Abstract
The classic epileptic encephalopathies, including infantile spasms (IS) and Lennox–Gastaut syndrome (LGS), are severe seizure disorders that usually arise sporadically. De novo variants in genes mainly encoding ion channel and synaptic proteins have been found to account for over 15% of patients with IS or LGS. The contribution of autosomal recessive genetic variation, however, is less well understood. We implemented a rare variant transmission disequilibrium test (TDT) to search for autosomal recessive epileptic encephalopathy genes in a cohort of 320 outbred patient–parent trios that were generally prescreened for rare metabolic disorders. In the current sample, our rare variant transmission disequilibrium test did not identify individual genes with significantly distorted transmission over expectation after correcting for the multiple tests. While the rare variant transmission disequilibrium test did not find evidence of a role for individual autosomal recessive genes, our current sample is insufficiently powered to assess the overall role of autosomal recessive genotypes in an outbred epileptic encephalopathy population.
Similar content being viewed by others
Introduction
Epileptic encephalopathies are severe and therapy-resistant epilepsies of childhood, which frequently lead to developmental delay and multiple associated medical issues. Infantile spasms (IS) and Lennox–Gastaut syndrome (LGS) represent two of the more common broad subtypes of epileptic encephalopathies. Many novel genes for epileptic encephalopathies have been discovered in the last 5 years, fueled by the access to whole-exome sequencing. In particular, exome sequencing has highlighted the important role of de novo variants with current estimates suggesting that over 15% of non-Dravet epileptic encephalopathy cases are explained by a disease-causing de novo variant in an established epileptic encephalopathy gene1, 2 with this estimate increased to over 80% among individuals diagnosed with Dravet syndrome.3 Up to a further 3% have been reported to be explained by likely clinically relevant de novo copy-number variants.4
While the role of de novo genetic variation in epileptic encephalopathies is increasingly understood, the role of recessive genetic variation, outside of recessive neurometabolic disorders such as lysosomal disorders, amino acid or organic acid imbalances, congenital disorders of glycosylation, and some mitochondrial diseases, remains unclear. In our current study, we systematically assessed autosomal recessive inheritance in 320 IS or LGS patient–parent trios who did not have a likely disease-causing de novo variant among one of the established dominant epileptic encephalopathy genes.1, 2 In general, the 320 cases studied here had already been intensively studied for neurometabolic disorders using biochemical assessments.
Subjects and methods
Cohort
Three-hundred and twenty epileptic encephalopathy trios were recruited through multiple international consortia, including 57 IS or LGS trios unpublished in our earlier studies.1, 2 Patients did not have a clearly identified metabolic or genetic cause for their epilepsy based on clinically available testing, which varied across institutions. This collection of 320 trios did not include: (a) patients previously found to have a disease-causing de novo variant in an established dominant epileptic encephalopathy gene, and (b) trios where exome sequencing was based on a lymphoblastoid cell line source for at least one of the three family members. The overall cohort was not enriched for consanguineous parents. Only two parent pairs showed an identity-by-descent >0.125, both <0.15, which is approximately equivalent to third degree relatives.5
Among the 320 trios; two families reported multiple affected children. For one of these families both the proband and affected sibling were investigated through exome sequencing, while for the second family only the proband and parents were studied. Sequencing methods used to generate the sequence data have been previously described.1, 2
Transmission disequilibrium tests
For the transmission test, we used two approaches that we have previously introduced.6, 7 First, we tested for an autosomal homozygous or compound heterozygous effect using core TDT.7 In computing the test, we selected loss-of-function and missense single-nucleotide substitution variants found at a global population minor allele frequency <5% (MAF<0.05). The loss-of-function variants were defined as stop gain, stop lost, start lost, and canonical splice acceptor and donor site variants. For the missense variants, we used our in-house Analysis Tool for Annotated Variants platform to identify the possibly and probably damaging variants based on a maximum Polyphen-2 HumDiv and HumVar prediction score8 of >0.4333. This test was applied to each autosomal gene individually as well as collectively across a set of 99 autosomal recessive neurometabolic genes published by van Karnebeek et al.9 The recessive neurometabolic gene-set analysis allowed us to assess whether there was evidence for elevated rate of recessive genotypes among recessive neurometabolic disorder genes beyond what had already been screened out by the conventional biochemical assessments performed on this patient sample.
