Abstract
Whole exome/genome sequencing (WES/WGS) is increasingly offered to ostensibly healthy individuals. Understanding the motivations and concerns of research participants seeking out personal WGS and their preferences regarding return-of-results and data sharing will help optimize protocols for WES/WGS. Baseline interviews including both qualitative and quantitative components were conducted with research participants (n=35) in the HealthSeq project, a longitudinal cohort study of individuals receiving personal WGS results. Data sharing preferences were recorded during informed consent. In the qualitative interview component, the dominant motivations that emerged were obtaining personal disease risk information, satisfying curiosity, contributing to research, self-exploration and interest in ancestry, and the dominant concern was the potential psychological impact of the results. In the quantitative component, 57% endorsed concerns about privacy. Most wanted to receive all personal WGS results (94%) and their raw data (89%); a third (37%) consented to having their data shared to the Database of Genotypes and Phenotypes (dbGaP). Early adopters of personal WGS in the HealthSeq project express a variety of health- and non-health-related motivations. Almost all want all available findings, while also expressing concerns about the psychological impact and privacy of their results.
Similar content being viewed by others
Introduction
Whole exome and genome sequencing (WES/WGS) are increasingly used in both research and clinical settings.1, 2, 3, 4, 5, 6, 7, 8 WGS offers the potential to implement a single genetic test that can capture much of the known genetic variation associated with rare monogenic and common polygenic disease, and medication safety and efficacy, along with non-health-related information, for example, ancestry. WGS is being applied in both disease-specific and general genomic medicine scenarios.4 In the latter, recipients are ostensibly healthy, the immediate goals include screening for disease risk rather than diagnosis, and potential benefits include empowering patients to take a more informed, personalized and preventative approach to their health and wellness.9, 10
The attitudes of research participants considering receiving personal WGS results will provide valuable data to inform the development of much needed standards and procedures for WGS in a general genomic medicine context.6, 11 An improved understanding of participants’ motivations, expectations, concerns, preferences regarding return of results, and choices regarding how their data is stored, shared and used will inform provider communication strategies and the design of the consent process and associated educational materials.
Multiple studies have examined baseline attitudes of early adopters of direct-to-consumer genotyping tests.12, 13, 14 Participants’ motivations and concerns were examined in the ClinSeq project, a cardiac-focused WES study at the NIH,15, 16, 17, 18 and are being assessed among participants in the WGS-based MedSeq project at Harvard.4 However, none of these studies focus on the scenario we expect to be increasingly common: health- and non-health-related WGS results offered to individuals outside their usual health-care providers. In addition, although multiple studies have examined individuals’ desire for personal genomic results17, 19, 20 and attitudes towards data sharing,21, 22, 23, 24 to date, little is known about the preferences or choices that are made among research participants actually considering receiving personal WGS results in a general genomic medicine context.
In this report, we assess the motivations and concerns about personal genome sequencing of research participants seeking out personal WGS, using both quantitative and qualitative methods. Our secondary aims are to describe participants’ preferences regarding return-of-results, and to report their choices in the informed consent document regarding data sharing. We utilize baseline data from the HealthSeq project, a pilot longitudinal cohort study to assess the psychosocial and health-related outcomes among an unselected population of ostensibly healthy individuals offered health- and non-health-related WGS results.
Methods
Study design
This was a mixed-methods longitudinal cohort study in which participants (n=35) had their genomes sequenced and received a variety of genetic results. Participants completed questionnaires and in-depth interviews at multiple timepoints throughout the study (see Supplementary Figure 1).
Participant recruitment and study procedures
Participants were recruited from the general population at the Mount Sinai Medical Center in New York City using recruitment flyers posted in common areas and via word-of-mouth (rather than from specific clinics or disease communities). The study was open to interested adults aged 18 years or older and who were English speaking. Staff and family members of the Icahn School of Medicine at Mount Sinai (ISMMS) Genetics and Genomics Department, students who had taken the ISMMS Practical Analysis of Your Personal Genome course, individuals who reported currently undergoing a genetics evaluation and/or whose motivation for participation was to diagnose a suspected genetic disorder via WGS, and women who reported being pregnant at the time of enrolment were not eligible to participate. In order to recruit individuals from a diverse educational background, the number of participants working or training in health care and the sciences was capped at 20. Financial compensation was not provided. The ISMMS Institutional Review Board approved this study.
Interested participants were informed that they had the option to receive their personal results from WGS and that they would be able to choose the types of information they wanted to learn. They were also given the option to receive their raw sequencing data. Participation involved attending two in-person appointments and completing a total of five interviews and six questionnaires (see Supplementary Figure 1). The initial appointment included informed consent, T1-pre interview and questionnaire, genetic counselling, blood draw, and T1-post questionnaire. During informed consent and genetic counselling, participants were informed that the research report would include the following categories of results, sequencing quality, ancestry, physical traits, pharmacogenomics, common polygenic disease risk including Alzheimer’s disease and monogenic disease variants, and were then asked to consider what types of information they would like to know. The focus of this report is the T1-pre interview and questionnaire.
