Abstract
We describe a three-generation Norwegian family with a novel X-linked intellectual disability (XLID) syndrome characterized by subtle facial dysmorphism, autism and severe feeding problems. By exome sequencing we detected a rare missense variant (c.1067A>G, p.(Tyr356Cys)) in the RLIM gene, in two affected male second cousins. Sanger sequencing confirmed the presence of the variant in the four affected males (none of whom were siblings) and in three mothers available for testing. The variant was not present in 100 normal Norwegian controls, has not been reported in variant databases and is deleterious according to in silico prediction tools. The clinical phenotype and the variant co-segregate, yielding a LOD score of 3.0 for linkage to the shared region (36.09 Mb), which contains 242 genes. No other shared rare variants on the X chromosome were detected in the two affected exome-sequenced individuals, and all female carriers had an extremely skewed X-chromosome inactivation pattern. RLIM encodes RING zinc finger protein 12 (RNF12), an ubiquitin ligase that is essential for X inactivation in mice and that acts as a co-regulator of a range of transcription factors, particularly those containing a LIM homeodomain. Tyrosine in position 356 in RNF12 is located within a highly conserved domain essential for binding such transcription factors. Expression of RNF12 is widespread during embryogenesis, and is particularly high in the outer layers of the cerebral cortex. Functional studies are needed to prove a definite causal relationship between the variant and the phenotype. Subsequent reports may confirm a role for RLIM variants in patients with XLID.
Similar content being viewed by others
Introduction
Intellectual disability (ID) affects 2–3% of the population with an estimated male excess of 30%.1 This gender difference is explained in part by the fact that an estimated 5–10% of ID in males is caused by pathogenic variants in X-linked genes.1, 2 A genetic cause has been identified for about 80 of the >150 known syndromic forms of X-linked ID (XLID).1 Variants in the same gene may give rise to syndromic or non-syndromic ID, and dysmorphic features in syndromic XLID may be subtle and variable even within a family. Autism is associated with at least 8% of the XLID types with a known genetic cause,1 and the frequent overlap of autism and ID is seen also with microdeletions/duplications.3 This observation probably reflects that these neurodevelopmental phenotypes often share common pathophysiological mechanisms during embryogenesis.4
We investigated a family with four affected males in three generations with a variable degree of ID, dysmorphic facial features, severe initial feeding difficulties and autism. The cases were related to each other exclusively through non-affected females, and no known XLID syndrome was diagnosed initially. We therefore performed exome sequencing in two of the affected males, and identified only one shared rare, novel missense variant in the RLIM gene on the X chromosome.
Materials and Methods
Subjects
The family structure is shown in Figure 1. IV-1, IV-2, IV-3, IV-4 and III-3 were examined by two of us (ET and TP). III-4 provided information about IV-5, IV-6 and III-5, as did II-4 for II-1 and II-2. The study was approved by the Western Norway regional ethics committee (application # 2011/2453). Written informed consent (including consent for publication of the pedigree and photographs) was obtained from all the individuals including non-affected adults. Parents consented on behalf of children under 16 years. Consent for participation for II-1 was obtained from his legal guardian (II-4).
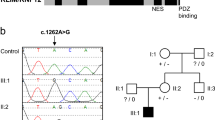
Overview of the pedigree. Black symbols denote affected individuals and symbols with a dot denote carriers. X-inactivation patterns (f.ex. 100:0) are shown underneath the identifier for individuals II:3, II:4, II:5, III:1, III:2 and III:4; for individuals II-5 and III-2 the analysis was performed twice. All individuals except I:1 and I:2 (*) were genotyped for the variant c.1067A>G in RLIM.
Whole-exome sequencing
DNA was extracted from blood using the QIAsymphony system (Qiagen, Hilden, Germany) and exome sequencing was performed using the SOLID 5500xl platform (Life Technologies, Waltham, MA, USA). Exonic sequences were enriched using the SureSelect Human All Exon v4 kit (Agilent Technologies, Santa Clara, CA, USA), which targets ~21 000 human genes and covers 51-Mb of genomic sequence. Color space reads were mapped to the hg19 reference genome using the LifeScope v2.5 software (Life Technologies), yielding on average 114 million mapped reads per exome (50- bp single end), with ~83% mapped reads on target or near target. Average depth of coverage within targets was 81.2 (IV-1) and 86.9 (IV-4), and 82% and 83% respectively, of the target bases were covered >20 times. SNPs were called using the LifeScope diBayes algorithm with high call stringency. Small insertions and deletions were detected using the LifeScope Small Indel Tool. Variants were annotated using a custom pipeline.5 All called variants on the X chromosome were filtered for known variants in dbSNP, intronic and UTR variants, synonymous variants and variants in our in-house database. Variants with<5 reads were excluded from further analysis.
