Abstract
Mutations in the gene encoding surfactant protein C (SFTPC) have led to a broad range of phenotypes from neonatal respiratory distress syndrome to adult interstitial lung disease. We previously identified the c.435G>C variant in the SFTPC gene associated with fatal neonatal respiratory distress syndrome in an infant girl. Although this variation is predicted to change glutamine (Q) at position 145 to histidine (H), its position at the last base of exon 4 and the severity of the phenotype suggested that it might also induce a splicing defect. To test this hypothesis, we used hybrid minigene, biochemical and immunofluorescence tools to decipher the molecular mechanism of the mutation. Immunoblotting and confocal imaging showed similar maturation and localization of wild-type and Q145H proteins, but hybrid minigene analysis showed complete exon 4 skipping. Since the exon 4 is in frame, a putative truncated protein of 160 amino acids would be produced. We have shown that this truncated protein had an altered intracellular trafficking and maturation. The c.435G>C mutation is deleterious not because of its amino acid substitution but because of its subsequent splicing defect and should be referred to as r.325_435del and p.Leu109_Gln145del. The absence of residual full-length transcripts fully explained the severity of the phenotype we observed in the infant.
Similar content being viewed by others
Introduction
The genetic surfactant disorders are a group of rare diseases responsible for interstitial lung disease (ILD) in both children and adults. Since the first description of ILD being associated with an altered SFTPC gene in 2001, about 50 different mutations have been reported, almost all with heterozygous status.1 Mutations in the SFTPC gene are defined by their location: the mutations within the BRICHOS domain, which alter the ability of proSP-C to fold correctly, causing aggresome formation and cellular dysfunction,2, 3, 4 and the non-BRICHOS mutations, which induce abnormal trafficking of proSP-C, for functional impairment of lipid uptake.5, 6 The phenotypes associated with the SFTPC mutations vary, from neonatal respiratory distress syndrome to adult ILD, and this wide phenotypic variability is not fully explained by the genotype. Severe phenotypes have been associated with two splicing mutations within the donor splice site at the exon–intron 4 junction of SFTPC gene, c.435+1 G>A and c.435G>A p.Gln145=.1, 7 Both mutations result in SFTPC exon 4 skipping.
The aim of this study was to decipher the molecular mechanism leading to the c.435G>C deleterious effect. Although this variation is a missense mutation, its position at the last base of exon 4 suggested that it might induce a splicing defect and lead to the skipping of exon 4. A minigene strategy was used to test this hypothesis, together with immunoblotting and confocal imaging to compare the cellular impact of the amino acid change at position 145 and the deleted protein, resulting from the splicing mutation, on the maturation and trafficking of proSP-C. The results obtained with the splicing mutant agree with the phenotype we found associated with this mutation: an extremely severe neonatal respiratory distress syndrome that led to death within the third month of life.8
Methods
Recommendations of the Human Genome Variation Society (http://www.hgvs.org/mutnomen/) were followed for mutation names, with +1 used as the A for the ATG translation initiation start site in the GenBank SFTPC reference sequence NM_003018.3. Exons are numbered like in NG_016968.1. The impact of c.435G>C, c.435G>A and c.435+1 G>A mutations on the strength of the splice donor site was analyzed by using free online software as previously described.9 In addition, the Shapiro and Senapathy nucleotide weight matrices were used to calculate each splice site score.10 The data of the study (variant, phenotype and possible functional consequence) have been submitted into the freely accessible public database LOVD (http://databases.lovd.nl/shared/individuals/00090135).
We constructed a hybrid minigene containing the SFTPC exon 4 and its flanking intronic regions (231 bp upstream and 334 bp downstream) PCR-amplified from genomic DNA and subcloned into the pET01 vector (Mobitec, Göttingen, Germany) as previously described.11 Human proSP-C wild-type (WT) cDNA (Origene, Herford, Germany) was subcloned into the pcDNA3.1/V5-His-TOPO vector. All mutations were generated by using a site-directed mutagenesis kit (Stratagene, Waghaeusel-Wiesental, Germany), and pET01 or pcDNA3.1/V5-His-TOPO constructs were transiently transfected in A549 cells by using Lipofectamine 2000 (Life Technologies, Courtaboeuf, France) according to the manufacturer’s instructions.