Second, we tested for a general effect of inherited autosomal variation by using a rare variant TDT that uses information from an independent collection of population controls (6503 EVS10 plus 1303 IGM sequenced controls) to weigh the contribution of variants to the final test statistic.6 In this analysis, qualifying variants were defined using the same PolyPhen-2 thresholds as above and were again required to have a global MAF <5%. Given that population stratification can impact the power of the test but not the type I error, we restricted this second analysis to trios with European ancestry (n=286 trios).
Power simulation
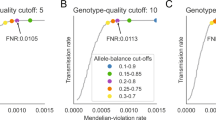
As we had not previously performed power simulations for the type of gene-set application conducted in our neurometabolic analysis, we conduct a new power simulation to evaluate the types of effects that we could exclude based on this analysis (Figure 1). In these simulations, we conditioned on the parental genotype information contained in this IS/LGS population sample and characterized the distribution of offspring genotypes, given this information and the fact that the offspring is affected. This distribution is a function of the number of causal genes, for which the family is informative, which is related to the density of causal genes within the actual gene set, and the relative risk of the offspring developing disease, given that they have two affected gene copies. We give the details of this procedure here.
coreTDT power simulation conditional on the parental genotype of 54 informative families and 20 informative genes in the compound heterozygous analysis. Presents the combination of the relative risk and the proportion of disease causal genes among these 20 informative genes as a single unit, under which the tests can achieve 80% power.
Let Gf, Gm, Go be the number of gene copies harboring a qualifying variant in the trio’s father, mother, and offspring, respectively. We condition our power analysis on the observed parental genotype and study our ability to identify signal given a differing proportion of causal genes (out of the total number of genes considered), γ and differing relative risks, R, of being diseased, given two gene copies (of a causal disease gene) are affected versus less than two copies are affected. Since the analysis is conditional on the observed parental data, only a subset of genes and families are informative.7 Specifically, only 20 genes across 54 families can have compound genotypes that lead to informative transmissions, that is, Gf=Gm=1, Gf=1, Gm=2 or Gf=2, Gm=1. A total of 46 families are informative for only one gene and eight families are informative for two genes. In each of these eight families, the two genes are located on different chromosomes, so we assume that the transmissions of each gene are independent.
Let Do=1 indicate the fact that the offspring is affected. Let C be an indicator of whether the gene whose transmission is being considered is among the set of disease causal genes or not. When a family is informative for two genes disease causal indicators are given for each gene by C1 and C2. Note, we assume the disease risk for samples with multiple affected disease genes are the same with those with only one affected disease gene.
To simulate trios under the alternative, we first randomly select 20γ genes as disease causal and then generate offspring as follows.
If the family is informative for only one gene, the distribution of both offspring’s gene copies being affected is given by:

If the family is informative for two genes and no more than one of them are disease causal, the compound genotype, the two genes, can be computed independently of one another using the equation above. When both genes are disease causal, their transmissions are not independent given the offspring is affected. In this case, the compound genotypes of the offspring, for the two genes, can be given by,

where Go1, Gm1, Gf1 and Go2, Gm2, Gf2 denotes the trio’s compound genotypes at the first and second gene, respectively. We apply coreTDT to each simulated data set, and for each combination of γ and R, we use 1000 replicates to estimate the power. The combination of γ and R that obtains 80% power are presented in Figure 1.
Accession numbers
The exome-sequencing data reported in this paper are deposited in the Database of Genotypes and Phenotypes (dbGaP) with the accession number phs000654.v2.p1. The EuroEPINOMICS-RES data are deposited in the European Genome-phenome Archive with the accession numbers: EGAS00001000190, EGAS00001000386, and EGAS00001000048.
Results
We assessed the role of inherited rare variation using the population control-weighted rare variant TDT.6 This test was applied to each autosomal gene across 320 eligible trios. No gene reached exome-wide significance after correcting for the 17 816 consensus coding sequence (CCDS release 14) autosomal genes (adjusted α=2.81−10−6, Table 1). Though population stratification cannot affect the false positive rate of the test, it can affect the power.6 We also conducted an analysis that was restricted to the 286 trios of European ancestry. Again, no gene reached the exome-wide significance level (Table 1).