Sanger sequencing
Verification of the shared variant was performed for all living individuals shown in the pedigree (Figure 1) using targeted Sanger sequencing according to the standard procedures.
SNP arrays
Affymetrix Cytoscan HD containing 2.7 million oligonucleotides was performed for the four affected and six unaffected males, and common hemizygous region(s) were determined using a combination of an in-house tool developed in the R program package, and manual comparison of genotypes in excel.
X-chromosome inactivation
X-chromosome inactivation analysis of DNA extracted from the blood was performed using methylation-sensitive fragment analysis of a polymorphic CAG repeat in the first exon of the androgen receptor gene on Xq11.2-q12.6 DNA was amplified and analyzed by capillary electrophoresis and digested with the methylation-sensitive HpaII endonuclease. The analysis was informative in all the six females who were tested.
Cases
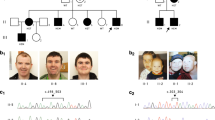
The four affected males (Figure 1), ages 5 ½–71 years have mild to moderate/severe ID. Expressive language development was particularly delayed. Limited phenotypic and historical data are available for the two older affected men. On the basis of the information from relatives, and in the case of III-3 also clinical examination, we concluded that it was highly likely that these two adults had the same disorder as IV-1 and IV-4, before targeted analysis of RLIM. Clinical features and results of supplementary investigations are summarized in Table 1. Facial features include a broad forehead, hypertelorism, fine hair with a receding hairline (starting in childhood), a square face and a rather square jaw (Figure 2). However, dysmorphic features may be quite subtle, as demonstrated by the childhood picture of III-3 (Figure 2e). Measurements at birth, where available, were within the normal range and microcephaly has not developed postnatally. Initial feeding difficulties in the two younger boys were strikingly severe given the degree of their developmental delay. They continue to have difficulties with chewing and swallowing. A characteristically stiff gait was evident in the youngest individuals. Behavioral issues have been noted in all affected individuals, with a preference for repetitive activities and fixed routines. Autism or autistic spectrum disorder was diagnosed in the two young boys and was suspected in the 45 year old man (III-3).
Results
Identification of one shared rare variant in two affected boys by exome sequencing
Exome sequencing revealed a single rare variant on the X chromosome in the genes FANCB, RLIM, ACSL4 and LRCH2 in individual IV-1 and a rare variant in RLIM and NHS in individual IV-4. Thus, these two affected boys shared only one rare variant in a single gene on the X chromosome, c.1067A>G in RLIM (NM_183353.2), g.73812083 bp from pter. The variant was not detected in samples from 100 healthy Norwegian blood donors and has not been reported in dbSNP or the NHLBI Exome Variant Server. The c.1067A>G change predicts substitution of a tyrosine residue with a cysteine, p.(Tyr356Cys). Tyrosine 356 is located within a highly conserved stretch of 100 amino acids7 (phyloP score 4.536; Figure 3). The detected variant is predicted to be damaging by PolyPhen2 (0,98), CADD (PHRED 11.83), SIFT (0,01), Align GVGD (Class C65) and MutationTaster (0,99). The variant has been reported to the ClinVar database (accession number SCV000172707, http://www.ncbi.nlm.nih.gov/clinvar/).
Alignment of the LIM binding domain of RLIM. The position of the variant c.1067A>G is marked with a yellow box, and is predicted to result in the substitution of the amino acid tyrosine in position 356 by cysteine: p.(Tyr356Cys) or p.(Y356C). Tyr (Y) 356 is located in a highly conserved region of the RNF12 protein, essential for binding to LIM-containing proteins,7 as shown by Ostendorff et al.7
The c.1067A>G missense variant co-segregates with ID in the family
Sanger sequencing of RLIM confirmed the presence of c.1067A>G in individuals IV-1 and IV-4. Further genotyping revealed that the two additional affected males (II-1 and III-3) were hemizygous for c.1067A>G and that the mothers of the three youngest affected males (II-4, III-1 and III-4), as well as the maternal grandmother (II-3) of IV-1, were heterozygous. The mother of II-1 is deceased. Six healthy males, (II-2, III-5, IV-2, IV-3, IV-5 and IV-6), and the mothers of III-5 and IV-2/IV-3 did not harbor the variant. In marked contrast to non-carriers (II-5 and III-2), all the heterozygous females had an extremely skewed X-inactivation pattern. The results of 1 or 2 separate assays are denoted under each individual in Figure 1. The variant segregated perfectly with the phenotype, and linkage analysis yielded a significant LOD score of 3.0 between the phenotype and the region shared by the four affected individuals.