After 24 h of transfection, total RNA was extracted with use of TRIzol Reagent (Life Technologies) according to the manufacturer’s instructions. RT-PCR involved use of the Maxima First Strand cDNA Synthesis kit for RT-qPCR with dsDNase (Thermo Scientific, St Leon-rot, Germany). Amplified products were separated by capillary electrophoresis as described.11
Ratios of splicing isoforms were determined as the peak area of the considered SFTPC isoform divided by the sum of peak areas of all isoforms for each variant.
Human WT proSP-C or mutated cDNA was transiently transfected in A549 cells for 24 h, then total cell protein extracts were collected after cell lysis in RIPA buffer containing protease inhibitors. In total, 30 μg protein was immunoblotted with the following primary antibodies rabbit polyclonal anti-proSP-C or mouse monoclonal anti-beta actin (both Abcam, Cambridge, UK), then the secondary antibodies horseradish peroxidase-conjugated goat anti-rabbit or goat anti-mouse (both Cell Signaling, Danvers, MA, USA).
For immunofluorescence, A549 cells seeded on coverslips were transfected for 24 h, then fixed with 4% paraformaldehyde. Immunostaining involved use of the primary antibodies rabbit polyclonal anti-proSP-C and mouse monoclonal anti-calnexin (both Abcam), followed by the secondary antibodies goat anti-rabbit Alexa Fluor 488 and goat anti-mouse Alexa Fluor 568 (both Life Technologies). Nuclei were detected with To-Pro-3 Iodide staining (Life Technologies) according to the manufacturer’s instructions. Laser scanning confocal microscopy involved use of a Zeiss confocal LSM 510 and Zeiss Zen 2009 software.
Results
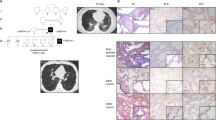
The index patient was an infant girl delivered by caesarian section at 38 weeks after an uneventful pregnancy. Within the first month of life, she experienced rapidly progressive respiratory failure that required invasive ventilation related to severe respiratory distress syndrome. High-resolution computed tomography scan revealed bilateral ground-glass opacity with anterior pneumothorax. Despite methylprednisolone pulses, exogenous surfactant administration and high-pressure ventilation, the respiratory condition worsened and the patient died at age 3 months.
The c.435G>C mutation was identified in one allele of the SFTPC gene in the index patient.8 Because this mutation is located at the last nucleotide of exon 4, we used splicing bioinformatics tools to investigate a putative change in the strength of this donor splice site at the exon–intron 4 junction (Table 1). Globally, the c.435G>C mutation induced a marked decrease or complete abolition of the predicted scores of this donor splice site (11 of 11), in the same manner as the two positive controls, which indicates that this mutation could alter the recognition of the exon 4 by the splicing machinery.
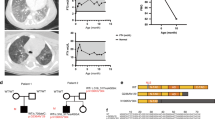
We then assessed in vitro splicing by using hybrid minigene assay. Capillary electrophoresis analysis revealed a single 245- bp transcript after empty-plasmid transfection (Figure 1a). With the WT construct, two peaks could be detected (Figure 1b), including a major peak of 355 bp corresponding to the full-length exon 4 (75%) and a minor peak of 245 bp corresponding to exon 4 skipping (25%). By contrast, use of the c.435+1G>A construct revealed a single 245-bp transcript (100%) (Figure 1c). A similar result was observed with the c.435G>C construct (Figure 1d), which demonstrates the effect of this exonic mutation on SFTPC splicing.
