campbell SL, Khosravi-Far R, Rossman KL, Clark GJ, Der CJ . Increasing complexity of Ras signaling. Oncogene 1998; 17: 1395–1413.
Marshall C . How do small GTPase signal transduction pathways regulate cell cycle entry? Curr Opin Cell Biol 1999; 11: 732–736.
Shields JM, Pruitt K, McFall A, Shaub A, Der CJ . Understanding Ras: ‘it ain’t over ‘til it’s over’. Trends Cell Biol 2000; 10: 147–154.
Hanahan D, Weinberg RA . The hallmarks of cancer. Cell 2000; 100: 57–70.
Downward J . Targeting RAS signalling pathways in cancer therapy. Nat Rev Cancer 2003; 3: 11–22.
Schubbert S, Shannon K, Bollag G . Hyperactive Ras in developmental disorders and cancer. Nat Rev Cancer 2007; 7: 295–308.
Bos JL . Ras oncogenes in human cancer: a review. Cancer Res 1989; 49: 4682–4689.
Downward J . Cancer biology: signatures guide drug choice. Nature 2006; 439: 274–275.
Roberts PJ, Der CJ . Targeting the Raf-MEK-ERK mitogen-activated protein kinase cascade for the treatment of cancer. Oncogene 2007; 26: 3291–3310.
Sebolt-Leopold JS, Herrera R . Targeting the mitogenactivated protein kinase cascade to treat cancer. Nat Rev Cancer 2004; 4: 937–947.
Marzo I, Brenner C, Zamzami N, Susin SA, Beutner G, Brdiczka D et al. The permeability transition pore complex: a tar for apoptosis regulation by caspase and bcl-2-related proteins. J Exp Med 1998; 187: 1261–1271.
Marzo I, Brenner C, Zamzami N, Jurgensmeier JM, Susin SA, Vieira HL et al. Bax and adenine nucleotide translocator cooperate in the mitochondrial control of apoptosis. Science 1998; 281: 2027–2031.
Kokoszka JE, Waymire KG, Levy S, Sligh JE, Cai J, Jones DP et al. The ADP/ATP translocator is not essential for the mitochondrial permeability transition pore. Nature 2004; 427: 461–465.
Faure VH, Heddi A, Giraud S, Chautard D, Stepien G . Expression of oxidative phosphorylation genens in renal tumors and tumoral cell lines. Mol Carcinog 1996; 16: 165–172.
Chevrollier A, Loiseau D, Chabi B, Renier G, Donay O, Malthiery Y et al. ANT2 isoform required for cancer cell glycolysis. J Bioenerg Biomembr 2005; 37: 307–316.
Chevrollier A, Loiseau D, Gautier F, Malthlery Y, Stepien G . ANT2 expression under hypoxic conditions produces opposite cell-cycle behavior in 143B and HepG2 cancer cells. Mol Carcinog 2005; 42: 1–8.
Chevrollier A, Loiseau D, Stepien G . What is the specific role of ANT2 in cancer cells? Med Sci (Paris) 2005; 21: 156–161.
Le Bras M, Borgne-Sanchez A, Touat Z, El Dein OS, Deniaud A, Maillier E et al. Chemosensitization by knockdown of adenine nucleotide translocase-2. Cancer Res 2006; 66: 9143–9152.
Jang JY, Kim MK, Jeon YK, Joung YK, Park KD, Kim CW . Adenovirus adenine nucleotide translocator-2 shRNA effectively induces apoptosis and enhances chemosensitivity by the down-regulation of ABCG2 in breast cancer stem-like cells. Exp Mol Med 2012; 44: 251–259.
Jang JY, Choi Y, Jeon YK, Kim CW . Suppression of adenine nucleotide translocase-2 by vector-based siRNA in human breast cancer cells induces apoptosis and inhibits tumor growth in vitro and in vivo. Breast Cancer Res 2008; 10: 1–14.
Murakami Y, Yasuda T, Saigo K, Urashima T, Toyoda H, Okanoue T et al. Comprehensive analysis of microRNA expression patterns in hepatocellular carcinoma and non-tumorous tissues. Oncogene 2006; 25: 2537–2545.
Huang YS, Dai Y, Yu XF, Bao SY, Yin YB, Tang M et al. Microarray analysis of microRNA expression in hepatocellular carcinoma and non-tumorous tissues without viral hepatitis. J Gastroenterol Hepatol 2008; 23: 87–94.
