Abstract
The concept of interferon-γ (IFN-γ) having a central role in cell-mediated immune defence to Mycobacterium tuberculosis has long been proposed. Observations made through early candidate gene studies of constituents of the IFN-γ pathway have identified moderately associated variants associated with resistance or susceptibility to tuberculosis (TB). By analysing 20 major genes whose proteins contribute to IFN-γ signalling we have assessed a large fraction of the variability in genes that might contribute to susceptibility to TB. Genetic variants were identified by sequencing the promoter regions and all exons of IFNG, IFNGR1, IFNGR2, IRF1, IL12A, IL12B, IL12RB1, IL12RB2, IL23A, IL23R, IL27, EBI3, IL27RA, IL6ST, SOCS1, STAT1, STAT4, JAK2, TYK2 and TBX21 in 69 DNA samples from Ghana. In addition, we screened all exons of IFNGR1 in a Ghanaian study group comprising 1999 TB cases and 2589 controls by high-resolution melting point analysis. The fine-mapping approach allows for a detailed screening of all variants, common and rare. Statistical comparisons of cases and controls, however, did not yield significant results after correction for multiple testing with any of the 246 variants selected for genotyping in this investigation. Gene-wise haplotype tests and analysis of rare variants did not reveal any significant association with susceptibility to TB in our investigation as well. Although this analysis was applied on a plausible set of IFN-γ pathway genes in the largest African TB cohort available so far, the lack of significant results challenges the view that genetic marker of the IFN-γ pathway have an important impact on susceptibility to TB.
Similar content being viewed by others
Introduction
Tuberculosis (TB) remains a major threat to human health. It is estimated that M. tuberculosis affects one third of the world’s population. The host defence against the pathogen is essentially determined by a cell-mediated immune response.1 On activation, the primary target cells for M. tuberculosis, human alveolar macrophages, are able to mount an antimicrobial programme enabling them to kill the intracellular pathogen. Studies showing that monozygotic twins have a higher risk of developing active TB compared with dizygotic twins indicate a central role of host genetic factors in susceptibility to TB.2 This is also emphasized by significant findings of studies attributing an important impact of several candidate genes on TB susceptibility or resistance,3 and more recently by genome-wide association studies (GWASs) in TB.4, 5
An essential mediator of macrophage activation is the proinflammatory cytokine interferon-γ (IFN-γ), which upregulates a large number of genes and their corresponding proteins that are needed to provide an effective anti-mycobacterial response. The importance of this cytokine and its signalling pathway in TB is underlined by studies of IFN-γ-deficient mice, which, when infected with M. tuberculosis, succumb early to the disease.6 In addition, rare deleterious mutations in genes of the pathway were found associated with increased susceptibility to mostly weakly virulent, non-tuberculous mycobacteria (NTM), and also to a small number of infections with M. tuberculosis.7 Together, these findings led to the view that human variability of genes of the IFN-γ pathway might influence susceptibility to infection with M. tuberculosis.
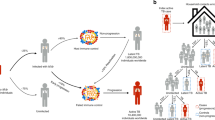
Recognition of M. tuberculosis by human macrophages results in the release of a multitude of cytokines, including the interleukins (ILs)-12, IL-23 and IL-27.7 These immunoregulatory mediators prime and activate Th1 cells through binding to their complementary receptors. Components of the JAK-STAT signalling pathway, such as the signal transducers and activators of transcription 1 and 4 (STAT1/4), the Janus kinase 2 (JAK2) and the tyrosine kinase 2 (TYK2) are important for intracellular IFN-γ signal transduction in Th1 cells.7 Signal transduction also involves the transcription factor T-box 21 (TBX21), which was shown to control expression of IFN-γ in Th1 cells.8 IFN-γ secreted by Th1 cells activates macrophages via binding to its heterodimeric receptor, which consists of two chains, IFN-γ receptor 1 and 2 (IFNGR1/2). Signal transduction in macrophages is then mediated by a cascade of signals, again involving JAK2, STAT1 and TYK2. Additional molecules modulating IFN-γ signalling in macrophages are the transcription activator IFN regulatory factor 1 (IRF1) and the negative regulator of IFN-γ production, the suppressor of cytokine signalling 1 (SOCS1).9, 10 Activation of macrophages by IFN-γ may induce the production of reactive oxygen and nitrogen intermediates, thus creating an unfavourable environment for intracellular mycobacteria.
