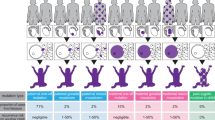
Abstract
In sporadic cases, a post-zygotic mutational event signifies a somatic mosaicism in the affected child only, which implies that these mutations affect only a portion of the body. Therefore siblings do not need follow-up. On the other hand, a pre-zygotic mutation transmitted by an unaffected mosaic parent implies recurrent risks in offspring. To better estimate the contribution of pre- and post-zygotic events, we analysed 124 consecutive bilateral retinoblastoma probands, carrying a heterozygous RB1 pathogenic variant and their unaffected, non-carrier parents. In order to evaluate somatic mosaicism in blood, the deleterious RB1 pathogenic variant identified in the proband, was searched for in the unaffected parents, using targeted deep sequencing. Observed recurrences, which represent an estimation of germline and somatic mosaicisms, were recorded and computed in the sibships. Deep sequencing revealed one mosaic-unaffected parent out of 124 tested couples, which provides an estimation of the maximal risk of recurrence, due to parental mosaicism, at 0.4%. Follow-up in the sibships showed one recurrence, providing a maximal recurrence risk, due to parental mosaicism, at 0.8%. Two different statistical strategies led to close estimates (0.4 and 0.8% risks) which appeared 266–533-fold higher, as compared with the general population. These recurrence estimates could be considered when counselling couples with retinoblastoma or diseases with a high de novo mutation rate.
Similar content being viewed by others
Introduction
Mosaicism is defined by the presence of different cell populations within the body and results from de novo, post-zygotic mutational events. Mosaicism has an important impact on the phenotype variability in first generation carriers but also on the recurrence risk and thus prenatal counselling.1 In sporadic cases, when unaffected parents are tested negative for the pathogenic variant identified in their affected child, it suggests that a de novo pathogenic variant has occurred. Unfortunately, as targeted testing is routinely performed using DNA extracted from blood, this negative result in both parents cannot exclude the possibility of parental mosaicism. With regards to prenatal counseling, it should therefore be determined whether the pathogenic variant is post-zygotic or pre-zygotic. A post-zygotic event signifies a somatic mosaicism in the affected child and as a result, parents and siblings are freed from genetic testing and clinical follow-up. On the other hand, a pre-zygotic pathogenic variant transmitted by an unaffected mosaic parent implies a recurrence risk in future offspring and thus prenatal diagnosis options might be discussed with the couple.
Retinoblastoma illustrates the difficulties raised by mosaics for geneticists. Retinoblastoma (Rb) [MIM 180200] is an embryonic neoplasm of retinal origin with an autosomal dominant mode of inheritance and is due to mutations in the RB1 tumour-suppressor gene. One or both eyes can be affected (unilateral Rb and bilateral Rb, respectively) and it should be emphasized that 90% of Rb cases are due to heterozygous de novo mutations whereas the remaining 10% are made of familial cases. More specifically, sporadic bilateral retinoblastoma stems from an early post-zygotic mutation in the affected child (ie no recurrence risk in siblings) or a pre-zygotic mutation in an unaffected parent (ie undetermined recurrence risk in siblings).2, 3, 4, 5 Consequently, it represents a highly relevant model to better estimate the relative contribution of pre- and post-zygotic events and thereby recurrence risks in offspring.
Patients and methods
Patients
Diagnosis of Rb was established on the basis of examinations by an ophthalmologist and a pediatrician and by histopathological criteria when the tumour was available. Rb patients were offered genetic counseling and individual written consent was obtained from all sampled individuals or their legal guardians. We included 124 consecutive bilateral Rb probands for whom blood samples from both unaffected parents were available. All parents also benefited from fundus examination and no sign of retinoblastoma or retinoma was found. To exclude false paternity, microsatellite analyses were performed in all families. Previous RB1 Sanger sequencing identified a heterozygous pathogenic variant in all Rb probands, but Sanger-targeted testing failed to identify this relevant variant in any of the unaffected parents. Overall, our analyses comprised the 124 sibships and included 248 siblings, that is, the 124 probands and their 124 sisters and/or brothers. For 75 sibships the third child (always unaffected) was not taken into account. The reason is that the ‘probability of having a second child affected knowing that the first is affected’ is different from ‘the probability of having a third child affected knowing that the first is affected’. Considering the second sibling as the prominent-and first- issue for parents, we did not consider the third sibling for analysis.