We then tested for the presence of a recessive effect in each autosomal gene across the 320 trios. After quality control, only 3472 autosomal genes were found to have at least one informative family, that is, contain qualifying variants within the gene and that could, potentially, lead to homozygous or compound heterozygous offspring. None of these 3472 genes achieved significance after correcting for the number of genes tested (adjusted α=1.44 × 10−5). The 10 most significant genes are listed in Table 2. To investigate whether there is any evidence of recessive neurometabolic involvement in this sample, we also applied the coreTDT to the set of 99 autosomal recessive neurometabolic genes,9 looking for an enrichment of homozygous or compound heterozygous offspring across the entire gene set as a single unit. No enrichment was found (P=0.51).
We investigated the power of this analysis. Since only 54 families are informative for at least one of the 99 autosomal recessive neurometabolic genes, and only 20 genes have at least one informative family, our analyses are effectively restricted to these 54 families and 20 genes. We vary the proportion of informative genes that are actually disease causal and the relative risk and identify combinations of these parameters that attain at least 80% power (see ‘Power simulation’ for details). The results of this analysis can be found in Figure 1. As can be seen, even when the compound heterozygous or homozygous qualifying variants are fully penetrant, the causal gene proportion must be >40% to attain 80% power. When the proportion of causal genes is larger, for example, 80%, we will have high power to detect an effect even with a relatively low relative risk.
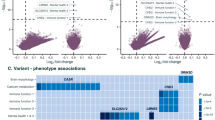
Using established standards to identify clinically relevant recessive genotypes,11, 12 one trio was found to have inherited two clinically relevant SPATA5 variants in a compound heterozygous manner.13 The proband’s phenotype is consistent with the SPATA5 disease literature, and both variants (NM_145207.2; c.1677C>A (p.(Tyr559*)) and c.251G>A (p.(Arg84Gln)) have previously been described as clinically relevant among other patients with SPATA5 encephalopathy.13
Discussion
A number of rare recessive disorders can present with an epileptic encephalopathy, particularly neurometabolic disorders; the latter are generally identified by biochemical analyses of blood, urine or CSF. We performed a global, hypothesis-free test to assess the role of autosomal recessive genetic variation in 320 patients with classic epileptic encephalopathies undiagnosed with standard clinical workups. Our sample of patient–parent trios did not identify a genome-wide significant departure in the observed number of offspring with recessive genotypes from that expected for any specific gene, or among 99 genes compiled for autosomal recessive neurometabolic disorders.
Many classical recessive metabolic disorders are routinely identified through biochemical screening prior to research study enrollment. Within our sample of 320 trios, we did not find any genetic neurometabolic disorders that were missed through the conventional biochemical screening. From a clinical perspective, this emphasizes that conventional biochemical screening for these treatable causes should continue to be pursued. We did identify a single case among the 320 with a clinically relevant recessive genotype in SPATA5,13 a recently described gene for a recessive condition characterized by seizures, microcephaly, intellectual disability, and hearing loss.
The role of various dominant epilepsy genes including ALG13, CDKL5, DNM1, GABRB3, SCN1A, SCN2A, and STXBP1, for epileptic encephalopathies was securely established through exome sequencing of 356 trios and subsequent genome-wide assessments for excess de novo variants identified in individual genes.1, 2 No single gene passes a comparable threshold among the 320 trios studied here when assessing autosomal recessive genotypes. We demonstrate that the current sample of 320 trios is insufficiently powered to appropriately estimate what overall contribution autosomal recessive epilepsy genes have on the epileptic encephalopathies; however, our power analyses show that we do have sufficient power to rule out a large role from known recessive neurometabolic genes among this patient sample that has been previously screened for such factors using conventional biochemical assessments. Using a similar approach, a recent study on 4125 patient–parent trios with various developmental disorders identified two novel autosomal recessive disease genes exceeding genome-wide significance,14 emphasizing the importance of acquiring larger numbers to more confidently interpret the current lack of signal for very rare genetic epilepsies with recessive inheritance. Large-scale collaborative initiatives like the epilepsy genetic initiative and the Epi25 effort will aid the efforts to analyze genomic data on this scale.
References
Epi4K Consortium. Epilepsy Phenome/Genome Project Epi4K Consortium. Epilepsy Phenome/Genome Project, Allen AS et al: De novo mutations in epileptic encephalopathies. Nature 2013; 501: 217–221.