Previous candidate gene studies have identified associations with single-nucleotide polymorphisms (SNPs) of the genes IFNG, IFNGR1, IL12A, IL12B, IL12RB1 and STAT4 in TB susceptibility.11, 12, 13, 14, 15, 16 However, small numbers only of SNPs were analysed, resulting in weak-to-moderate associations in most of these studies. Many of these studies were underpowered for the detection of significant effects.
Here we present the results of a TB case–control association study including 20 essential genes of the IFN-γ signalling pathway. Variants of IFNG, IFNGR1, IFNGR2, IRF1, IL12A, IL12B, IL12RB1, IL12RB2, IL23A, IL23R, IL27, EBI3, IL27RA, IL6ST, SOCS1, STAT1, STAT4, JAK2, TYK2 and TBX21 were analysed. Promoters and exonic regions of these genes were sequenced in 23 TB cases and 46 controls from Ghana, West Africa. With regard to the particular relevance of IFNGR1 we focused also on all variants of this gene by screening the entire study group applying high-resolution melting point (HRM) analyses. In total, 246 variants were genotyped and tested for association with susceptibility or resistance to TB in 1999 HIV-negative patients with pulmonary TB and 2589 exposed, healthy controls.
Materials and methods
Study group
The study design and the enrolment procedure have been described previously.17 In brief, the two major Ghanaian teaching hospitals in Accra and Kumasi, and additional hospitals and polyclinics and regional district hospitals were selected as study sites. We recruited 1999 HIV-negative individuals with smear- and/or culture-positive pulmonary TB as cases. As controls, 2589 individuals were selected. There was no significant difference in the proportion of ethnic groups between cases and controls. The ethnic groups included Akan, Ga-Adangbe, Ewe and groups from northern Ghana (Northerners). Phenotype characterization was based on a detailed documentation of the medical history of cases using structured questionnaires, two independent examinations of non-induced sputum specimens, serological determination of the HIV status, culturing and molecular differentiation of phylogenetic lineages of mycobacterial clades and posterior–anterior chest X-rays. An alternate test system was applied to confirm positive HIV test results. TB patients were enroled into the DOTS programme (Directly Observed Treatment Short-Course strategy) organized by the Ghanaian National Tuberculosis Programme.
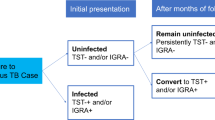
The medical history of the control group was obtained and a clinical examination was performed. Chest X-rays did not reveal any signs of actual or past pulmonary TB. In addition, a tuberculin skin test (TST, Tuberculin Test PPD Mérieux, bioMérieux, Nürtingen, Germany) was performed.
Ethical approval of the study design was obtained by the Committee on Human Research, Publications and Ethics, College of Health Sciences, Kwame Nkrumah University of Science and Technology, Kumasi, Ghana, and the Ethics Committee of the Ghana Health Services, Accra, Ghana. Informed consent was provided by study participants either by signature or, in case of illiteracy, by thumbprint in the presence of a witness. The aims of the study were explained in detail before blood samples were taken.
Sequencing and genetic analyses of variants selected for genotyping
Genomic DNA was extracted from total venous blood using the AGOWA mag Maxi DNA Isolation Kit (Macherey & Nagel, Düren, Germany) according to the instructions of the manufacturer. In total, DNA of 69 individuals was sequenced for 20 genes (IFNG, IFNGR1, IFNGR2, IRF1, IL12A, IL12B, IL12RB1, IL12RB2, IL23A, IL23R, IL27, EBI3, IL27RA, IL6ST, SOCS1, STAT1, STAT4, JAK2, TYK2 and TBX21). Sequence analyses were performed from 23 TB patients, 23 TST-positive and 23 TST-negative controls, allowing to identify variants with allele frequencies >2.2% in each of the subgroups. For this purpose we designed PCR primers covering the exonic regions as well as the promoter and the 3′-UTR of each gene with the PRIMER3 software (http://primer3.ut.ee) using the GenBank reference sequences of each gene. SNPs identified by DNA sequencing in only one sample of the study group were confirmed in independent PCR and sequencing reactions. Markers selected for genotyping in the entire study group included all non-synonymous variants, variants located in splice-site junctions and those identified in known or putative transcription factor bindings sites of the promoter. In addition, variants were analysed exhibiting obvious differences after sequencing in genotype frequencies between TB cases and TST-positive or -negative controls. All variants identified in this study were submitted to the dbSNP database (http://www.ncbi.nlm.nih.gov/SNP/). In order to capture common haplotypes and to assess relevant haplotype-related risks, additional tagging SNPs were selected by the Haploview software (http://www.broadinstitute.org/scientific-community/science/programs/medical-and-population-genetics/haploview/haploview) at r2 thresholds of 0.5 and minor allele frequencies of 10%. Variants were either analysed with the SNPlex genotyping system (Applied Biosystems, Foster City, CA, USA) or with in-house assays using the fluorescence resonance energy transfer (LightTyper; Roche Diagnostics, Mannheim, Germany) with dynamic allele-specific hybridization. A list of all primers that were used in the reactions can be obtained on request.