Firstly, and in order to evaluate somatic mosaicism in blood, the deleterious pathogenic variant identified in the proband was systematically searched for in the unaffected parents using targeted, high-sensitive deep sequencing. Secondly, observed recurrences for the sibships were recorded and computed to estimate germline mosaicism in the parents. Both approaches were then used to estimate recurrence risks to be used in genetic counseling.
Methods
DNAs were extracted from blood samples using the Quickgene 610-L automated system from FujiFilm (Courbevoie, France) according to the manufacturer’s instructions and calibrated to 50 ng/μl by UV spectrophotometric assay (Nanodrop, Thermo Fisher Scientific, Villebon sur Yvette, France). PCR amplicons targeting each point pathogenic variant identified in a proband were barcoded, pooled in equimolar ratio and libraries were prepared using the Library Builder (Life Technologies, Villebon sur Yvette, France) to obtain 300 bp DNA fragments flanked by adaptor and barcode sequences, allowing sequencing and sample identification, respectively. Libraries were then pooled and submitted to 10 PCR cycles in order to select and amplify relevant constructions, for example, DNA fragments with correct barcode and adaptor ligation. Amplified libraries were controlled for primer dimers and size range using LabChip devices (Caliper, Villepinte, France) and were then submitted to emulsion PCR with the Ion Xpress template kit using the Ion One Touch system (Life Technologies). Ion Sphere Particles were enriched using the E/S module and sequenced with an Ion Personal Genome Machine (PGM) in a 300 bp configuration run using a 318 chip (Life Technologies). Bioinformatic analysis was performed using NextGene software (Softgenetics, State College, PA, USA).6 In addition, two serial dilution assays were performed for defining the sensitivity of the method. Serial dilutions of two DNAs which were heterozygous for a substitution and a deletion, respectively, were mixed with wild-type DNA to mimic mutation levels of 50%, 25%, 12.5%, 6.25%, 4%, 2% and 1%. Experiments were repeated in two different runs. Serial dilution experiments demonstrated that mutations as low as 1% were correctly identified and quantitation was in accordance with theoretical, expected values. Moreover, 13 RB1 mosaic pathogenic variants previously identified by Sanger sequencing or denaturing high-performance liquid chromatography were correctly identified.
The statistical model to define a risk of occurence is based on the likelihood of the germline hypothesis H1 and the likelihood of the developmental hypothesis H2 (see the Result section for details). By definition, The likelihood of the germline hypothesis H1, considering the observed data, L(H1/dataobs) is equivalent to the probability PH1=P(dataobs/H1) to have the observed data if the H1 hypothesis is true. Because H1 is related to the proportion p1, the likelihood of H1 is equivalent to the likelihood of p1 L(H1/dataobs)=L(p1/dataobs). In addition, the probability PH1 to have 122 children not heterozygous and one second child heterozygous can be modelized by a binomial law B(n=123, p1), with p1 being the probability of success (ie, the probability to have a second child heterozygous), which leads to the equation provided in full in the Result section (the population providing the 123 families is supposed to be infinite). The derivative of PH1 provides the maximum likelihood of p1, which is equal to k/n, with k being the number of success. According to the observed data, k=1. Thus, the maximum likelihood of H1 is obtained for p1=1/123=0.008. This reasoning is also valid for the H2 hypothesis and the p2 proportion, except that the binomial law is B(n=123, p2=1.5e−5), and that this modelisation is a constant. Thus, the maximum likelihood ratio L(H1/dataobs)/L(H2/dataobs)=PH1/PH2 is obtained for p1=1/123=0.008.
Results
Targeted deep sequencing data obtained in the blood of the unaffected parents showed a minimum × 1000 depth of coverage for all amplicons. Only one of the unaffected parents, out of 124 tested couples, carried the pathogenic variant identified in her affected child and the degree of mosaicism of the mutant allele was 11% in her leucocyte DNA. This indicates a 0.8% (1/124) risk of mosaicism in one of the two parents, when the couple has a first child affected by a bilateral Rb. This also implicates a theoretical 0.4% (0.8%/2) maximum risk of recurrence (the maximum risk corresponding to the presence of the mutation in all germline cells of the mosaic mutation carrier).