Meng F, Henson R, Wehbe-Janek H, Ghoshal K, Jacob ST, Patel T . MicroRNA-21 regulates expression of the PTEN tumor suppressor gene in human hepatocellular cancer. Gastroenterology 2007; 133: 647–658.
Budhu A, Jia HL, Forgues M, Liu CG, Goldstein D, Lam A et al. Identification of metastasis-related microRNAs in hepatocellular carcinoma. Hepatology 2008; 47: 897–907.
Michael AT, Ashley LB, Nancy LK, Steven TR, Preethi HG . Role for MicroRNAs in regulating glucocorticoid response and resistance in multiple myeloma. Horm Cancer 2011; 2: 182–189.
Moodie SA, Willumsen BM, Weber MJ, Wolfman A . Complexes of Ras.GTP with Raf-1 and mitogen-activated protein kinase kinase. Science 1993; 260: 1658–1661.
Warne PH, Viciana PR, Downward J . Direct interaction of Ras and the amino-terminal region of Raf-1 in vitro. Nature 1993; 364: 352–355.
Vojtek AB, Hollenberg SM, Cooper JA . Mammalian Ras interacts directly with the serine/threonine kinase Raf. Cell 1993; 74: 205–214.
Zhang ZF, Settleman J, Kyriakis JM, Takeuchi-Suzuki E, Elledge SJ, Marshall MS et al. Normal and oncogenic p21ras proteins bind to the amino-terminal regulatory domain of c-Raf-1. Nature 1993; 364: 308–313.
Roux PP, Blenis J . ERK and p38 MAPK activated protein kinases: a family of protein kinases with diverse biological functions. Microbiol Mol Biol Rev 2004; 68: 320–344.
Vivanco I, Sawyers CL . The phosphatidylinositol 3-kinase AKT pathway in human cancer. Nat Rev Cancer 2002; 2: 489–501.
Castellano E, Downward J . Role of RAS in the regulation of PI 3-kinase. Curr Top Microbiol Immunol 2011; 346: 143–169.
Cantley LC . The phosphoinositide 3-kinase pathway. Science 2002; 296: 1655–1657.
Volinia S, Hiles I, Ormondroyd E, Nizetic D, Antonacci R, Rocchi M et al. Molecular cloning, cDNA sequence, and chromosomal localization of the human phosphatidylinositol 3-kinase p110 alpha (PIK3CA) gene. Genomics 1994; 24: 472–477.
Fruman DA, Meyers RE, Cantley LC . Phosphoinositide kinases. Annu Rev Biochem 1998; 67: 481–507.
Courtney KD, Corcoran RB, Engelman JA . The PI3K pathway as drug target in human cancer. J Clin Oncol 2010; 28: 1075–1083.
Cox AD, Der CJ . The dark side of Ras: regulation of apoptosis. Oncogene 2003; 22: 8999–9006.
Downward J . Ras signalling and apoptosis. Curr Opin Genet Dev 1998; 8: 49–54.
Downward J . PI 3-kinase, Akt and cell survival. Semin Cell Dev Biol 2004; 15: 177–182.
Acknowledgements
This work was supported by the Global Core Research Center (GCRC) Grant No. 2012-0001190 from the National Research Foundation (NRF), Ministry of Education, Science and Technology (MEST), Republic of Korea, also by Grant No SS100003 from the Seoul R&BD Program.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Experimental & Molecular Medicine website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Jang, JY., Lee, YS., Jeon, YK. et al. ANT2 suppression by shRNA restores miR-636 expression, thereby downregulating Ras and inhibiting tumorigenesis of hepatocellular carcinoma. Exp Mol Med 45, e3 (2013). https://doi.org/10.1038/emm.2013.1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/emm.2013.1
Keywords
This article is cited by
-
LncRNA MONC suppresses the malignant phenotype of Endometrial Cancer Stem Cells and Endometrial Carcinoma Cells by regulating the MiR-636/GLCE axis
Cancer Cell International (2021)
-
LncRNA-HNF1A-AS1 functions as a competing endogenous RNA to activate PI3K/AKT signalling pathway by sponging miR-30b-3p in gastric cancer
British Journal of Cancer (2020)
-
The pleiotropic regulation of cyclin D1 by newly identified sesaminol-binding protein ANT2
Oncogenesis (2017)
-
ANT2 shRNA downregulates miR-19a and miR-96 through the PI3K/Akt pathway and suppresses tumor growth in hepatocellular carcinoma cells
Experimental & Molecular Medicine (2016)
-
microRNA: A Promising Diagnostic Biomarker and Therapeutic Target for Hepatocellular Carcinoma
Digestive Diseases and Sciences (2014)