Hybrid minigene splicing assays. Left: examples of capillary electrophoresis analysis of RT-PCR products obtained after 25 cycles. RNA was purified from A549 cells transfected with empty plasmid (pET01) (a), minigene containing exon 4 WT (b) or mutant exon 4: c.435+1G>A as a positive control (c) and c.435G>C (d). RT-PCR with use of a FAM-labeled forward primer located within the splice donor exon and a reverse primer within the splice acceptor exon of the pET01 plasmid. The corresponding size and relative amount of each peak is indicated. The peak at 245 bp corresponds to exon 4 skipped mRNA and 355 bp to full-length mRNA. Right: schematic representation of the minigenes used in the splicing assay. The minigenes contain wild-type (WT) or mutated SFTPC exon 4 flanked by their natural intronic sequences (231 bp of the upstream intron 3 and 334 bp of the downstream intron 4). Data are mean percentage of at least two independent measurements performed four times.
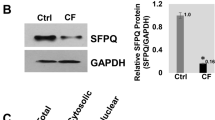
Minigenes may not precisely recapitulate the in vivo situation, and alternative splicing could possibly occur in vivo. Hence, some amount of normally processed transcript encoding the missense change may be produced in patient’s cells. Therefore, we investigated the effect of proSP-CQ145H and proSP-CΔExon4, the resulting protein of the splicing defect, at the protein level. Immunoblot analysis with anti-proSP-C antibody revealed two slight bands at 21 and 14 kDa with the empty-vector transfection and no transfection (Figure 2a). With proSP-CWT transfection, lysates contained a strong band of 21 kDa corresponding to a primary translation product, and higher and smaller processed intermediates at 25 and 14 kDa (Figure 2a). ProSP-CQ145H transfection resulted in a similar protein expression profile, which suggests normal processing, whereas proSP-CΔExon4 transfection resulted in a smaller primary translation product, ≈17 kDa, without post-translational processing.
ProSP-C processing and trafficking. (a) Representative immunoblotting of proSP-C in whole cell lysates of A549 cells untransfected (Ctr.) or transiently transfected with empty plasmid (pcDNA3.1V5HisTOPO), proSP-CWT, proSP-CQ145H and proSP-CΔEx4. (b) Representative confocal images of A549 cells after 24 h expression of empty plasmid, proSP-C WT, proSP-CQ145H and proSP-CΔEx4. Immunostaining was with rabbit polyclonal anti-proSP-C (green) and mouse monoclonal anti-calnexin antibodies (red). Nuclei were detected by ToPro-3 iodide (blue).
To assess cell trafficking modifications, we used immunostaining with anti-proSP-C antibody (Figure 2b) after transfection of the empty-vector plasmid (pcDNA3.1-V5-His) or WT (proSP-CWT) or mutated (proSP-CΔExon4 and proSP-CQ145H) plasmids into A549 cells. Confocal microscopy revealed no signal with empty-vector transfection, which suggests very weak proSP-C endogenous expression. With proSP-CWT, the proprotein predominantly localized in cytosolic vesicles and did not co-localize with endoplasmic reticulum. A similar intracellular distribution was observed with proSP-CQ145H transfection. By contrast, the proSP-CΔExon4 protein was retained in the endoplasmic reticulum (Figure 2b).
Discussion
We have shown that, in vitro, the c.435G>C mutation, initially interpreted as a missense mutation p.(Gln145His), results in the complete skipping of exon 4 r.325_435del. Since the exon 4 is in frame, a putative stable truncated protein of 160 amino acids p.Leu109_Gln145del would be produced. We observed a similar intracellular processing and trafficking of proSP-CQ145H and proSP-CWT, which suggests that if this protein was produced, this substitution would not be responsible for the severe lung phenotype observed in the infant girl.