Since deep sequencing already proved mosaicism in one unaffected parent (see above), this family was excluded from further calculations. Consequently, the modelisation approach included 123 sibships. In these 123 families, one retinoblastoma recurrence was observed during follow-up in the sibships. Both affected siblings carried the same RB1 pathogenic variant but we were unable to detect blood mosaicism in any of the parents, suggesting that this de novo mutation should have arisen in the germline only, after its separation from the soma. In other words, in 1 family out of 123, the second child is heterozygous in the leucocytes, while in the other 122 families, the second child has no RB1 pathogenic variant detected in the leucocytes. From these observations, a second recurrence risk was estimated. The model is based on two exclusive hypotheses that explain the absence of pathogenic variant in the leucocytes of the two parents: (1) in the germline hypothesis (H1), one of the two parents has the RB1 pathogenic variant in the germline cells (the unlikely hypothesis of two parents with an RB1 pathogenic variant in the germline cells was not considered); (2) in the developmental hypothesis (H2) the elder children have been mutated in the RB1 gene during embryonic development. The likelihood of the germline hypothesis H1 can be defined as the probability PH1 to have 122 children not heterozygous and one second child heterozygous (ie, what we observed), considering that H1 is true. This probability is written as

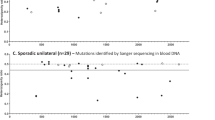
where PH1 is the likelihood of H1 and p1 is the probability to have a second child heterozygous, which is equivalent to the proportion of mutated gametes in the unaffected parent (0≤p1≤0.5). For instance, p1=0.5 means that all the germline cells of the parent are heterozygous. Results of the PH1 probability value according to the parental germline mosaicism is shown in Figure 1. According to the observed data, The likelihood of the germline hypothesis H1 is close to zero when p1=0.5 (PH1≈0) and it reaches a maximum for p1=0.008.
Likelihood of the germline hypothesis according to the observed data. Y-axis, left: probability PH1 that the germline hypothesis fit the observed data, which depends on the proportion p1 of mutated gametes in one of the two parents. Y-axis, right: ratio of the two PH1 and PH2 (developmental hypothesis) probabilities. The curve is based on our observations and the proposed hypotheses (see text for details).
Following the reasoning used for the germline hypothesis, the likelihood of the developmental hypothesis H2 is the probability PH2 to have a second child heterozygous considering that H2 is true, except that PH2 is related to the risk of a de novo RB1 pathogenic variant. The frequency of heterozygous people, carrying a RB1 pathogenic variant, is estimated between 1/15 000 and 1/20 000 in the general population.5, 7 Among the heterozygous population, 30% have bilateral Rb with no familial history, which provides a probability p2 of being heterozygous due to a de novo pathogenic variant and developing a bilateral Rb equal to 1/20 000 × 0.3=1.5 × 10−5. Hence, according to the developmental hypothesis, the probability to observe one recurrence in the 123 sibships is

Then, we compared the two hypotheses. As shown in Figure 1, the interval of p1 in favour of the germline hypothesis H1 is ]0–0.066]. In addition, the likelihood of the germline hypothesis is 200-fold higher than the likelihood of the developmental hypothesis (given by the ratio PH1/PH2) when p1=0.8%. According to our observed data, this indicates that the risk of recurrence can reach p1=0.8% in the case of germline mosaicism.
Discussion
Data on parental mosaicism are scarce despite their importance in recurrence risks of genetic diseases in sibships. In sporadic cases, mosaicism in unaffected parents has to be taken into account to discuss recurrence risks in offspring. A recent study estimated the proportion of low-level deleterious copy number variant mosaicism in blood over 4% and considered low-level mosaicism in parents as an under recognized cause of disease.8 Unfortunately, and as opposed to chromosomal mosaicism, there are limited data on the role of mosaicism of single gene mutations. A theoretical risk of 5% for mosaic parents undetected from blood analysis was reported for neurofibromatosis 2.9 With regards to retinoblastoma, two previous studies based on single strand conformation polymorphism, Sanger sequencing or allele-specific PCR, suggested that germline mosaicism in an unaffected parent of a sporadic Rb proband is rare i.e. 0.7%10 to 1.3%.4
To shed light on this issue, we specifically designed a study to address the recurrence risk in unaffected parents tested negative for the mutation identified in their affected sibling. We took advantage of deep sequencing which increases the sensitivity of mutation detection, increases the number of mosaics detected11 and as a result, may refine recurrence risk rates. We added a complementary analysis of recurrence based on a new predictive model. Both approaches provided close estimates, that is, 0.4 and 0.8%, respectively. That said, such strategies have their limitations (i) demonstrating mosaicism in blood does not necessarily imply a recurrence event ; and (ii) on the other hand, our data show that the hematopoietic lineage is not the most relevant tissue to study, as no mosaicism was found in the couple where a recurrence occurred. In any case, our predictive model is based on observed recurrences and not on tissue analyses, consequently tissue origin is not an issue. Overall, and taking into account that the recurrence risk due to a de novo pathogenic variant is p2=1.5 × 10−5, we conclude that the recurrence risk for unaffected parents of bilateral Rb sporadic cases, due to mosaicism, can be estimated as 266–533-fold higher, as compared with the general population (0.004/p2 to 0.008/p2). These values justify surveillance protocols in use for retinoblastoma5 and provides geneticists with reliable recurrence risks that could be helpful in counseling couples for prenatal diagnostic options. We believe this evaluation method or even this recurrence risk could be considered in other diseases with a high de novo mutation rate and should be used for genetic counseling and especially for improved prenatal options.