EuroEPINOMICS-RES Consortium. Epilepsy Phenome/Genome Project. Epi4K Consortium: De novo mutations in synaptic transmission genes including DNM1 cause epileptic encephalopathies. Am J Hum Genet 2014; 95: 360–370.
Harkin LA, McMahon JM, Iona X et al: The spectrum of SCN1A-related infantile epileptic encephalopathies. Brain 2007; 130 (Pt 3): 843–852.
Epilepsy Phenome/Genome Project Epi4K Consortium: Copy number variant analysis from exome data in 349 patients with epileptic encephalopathy. Ann Neurol 2015; 78: 323–328.
Purcell S, Neale B, Todd-Brown K et al: PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 2007; 81: 559–575.
Jiang Y, Satten GA, Han Y et al: Utilizing population controls in rare-variant case-parent association tests. Am J Hum Genet 2014; 94: 845–853.
Jiang Y, McCarthy JM, Allen AS : Testing the effect of rare compound-heterozygous and recessive mutations in case—parent sequencing studies. Genet Epidemiol 2015; 39: 166–172.
Adzhubei IA, Schmidt S, Peshkin L et al: A method and server for predicting damaging missense mutations. Nat Methods 2010; 7: 248–249.
van Karnebeek CD, Shevell M, Zschocke J, Moeschler JB, Stockler S : The metabolic evaluation of the child with an intellectual developmental disorder: diagnostic algorithm for identification of treatable causes and new digital resource. Mol Genet Metab 2014; 111: 428–438.
Exome Variant Server (EVS), NHLBI GO Exome Sequencing Project (ESP), Seattle, WA, USA.
Zhu X, Petrovski S, Xie P et al: Whole-exome sequencing in undiagnosed genetic diseases: interpreting 119 trios. Genet Med 2015; 17: 774–781.
Need AC, Shashi V, Hitomi Y et al: Clinical application of exome sequencing in undiagnosed genetic conditions. J Med Genet 2012; 49: 353–361.
Tanaka AJ, Cho MT, Millan F et al: Mutations in SPATA5 are associated with microcephaly, intellectual disability, seizures, and hearing loss. Am J Hum Genet 2015; 97: 457–464.
Akawi N, McRae J, Ansari M et al: Discovery of four recessive developmental disorders using probabilistic genotype and phenotype matching among 4,125 families. Nat Genet 2015; 47: 1363–1369.
Acknowledgements
We are deeply grateful to the probands, their families, clinical research coordinators, and referring physicians for their participation and provision of phenotype data, and DNA samples used in this study. We thank the EPGP Administrative (C Freyer, K Fox, R Fahlstrom, S Cristofaro, and K McGovern), Bioinformatics Core (G Nesbitt, K McKenna, and V Mays), staff at Coriell Institute – NINDS Genetics Repository, and members of the Institute for Genomic Medicine, Columbia University (P Cansler, J Charoensri, B Copeland, S Kamalakaran, J Keebler, B Krueger, C Malone, C Mebane, and M Cook) for their dedication and commitment to this work. We also thank R Stewart, K Gwinn, R Corriveau, B Fureman, and V Whittemore from the National Institute of Neurological Disorders and Stroke for their careful oversight and guidance of both EPGP and Epi4K. We thank the following organizations for assistance in publicizing EPGP; enabling us to recruit participants effectively: AED Pregnancy Registry, American Epilepsy Society, Association of Child Neurology Nurses, California School Nurses Organization, Child Neurology Society, Citizens United for Research in Epilepsy, Dravet Syndrome Foundation, Epilepsy Alliance of Orange County, Epilepsy Foundation, Epilepsy Therapy Project, Finding a Cure for Epilepsy and Seizures, IDEA League, InfantileSpasms.com, Lennox–Gastaut Syndrome Foundation, PatientsLikeMe, People Against Childhood Epilepsy, PVNH Support & Awareness, and Seizures & Epilepsy Education. We would like to acknowledge the following individuals or groups for the contributions of control samples: D Daskalakis; P Lugar; J Milner; T Young and K Whisenhunt; Z Farfel, D Lancet, and E Pras; W Lowe; R Gbadegesin and M Winn; K Schmader, S McDonald, HK White, and M Yanamadala; A Holden; E Behr; C Moylan; AM Diehl and M Abdelmalek; S Palmer; G Nestadt; J Samuels; Y Wang; M Carrington; M Harms; T Miller; A Pestronk; R Bedlack; R Brown; N Shneider; S Gibson; J Ravits; A Gilter; J Glass; F Baas; E