HRM-PCR of IFNGR1
In this study, a special focus was laid on the analysis of the IFNGR1 gene, as deleterious variants of this gene had been found to be associated with increased susceptibility to infections with NTM.18 Therefore, we screened the coding regions as well as the promoter and 3′-UTR regions of IFNGR1 in all samples of this study by HRM analyses using the LightCycler 480 device (Roche Diagnostics) as described elsewhere.19 A compilation of primer sequences used in HRM is available on request. Samples with unusual melting profiles were sequenced for confirmation of genetic variation applying the same primer sets as used in the HRM analyses.
Databases and statistical analyses
Statistical analyses including estimation of significant differences in genotype frequencies between cases and controls, as well as of deviations from the Hardy–Weinberg equilibrium (HWE) were performed with STATA v10.2 (STATA Corporation, College Station, TX, USA) and PLINK v1.07 (http://pngu.mgh.harvard.edu/~purcell/plink/). Significance was assessed for single variants by logistic regression adjusted for age, gender and ethnic groups. Haplotype-specific risks were analysed with the UNPHASED v3.1.6 software (https://sites.google.com/site/fdudbridge/software/unphased-3-1) for an entire gene when <11 SNPs were genotyped. Otherwise, a sliding window approach was carried out analysing 10 SNPs simultaneously.
We also applied methods for the joint analysis of rare variants obtained by HRM of IFNGR1. We calculated fixed-threshold tests (T1 and T5) and a variable-threshold test. In addition, we carried out the SNP-set kernel association test (SKAT) and the estimated regression coefficient test (EREC), which allow to assess associations between cases and controls with SNPs, indicating opposite directions of genetic effects. All tests were computed with the SCORE-Seq software (http://dlin.web.unc.edu/software/score-seq/). Power calculation was conducted with the CaTS power calculator software.
Results
Re-sequencing and association results of single variants
We selected 223 variants identified by re-sequencing of the 20 pathway genes and additional 23 non-synonymous variants by the HRM screen of IFNGR1 for genotyping in the entire study group on the basis of the defined criteria. Single-variant analyses gave the strongest nominal association results with two SNPs of STAT1, rs16824035 and rs7562024 (OR=1.19, Padd=0.002 and OR=0.87, Padd=0.004, respectively), and one SNP of IFNGR1 rs7749390 (OR=0.88, Padd=0.008) applying an additive model (Table 1). However, after correction for multiple testing none of the findings remained statistically significant.
Haplotype analysis, rare variant analysis and IFNGR1 HRM analysis
For haplotype analyses, either all variants or only non-synonymous variants of each gene were selected for re-construction. No statistically significant differences could be observed. Haplotype analysis was restricted to variants with minor allele frequencies >1%. When comparing the frequencies between cases and controls, no significant association was observed for any of the 20 genes (data not shown).
In addition, we jointly analysed all rare non-synonymous variants with an MAF <5% of the IFNGR1 gene identified by HRM analysis, applying tests assuming the same direction of effect for each SNP, and the SKAT and EREC tests that allow to assess associations with rare variants even with different effect directions for each SNP. However, no significant result with all non-synonymous variants of IFNGR1 was observed (data not shown). This also applied to the subgroup of non-synonymous variants with a predicted structural impact on the protein structure assessed by a PolyPhen-2 (http://genetics.bwh.harvard.edu/pph2/) score of >0.7. Single-variant analyses of IFNGR1 SNPs did yield significant associations as well (Table 2).
Power calculation and HWE
With the sample size analysed, we achieved >90% power for multiplicative and additive models, assuming an TB prevalence of 0.004 in West Africa, a frequency of 0.1 for high-risk alleles and a genotype-relative risk of 1.3 (α=0.05). We excluded 18 variants not in HWE among control groups applying a threshold of P=0.01.