SNP arrays
Haplotype analysis using SNP arrays in the four affected and six healthy males (II-2, III-5, IV-2, IV-3, IV-5 and IV-6) revealed that the hemizygous males share a region spanning 36.09 Mb (g.49486929 to g.85574677 bp from pter) and containing 242 annotated genes. Variants affecting 26 of these genes have been reported as causative for ID (syndromic and/or non-syndromic; OMIM, http://omim.org/). For these genes, median coverage was 46X and 49X (range 10–89 and 10–97) in the samples from individuals IV-1 and IV-4, respectively.
Discussion
We here describe a new XLID syndrome in a family with four affected males aged 5–71 years, who share a rare hemizygous variant in the RLIM gene at chromosome Xq13. Syndromic features include eating difficulties (in the three youngest individuals), autism/autistic spectrum disorder (confirmed in IV-1 and IV-4, and suspected in III-3) and subtle dysmorphic facial features, such as a broad forehead, fine hair with a receding hairline bitemporally and hypertelorism (IV-1 and IV-4). Facial features can also be unremarkable in childhood, as in individual III-3.
The skewed X-inactivation pattern observed in unaffected heterozygous carriers is in keeping with X-linked recessive inheritance. The missense variant c.1067A>G segregates with the disorder with a LOD score of 3.0. Thus, there is only a 1 in a 1000 chance that this 36.09-Mb shared region on the X chromosome does not contain the causative variant. The possibility that c.1067A>G in RLIM is a neutral variant linked to an undetected causal variant in RLIM or in another gene is quite unlikely, but cannot be excluded. No other rare variants in the shared region on Xq13 were observed, and the affected subjects did not share any other segments on the X chromosome. Coverage across the exome varies and not all exonic variants are reliably detected.8 In the analyzed samples, 18% (IV-1) and 17% (IV-4), respectively, of target bases were covered <20 times, so rare variants within targets with low coverage could have escaped detection.
RLIM encodes an ubiquitin protein ligase RNF12,9, 10 which is widely expressed during embryogenesis.11 It is crucial for normal embryonic development in some species12 and for normal X inactivation in mice.9, 13 RNF12 mainly localizes to the cell nucleus,11 where it regulates the levels of many proteins, including CLIM, LMO, HDAC2, TRF1, SMAD7 and REX1, by proteasomal degradation.7, 9, 12, 14 Thus, it modulates the activity of transcription factors, in particular those containing a LIM homeodomain,15 and the formation of transcriptional multiprotein complexes. The ubiquitin system has a critical role in the remodeling of synapses and neurodevelopment,16 and variants/CNV's in various E3 ubiquitin ligases may cause autism,17, 18, 19 ID19 or both.
The variant c.1067A>G is predicted to alter an amino acid (p.(Tyr356Cys)) in a stretch of 100 highly conserved amino acids crucial for binding to LIM domains (Figure 3), as shown by Ostendorff et al.7 Thus, protein–protein interactions could be affected by the variant. The most studied role of RNF12 is its essential function in the first wave of paternally imprinted X-chromosome inactivation in mice.9, 13 All female carriers available for testing in our family have a completely skewed X-chromosome inactivation pattern in contrast to non-carriers. This could indicate that RLIM is essential for normal X-chromosome inactivation in humans as well. However, X-chromosome inactivation is not imprinted in humans. Thus, in these female carriers, impairment of the important unknown roles of RNF12 in neuronal development and function20 could result in selection against the cell line in which the X chromosome harboring the variant is active. RLIM is expressed in specific regions of the neural tube in rat embryos,20 including the superficial layer of the developing telencephalon.21 In humans, RLIM is highly expressed in the prefrontal cortex throughout life (data from the braincloud database at http://braincloud.jhmi.edu).
In conclusion, we here report a missense variant in the RLIM gene encoding the ubiquitin ligase RNF12, in four affected males with ID from a three-generation family. Additional features include autistic spectrum disorder/autism, eating difficulties and subtle dysmorphic features. The detected variant likely disrupts an essential function of RLIM in neurodevelopment and raises the possibility of important, previously unknown roles for this ubiquitin ligase.
Note added to proof
Since this manuscript was accepted, three further families with X-linked ID and different missense mutations in RLIM have been reported by Kalscheuer et al (Mol Psychiatry; in press), confirming that RLIM missense mutations can cause XLID.