Simpson; and G Rouleau; K Welsh-Bomer, C Hulette, J Burke; The ALS Sequencing Consortium; The Murdock Study Community Registry and Biorepository; M Connors, L Morris, and the CHAVI investigators; the Carol Woods and Crosdaile Retirement Communities; and DUHS (Duke University Health System) Nonalcoholic Fatty Liver Disease Research Database and Specimen Repository. The collection of control samples and data was funded in part by: Biogen Idec.; The Duke Chancellor’s Discovery Program Research Fund 2014; Bill and Melinda Gates Foundation; The Division of Intramural Research; B57 SAIC-Fredrick Inc M11-074; Bryan ADRC NIA P30 AG028377; The Ellison Medical Foundation New Scholar award AG-NS-0441-08; National Institute of Mental Health (K01MH098126, R01MH097993); National Institute of Allergy and Infectious Diseases (1R56AI098588-01A1); and National Institute of Allergy and Infectious Diseases Center (U19-AI067854, UM1-AI100645). We thank the NHLBI GO Exome Sequencing Project and its ongoing studies that produced and provided exome variant calls for comparison: the Lung GO Sequencing Project (HL-102923), the WHI Sequencing Project (HL-102924), the Broad GO Sequencing Project (HL-102925), the Seattle GO Sequencing Project (HL-102926), and the Heart GO Sequencing Project (HL-103010). This work was supported by grants from the National Institute of Neurological Disorders and Stroke (The Epilepsy Phenome/Genome Project NS053998; Epi4K NS077364, NS077274, NS077303, and NS077276), The Andrew’s Foundation, Finding a Cure for Epilepsy and Seizures, the Richard Thalheimer Philanthropic Fund, and the Eurocores program EuroEPINOMICS-RES of the European Science Foundation. The project received further support through grants from the Fund for Scientific Research Flanders (FWO); the Academy of Finland (141549); the Folkhälsan Research Foundation; the program ‘Investissements d’avenir’ ANR-10-IAIHU-06; the Federal Ministry for Education and Research (IonNeurONet: 01GM1105), the German Research Foundation (DFG: HE5415/3-1; Le1030/11-1; RO3396/2-1), the German Society for Epileptology (DGfE), the Foundation noepilep.; the Swiss National Science Foundation (SNF: 32EP30_136042/1); the Wellcome Trust (09805); intramural funds of the University of Kiel; the Popgen 2.0 network (P2N) through the German Ministry for Education and Research (01EY1103); and the European Union through Seventh Framework Programme (FP7) under the project DESIRE (N602531). The project also received infrastructural support through the Institute of Clinical Molecular Biology in Kiel, supported in part by DFG Cluster of Excellence ‘Inflammation at Interfaces’ and ‘Future Ocean’. Orrin Devinsky, David B Goldstein, Steve Petrou and Slavé Petrovski have interests in companies related to epilepsy precision medicine.
Author contributions
Sequence analysis and statistical interpretation: YJ (lead analyst), ASA, and DBG. Bioinformatics processing: JBr, ELH, YJ, SlP, ZR, and QW. Clinical expert panel: IH, HCM, AP, and SW. Writing of manuscript: ASA, SFB, PDJ, DBG, ELH, IH, YJ, DHL, and SlP. Epi4K Steering Committee: ASA, SFB, PC, ND, DD, EEE, MPE, TG, DBG, ELH, MRJ, RuKu, DHL, AGM, HCM, TJO, RO, StP, SlP, AP, IES, and ES. EuroEPINOMICS-RES consortium leadership and study coordination: IH, PDJ, and SW. EuroEPINOMICS study design: A-EL, AS, BK, HL, IH, PG, PDJ, and SW. EPGP study design: BKA, OD, DD, MPE, RuKu, DHL, RO, ES, and MRW. Epi4K epileptic encephalopathy phenotyping: SFB, PC, DD, RuKu, DHL, HCM, RO, AP, IES, and ES. EuroEPINOMICS-RES proband recruitment and phenotyping: A-EL, AR, CM, CD, DC, DP, DH, EL, FZ, FRos, HC, HH, HM, HL, IH, JJ, JL, JS, KSe, KMK, KSt, NB, PS, RG, RSM, SvS, SW, SB, TL, TT, US, VK, and YW. EPGP proband recruitment and phenotyping: BA, EA, FA, DA, JFB, SFB, GDC, DCo, PCr, OD, DD, MEF, NBF, DF, EBG, TG, SG, SRH, JH, KH, SLH, HEK, RCKn, EK, RaKu, RuKu, DHL, SMM, PVM, EJN, JMP, JP, KP, AP, IES, JJS, RAS, JSi, LS, MS, LLT, AV, EPGV, GKV, JW, and PW. EPGP phenotype data analysis: BA, BKA, AB, JB, GDC, OD, DD, MPE, JF, TG, SJ, AK, RCKn, RuKu, DHL, RO, JMP, AP, IES, RAS, RS, ES, JJS, JSu, PW, and MRW.