Discussion
We have systematically screened 20 major genes of the IFN-γ signalling pathway by re-sequencing and HRM analyses in more than 4500 individuals of a Ghanaian TB case–control study. The genes selected represent a plausible set of constituents of the IFN-γ-dependent immune response and include most of those genes of which previous studies had reported positive associations. Although sufficient power to detect significant differences in the genotype frequencies of SNPs of IFN-γ pathway genes was achieved and our analysis covered all non-synonymous variants of the genes with minor allele frequencies >2%, no statistically significant signals could be observed after correction for multiple testing. This was also the case when we compared the haplotype frequencies and rare variants of each gene between cases and controls. Although we screened the entire study group for rare exonic variants of the IFNGR1 gene, none of the deleterious mutations linked to increased susceptibility to NTM in other study groups was identified.
The present study group was part of that recently applied in a GWAS.4, 5 Based on previous evidence of a role of the IFN-γ signalling pathway genes, we designed a protocol with a far higher SNP coverage of these pathway genes, including sequencing of promoters, exons and the 3′-UTR regions, and included haplotype tagging SNPs as well as HRM typing. As it has previously been shown that with the same study group significant results could be obtained in the GWAS, the current findings of the pathway analysis appear to be valid and credible.
Several of the previous studies analysing single components of IFN-γ pathway genes were performed with insufficient sample sizes. This might have led to spurious and non-reproducible findings, as illustrated by two recent meta-analyses exploring findings of distinct variants of the IL12 and IFNG. No consistent patterns of association were observed.9, 14
Studies of IFN-γ pathways genes in TB are largely motivated and driven by previous findings of deleterious mutations associated with an increased risk of NTM infections. The reasons why the majority of individuals with deleterious mutations were diagnosed with NTM and not with M. tuberculosis remain unclear. 20 Apart from the presumed explanation that infants carrying variants conferring primary immunodeficiency might die early from TB and, therefore, are not accessible to genetic studies, and the infrequent incidence of TB in these individuals might indicate that IFN-γ signalling does not have an equal relevance in M. tuberculosis infections as it has in those with weakly pathogenic mycobacteria. In fact, M. tuberculosis is known to be able to suppress signal transduction of IFN-γ in several ways and, as a result, may effectively inhibit the bactericidal effects of activated macrophages.21, 22, 23 Further indicators of IFN-γ effects in TB susceptibility came from studies of IFN-γ KO mice, which succumb soon after M. tuberculosis infection. It is known, however, that mouse models do not reflect appropriately the pathology of TB as observed in humans. Equivalents of TB granuloma in human lungs are not seen in the lungs of infected mice and mice ultimately die of an infection with M. tuberculosis, whereas only 5%–10% of humans develop active disease after infection.24 This indicates that the observations of the impact of IFN-γ signalling made in mice can not readily be assigned to human TB pathology.
Our results, namely the absence of associations of TB with genes of the IFN-γ signalling pathway, strongly suggest that genetic variation of these genes may be less important in immunity to M. tuberculosis, at least in our large Ghanaian study group. Although relying on the current comprehension of TB immunity, the problems to develop effective TB vaccines reflect an incomplete understanding of immunological processes in TB.25 Other, yet unknown pathways might be of more relevance in the pathophysiology and defence of the disease as preliminarily suggested by the GWASs undertaken in TB. The unbiased approach of GWASs has allowed the identification of two genomic regions associated with resistance and susceptibility. Regulatory elements in these regions might point to unrecognized metabolic pathway with considerable impact on the pathophysiology of TB.3
The present study has limitations. Only variants of the promoter, the exons, the 3′-UTR and tagging variants, as well as variants that have previously been found associated with TB, were included. Although intronic variation was partly covered by the haplotype tagging SNPs, variants of intronic regions as well as unknown regulatory variants located distant from the genes studied might have been missed. Nevertheless, with this analysis we have tested a substantial share of the coding region of IFN-γ pathway genes in a most vulnerable sub-Saharan population.
References
Cooper AM : Cell-mediated immune responses in tuberculosis. Annu Rev Immunol 2009; 27: 393–422.
Comstock GW : Tuberculosis in twins: a re-analysis of the Prophit survey. Am Rev Respir Dis 1978; 117: 621–624.
Meyer CG, Thye T : Host genetic studies in adult pulmonary tuberculosis. Semin Immunol 2014; 26: 445–453.
Thye T, Vannberg FO, Wong SH et al: Genome-wide association analyses identifies a susceptibility locus for tuberculosis on chromosome 18q11.2. Nat Genet 2010; 42: 739–741.
Thye T, Owusu-Dabo E, Vannberg FO et al: Common variants at 11p13 are associated with susceptibility to tuberculosis. Nat Genet 2012; 44: 257–259.