As the c.435G>C induces complete exon 4 skipping, the effect is similar to the substitution at the first nucleotide of intron 4 (c.435+1G>A) described by Nogee et al.1 Recently, severe lung phenotype with onset of symptoms in early infancy was shown in a full-term infant carrying the c.435G>A mutation, again affecting the donor splice site at the exon–intron 4 junction.7 In this case, analysis of mRNA from frozen lung tissue from the patient confirmed exon 4 skipping. The severity of the phenotype is in accordance with our results. Those results highlight that the donor splice site at the exon–intron 4 junction is a hot-spot mutation in SFTPC gene leading to a severe phenotype. We and others show that unmasking the splicing defect hidden behind missense or nonsense mutations is important for deciding appropriate therapeutic strategies for genetic surfactant disorders and other rare genetic diseases.11, 12, 13
References
Nogee LM, Dunbar AE 3rd, Wert SE, Askin F, Hamvas A, Whitsett JA : A mutation in the surfactant protein C gene associated with familial interstitial lung disease. N Engl J Med 2001; 344: 573–579.
Mulugeta S, Maguire JA, Newitt JL, Russo SJ, Kotorashvili A, Beers MF : Misfolded BRICHOS SP-C mutant proteins induce apoptosis via caspase-4 and cytochrome c-related mechanisms. Am J Physiol Lung Cell Mol Physiol 2007; 293: L720–L729.
Thurm T, Kaltenborn E, Kern S, Griese M, Zarbock R : SFTPC mutations cause SP-C degradation and aggregate formation without increasing ER stress. Eur J Clin Invest 2013; 43: 791–800.
Wang WJ, Mulugeta S, Russo SJ, Beers MF : Deletion of exon 4 from human surfactant protein C results in aggresome formation and generation of a dominant negative. J Cell Sci 2003; 116: 683–692.
Beers MF, Hawkins A, Maguire JA et al: A nonaggregating surfactant protein C mutant is misdirected to early endosomes and disrupts phospholipid recycling. Traffic 2011; 12: 1196–1210.
Stevens PA, Pettenazzo A, Brasch F et al: Nonspecific interstitial pneumonia, alveolar proteinosis, and abnormal proprotein trafficking resulting from a spontaneous mutation in the surfactant protein C gene. Pediatr Res 2005; 57: 89–98.
Litao MK, Hayes Jr D, Chiwane S, Nogee LM, Kurland G, Guglani L : A novel surfactant protein C gene mutation associated with progressive respiratory failure in infancy. Pediatr Pulmonol 2017; 52: 57–68.
Guillot L, Epaud R, Thouvenin G et al: New surfactant protein C gene mutations associated with diffuse lung disease. J Med Genet 2009; 46: 490–494.
Aissat A, de Becdelievre A, Golmard L et al: Combined computational-experimental analyses of CFTR exon strength uncover predictability of exon-skipping level. Hum Mutat 2013; 34: 873–881.
Shapiro MB, Senapathy P : RNA splice junctions of different classes of eukaryotes: sequence statistics and functional implications in gene expression. Nucleic Acids Res 1987; 15: 7155–7174.
Hinzpeter A, Aissat A, Sondo E et al: Alternative splicing at a NAGNAG acceptor site as a novel phenotype modifier. PLoS Genet 2010; 6: e1001153.
Perez-Carro R, Sanchez-Alcudia R, Perez B et al: Functional analysis and in vitro correction of splicing FAH mutations causing tyrosinemia type I. Clin Genet 2014; 86: 167–171.
Scotti MM, Swanson MS : RNA mis-splicing in disease. Nat Rev Genet 2016; 17: 19–32.
Acknowledgements
We thank Natacha Martin for her help during transcript analyses. This work was supported by public grants from the Institut National de la Santé et de la Recherche Médicale and Chancellerie des Universités de Paris.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Delestrain, C., Simon, S., Aissat, A. et al. Deciphering the mechanism of Q145H SFTPC mutation unmasks a splicing defect and explains the severity of the phenotype. Eur J Hum Genet 25, 779–782 (2017). https://doi.org/10.1038/ejhg.2017.36
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2017.36