Author information
Authors and Affiliations
Consortia
Ethics declarations
Competing interests
The sponsors of the study had no role in study design, data collection, data analysis, data interpretation, or writing of the report.
Additional information
Andrew S Allen, Samuel F Berkovic, Joshua Bridgers, Patrick Cossette, Dennis Dlugos, Michael P Epstein, Tracy Glauser, David B Goldstein, Erin L Heinzen, Yu Jiang, Michael R Johnson, Ruben Kuzniecky, Daniel H Lowenstein, Anthony G Marson, Heather C Mefford, Terence J O’Brien, Ruth Ottman, Steven Petrou, Slavé Petrovski, Annapurna Poduri, Zhong Ren, Ingrid E Scheffer, Elliott Sherr, Quanli Wang
Rudi Balling, Nina Barisic, Stéphanie Baulac, Hande Caglayan, Dana Craiu, Peter De Jonghe, Christel Depienne, Renzo Guerrini, Ingo Helbig, Helle Hjalgrim, Dorota Hoffman-Zacharska, Johanna Jähn, Karl Martin Klein, Bobby Koeleman, Vladimir Komarek, Roland Krause, Eric Leguern, Anna-Elina Lehesjoki, Johannes R Lemke, Holger Lerche, Tarja Linnankivi, Carla Marini, Patrick May, Rikke S Møller, Hiltrud Muhle, Deb Pal, Aarno Palotie, Felix Rosenow, Kaja Selmer, Jose M Serratosa, Sanjay Sisodiya, Ulrich Stephani, Katalin Sterbova, Pasquale Striano, Arvid Suls, Tiina Talvik, Sarah von Spiczak, Yvonne Weber, Sarah Weckhuysen, Federico Zara
Bassel Abou-Khalil, Brian K Alldredge, Dina Amrom, Eva Andermann, Frederick Andermann, Jocelyn F Bautista, Samuel F Berkovic, Judith Bluvstein, Gregory D Cascino, Damian Consalvo, Patricia Crumrine, Orrin Devinsky, Dennis Dlugos, Michael P Epstein, Miguel E Fiol, Nathan B Fountain, Jacqueline French, Daniel Friedman, Tracy Glauser, Kevin Haas, Sheryl R Haut, Jean Hayward, Sucheta Joshi, Andres Kanner, Heidi E Kirsch, Eric H Kossoff, Rachel Kuperman, Ruben Kuzniecky, Daniel H Lowenstein, Shannon M McGuire, Paul V Motika, Edward J Novotny, Ruth Ottman, Juliann M Paolicchi, Jack Parent, Kristen Park, Annapurna Poduri, Ingrid E Scheffer, Renée A Shellhaas, Elliott Sherr, Joseph Sirven, Michael C Smith, Joseph Sullivan, Liu Lin Thio, Anu Venkat, Eileen PG Vining, Gretchen K Von Allmen, Judith L Weisenberg, Peter Widdess-Walsh, Melodie R Winawer
Rights and permissions
About this article
Cite this article
Epi4K Consortium., EuroEPINOMICS-RES Consortium. & Epilepsy Phenome Genome Project. Application of rare variant transmission disequilibrium tests to epileptic encephalopathy trio sequence data. Eur J Hum Genet 25, 894–899 (2017). https://doi.org/10.1038/ejhg.2017.61
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2017.61
This article is cited by
-
The role of recessive inheritance in early-onset epileptic encephalopathies: a combined whole-exome sequencing and copy number study
European Journal of Human Genetics (2019)