References
Castera L, Gauthier-Villars M, Dehainault C et al: Mosaicism in clinical practice exemplified by prenatal diagnosis in retinoblastoma. Prenat Diagn 2011; 31: 1106–1108.
Lohmann D, Gallie B, Dommering C, Gauthier-Villars M : Clinical utility gene card for: retinoblastoma. Eur J Hum Genet, e-pub ahead of print 19 March 2011 doi:10.1038/ejhg.2010.200.
Price EA, Price K, Kolkiewicz K et al: Spectrum of RB1 mutations identified in 403 retinoblastoma patients. J Med Genet 2014; 51: 208–214.
Sippel KC, Fraioli RE, Smith GD et al: Frequency of somatic and germ-line mosaicism in retinoblastoma: implications for genetic counseling. Am J Hum Genet 1998; 62: 610–619.
Houdayer C, Gauthier-Villars M, Castéra L, Desjardins L, Doz F, Stoppa-Lyonnet D 2012, Retinoblastoma - Genetic Counseling and Molecular Diagnosis. Available from: http://www.intechopen.com/books/retinoblastoma-an-update-on-clinical-genetic-counseling-epidemiology-and-molecular-tumor-biology/retinoblastoma-genetic-counseling-protocols-and-molecular-diagnostic-protocols-.
Tarabeux J, Zeitouni B, Moncoutier V et al: Streamlined ion torrent PGM-based diagnostics: BRCA1 and BRCA2 genes as a model. Eur J Hum Genet 2015; 22: 535–541.
Vogel F : Genetics of retinoblastoma. Hum Genet 1979; 52: 1–54.
Campbell IM, Yuan B, Robberecht C et al: Parental somatic mosaicism is underrecognized and influences recurrence risk of genomic disorders. Am J Hum Genet 2014; 95: 173–182.
Evans DG, Wallace A : An update on age related mosaic and offspring risk in neurofibromatosis 2 (NF2). J Med Genet 2009; 46: 792.
Rushlow D, Piovesan B, Zhang K et al: Detection of mosaic RB1 mutations in families with retinoblastoma. Hum Mutat 2009; 30: 842–851.
Chen Z, Moran K, Richards-Yutz J et al: Enhanced sensitivity for detection of low-level germline mosaic RB1 mutations in sporadic retinoblastoma cases using deep semiconductor sequencing. Hum Mutat 2014; 35: 384–391.
Acknowledgements
We thank Allyson Holmes for critical reading of the manuscript, Elsa Bernard for technical advices and Camille Benoist for helpful discussion. This work was supported by RETINOSTOP and Programme Incitatif et Coopératif ‘Rétinoblastome’ Institut Curie.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Dehainault, C., Golmard, L., Millot, G. et al. Mosaicism and prenatal diagnosis options: insights from retinoblastoma. Eur J Hum Genet 25, 381–383 (2017). https://doi.org/10.1038/ejhg.2016.174
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2016.174
This article is cited by
-
Genetics in ophthalmology: molecular blueprints of retinoblastoma
Human Genomics (2023)
-
Molecular tumor testing in patients with Lynch-like syndrome reveals a de novo mosaic variant of a mismatch repair gene transmitted to offspring
European Journal of Human Genetics (2020)
-
Frequency of low-level and high-level mosaicism in sporadic retinoblastoma: genotype–phenotype relationships
Journal of Human Genetics (2020)
-
Parent-of-origin effect of hypomorphic pathogenic variants and somatic mosaicism impact on phenotypic expression of retinoblastoma
European Journal of Human Genetics (2018)