Cooper AM, Dalton DK, Stewart TA, Griffin JP, Russell DG, Orme IM : Disseminated tuberculosis in interferon gamma gene-disrupted mice. J Exp Med 1993; 178: 2243–2247.
van de Vosse E, Hoeve MA, Ottenhoff TH : Human genetics of intracellular infectious diseases: molecular and cellular immunity against mycobacteria and salmonellae. Lancet Infect Dis 2004; 4: 739–749.
Lazarevic V, Glimcher LH : T-bet in disease. Nat Immunol 2011; 12: 597–606.
Pine R : IRF and tuberculosis. J Interferon Cytokine Res 2002; 22: 15–25.
Krebs DL, Hilton DJ : SOCS proteins: negative regulators of cytokine signaling. Stem Cells 2001; 19: 378–387.
Kusuhara K, Yamamoto K, Okada K, Mizuno Y, Hara T : Association of IL12RB1 polymorphisms with susceptibility to and severity of tuberculosis in Japanese: a gene-based association analysis of 21 candidate genes. Int J Immunogenet 2007; 34: 35–44.
Sahiratmadja E, Baak-Pablo R, de Visser AW et al: Association of polymorphisms in IL-12/IFN-gamma pathway genes with susceptibility to pulmonary tuberculosis in Indonesia. Tuberculosis (Edinb) 2007; 87: 303–311.
Moller M, Nebel A, van Helden PD, Schreiber S, Hoal EG : Analysis of eight genes modulating interferon gamma and human genetic susceptibility to tuberculosis: a case-control association study. BMC Infect Dis 2010; 10: 154.
de Albuquerque AC, Rocha LQ, de Morais Batista AH et al: Association of polymorphism +874A/T of interferon-gamma and susceptibility to the development of tuberculosis: meta-analysis. Eur J Clin Microbiol Infect Dis 2012; 31: 2887–2895.
Xu Z, Pei L, Zhang F et al: A functional variant in IL12B promoter modulates its expression and associates with increased risk of allergic asthma. Genes Immun 2013; 14: 238–243.
Sabri A, Grant AV, Cosker K et al: Association study of genes controlling IL-12-dependent IFN-gamma immunity: STAT4 alleles increase risk of pulmonary tuberculosis in Morocco. J Infect Dis 2014; 210: 611–618.
Thye T, Browne EN, Chinbuah MA et al: No associations of human pulmonary tuberculosis with Sp110 variants. J Med Genet 2006; 43: e32.
Sologuren I, Boisson-Dupuis S, Pestano J et al: Partial recessive IFN-gammaR1 deficiency: genetic, immunological and clinical features of 14 patients from 11 kindreds. Hum Mol Genet 2011; 20: 1509–1523.
Thye T, Browne EN, Chinbuah MA et al: IL10 haplotype associated with tuberculin skin test response but not with pulmonary TB. PLoS One 2009; 4: e5420.
Nunes-Alves C, Booty MG, Carpenter SM, Jayaraman P, Rothchild AC, Behar SM : In search of a new paradigm for protective immunity to TB. Nat Rev Microbiol 2014; 12: 289–299.
Fortune SM, Solache A, Jaeger A et al: Mycobacterium tuberculosis inhibits macrophage responses to IFN-gamma through myeloid differentiation factor 88-dependent and -independent mechanisms. J Immunol 2004; 172: 6272–6280.
Kincaid EZ, Ernst JD : Mycobacterium tuberculosis exerts gene-selective inhibition of transcriptional responses to IFN-gamma without inhibiting STAT1 function. J Immunol 2003; 171: 2042–2049.
Ting LM, Kim AC, Cattamanchi A, Ernst JD : Mycobacterium tuberculosis inhibits IFN-gamma transcriptional responses without inhibiting activation of STAT1. J Immunol 1999; 163: 3898–3906.
O'Toole R : Experimental models used to study human tuberculosis. Adv Appl Microbiol 2010; 71: 75–89.
Orme IM : Vaccine development for tuberculosis: current progress. Drugs 2013; 73: 1015–1024.
Acknowledgements
The participation of patients and the volunteers who served as controls is gratefully acknowledged, also the contributions of field workers, nurses and physicians involved in the recruitment of participants, the staff of the Kumasi Centre for Collaborative Research in Tropical Medicine (KCCR) and the excellent assistance of Emmanuel Abbeyquaye, Lincoln Gankpala, Birgit Muntau, Christa Ehmen, Gerd Ruge and Jürgen Sievertsen.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Meyer, C., Intemann, C., Förster, B. et al. No significant impact of IFN-γ pathway gene variants on tuberculosis susceptibility in a West African population. Eur J Hum Genet 24, 748–755 (2016). https://doi.org/10.1038/ejhg.2015.172
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2015.172