Accession codes
References
Lubs HA, Stevenson RE, Schwartz CE : Fragile X and X-linked intellectual disability: four decades of discovery. Am J Hum Genet 2012; 90: 579–590.
Jacquemont S, Coe BP, Hersch M et al: A higher mutational burden in females supports a "female protective model" in neurodevelopmental disorders. Am J Hum Genet 2014; 94: 415–425.
Girirajan S, Brkanac Z, Coe BP et al: Relative burden of large CNVs on a range of neurodevelopmental phenotypes. PLoS Genet 2011; 7: e1002334.
Parikshak NN, Luo R, Zhang A et al: Integrative functional genomic analyses implicate specific molecular pathways and circuits in autism. Cell 2013; 155: 1008–1021.
Gilissen C, Arts HH, Hoischen A et al: Exome sequencing identifies WDR35 variants involved in Sensenbrenner syndrome. Am J Hum Genet 2010; 87: 418–423.
Allen RC, Zoghbi HY, Moseley AB, Rosenblatt HM, Belmont JW : Methylation of HpaII and HhaI sites near the polymorphic CAG repeat in the human androgen-receptor gene correlates with X chromosome inactivation. Am J Hum Genet 1992; 51: 1229–1239.
Ostendorff HP, Peirano RI, Peters MA et al: Ubiquitination-dependent cofactor exchange on LIM homeodomain transcription factors. Nature 2002; 416: 99–103.
Gilissen C, Hoischen A, Brunner HG, Veltman JA : Disease gene identification strategies for exome sequencing. Eur J Hum Genet 2012; 20: 490–497.
Gontan C, Achame EM, Demmers J et al: RNF12 initiates X-chromosome inactivation by targeting REX1 for degradation. Nature 2012; 485: 386–390.
Ostendorff HP, Bossenz M, Mincheva A et al: Functional characterization of the gene encoding RLIM, the corepressor of LIM homeodomain factors. Genomics 2000; 69: 120–130.
Jiao B, Taniguchi-Ishigaki N, Gungor C et al: Functional activity of RLIM/Rnf12 is regulated by phosphorylation-dependent nucleocytoplasmic shuttling. Mol Biol Cell 2013; 24: 3085–3096.
Zhang L, Huang H, Zhou F et al: RNF12 controls embryonic stem cell fate and morphogenesis in zebrafish embryos by targeting Smad7 for degradation. Mol Cell 2012; 46: 650–661.
Shin J, Wallingford MC, Gallant J et al: RLIM is dispensable for X-chromosome inactivation in the mouse embryonic epiblast. Nature 2014; 511: 86–89.
Her YR, Chung IK : Ubiquitin Ligase RLIM Modulates Telomere Length Homeostasis through a Proteolysis of TRF1. J Biol Chem 2009; 284: 8557–8566.
Hunter CS, Rhodes SJ : LIM-homeodomain genes in mammalian development and human disease. Mol Biol Rep 2005; 32: 67–77.
Bingol B, Sheng M : Deconstruction for reconstruction: the role of proteolysis in neural plasticity and disease. Neuron 2011; 69: 22–32.
O'Roak BJ, Vives L, Girirajan S et al: Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations. Nature 2012; 485: 246–250.
Sanders SJ, Murtha MT, Gupta AR et al: De novo mutations revealed by whole-exome sequencing are strongly associated with autism. Nature 2012; 485: 237–241.
Glessner JT, Wang K, Cai G et al: Autism genome-wide copy number variation reveals ubiquitin and neuronal genes. Nature 2009; 459: 569–573.
Ostendorff HP, Tursun B, Cornils K et al: Dynamic expression of LIM cofactors in the developing mouse neural tube. Dev Dyn 2006; 235: 786–791.
Furuyama T, Inagaki S, Iwahashi Y, Wanaka A, Tohyama M : Localization of mRNAs for Rlim-1, the rat Xlim-1 homolog, in the developing rat brain. Brain Res Mol Brain Res 1996; 36: 152–156.
Acknowledgements
We are especially grateful to the family for participating in this study. Technical assistance from Hilde Rusaas and contributions from Bjørn Ivar Haukanes, Ketil Heimdal, Yngve Sejersted and Tomasz Stokowy were highly appreciated. The study has been supported by a grant from Bergen Medical Research Foundation (BMFS) and The University of Bergen.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Tønne, E., Holdhus, R., Stansberg, C. et al. Syndromic X-linked intellectual disability segregating with a missense variant in RLIM. Eur J Hum Genet 23, 1652–1656 (2015). https://doi.org/10.1038/ejhg.2015.30
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2015.30