Qualitative topic guide content
The qualitative topic guide for the T1-pre interview was designed to explore participants’ baseline motivations for participating in HealthSeq and interest in personal WGS results, expectations regarding receiving WGS results, concerns surrounding receiving results, and baseline understanding of WGS, using questions and probes. See Supplementary Materials for the questionnaire, including the short qualitative topic guide.
Quantitative measures
Supplementary Table 1 and Supplementary Table 2 present a summary of the quantitative and qualitative measures. In brief, the T1-pre questionnaire measures included: socio-demographic characteristics, motivations, concerns and intentions regarding personal WGS results. In addition, choices regarding WGS data storage and data sharing were recorded in the informed consent document (included in the Supplementary Materials) and procedure, and were also included in the present report. See Supplementary Material for further details of the measures used.
Qualitative analyses
The baseline T1-pre interviews assessing participants’ motivations and concerns were audio-recorded and transcribed verbatim. Only the transcript sections that were relevant to the primary aims of this study, that is, the motivations, expectations and concerns about personal genome sequencing, were coded. Although we attempted to analyze expectations, we found that with the possible exception of those with higher levels of familiarity with genomics research, participants struggled to answer the questions on this topic, and in most cases, answered by discussing their motivations rather than expectations. Therefore, we did not include participants’ expectations in our final analysis, and do not discuss it further here. The interview data were analyzed using thematic analysis.25 In brief, transcripts were read and an initial set of codes were developed. Data were then coded and organized according to content into categories by two investigators. New categories were created until a repeating category was apparent. Differences between the two investigators were discussed and reconciled, and codes were revised. Themes and sub-themes were collaboratively identified from the coded categories. Frequencies were calculated to better describe and understand the relationships between the themes and sub-themes.
Quantitative analyses
We described the socio-demographics of the sample using frequencies, means and standard deviations (SDs). We described the 10 individual motivations and 6 individual concerns items using frequencies. A composite ‘overall concerns’ score was calculated by creating a new dichotomous variable indicating whether or not each participant had endorsed at least one of the concerns. Intentions regarding personal results and informed consent choices regarding data-sharing were described using frequencies. All statistical analyses were conducted using statistical software package SPSS v. 20 (IBM).
Results
Socio-demographic characteristics of the participants
As Table 1 shows, 46% of the participants were female; mean age was 48 years; 71% were White non-Hispanic; 31% had a PhD/MD/JD; 83% were employed full-time; and half had an annual household income of less than $150 000.
Qualitative results for motivations and concerns
Motivations (qualitative)
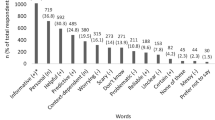
As Table 2 shows, eight overarching motivation themes emerged in the qualitative portions of the T1-pre interviews: (1) To learn personal disease risk or health-related information; (2) Curiosity; (3) To contribute to research; (4) Interested in their ancestry; (5) To provide disease risk information for their children and other family members; (6) Self-exploration; (7) Novel opportunity; and (8) Useful professionally.
To learn personal disease risk or health-related information. The first overarching theme, to learn personal disease risk or health-related information, was reported by the majority of participants. Participants primarily talked about wanting this information so they could avoid or reduce their risk of disease. Of these, almost half discussed how having this information might motivate changes to their lifestyle, diet and/or exercise to reduce their risk. Some also said the information could allow for reduced disease risk via medical intervention.
A quarter of participants discussed wanting personal disease risk information so they could prepare or plan for the future, even if they could not do anything to treat or prevent the disease. Participants talked about this in terms of both wanting to be able to emotionally prepare, and/or to be able to financially plan, for the future. When discussing their beliefs about the value of having their genomes sequenced, some participants said they believed their stored personal information could be of potential value to them in the future. In most of these cases, participants talked about how stored personal information may be useful in the future for medication prescribing purposes (ie, pharmacogenomics). One participant speculated that stored personal information might be useful in the future for diagnostic purposes. Other participants’ responses reflected a belief that stored personal information could be useful to them in the future as the science advances, for example, ‘I think that there’s potential future medical benefit for myself as medical and genetic information advances, so I believe that’ll be the main benefit’ [Participant 15].
A third of participants said they were interested in having their genomes sequenced because of a known family history of disease. Fewer said they were interested in having their genomes sequenced because of a known or suspected personal history of a disease such as heart disease or cancer. A few said they were motivated because of a lack of information about family history.
Curiosity. Curiosity was the second major overarching theme. Of the two-thirds who expressed curiosity as a motivator, some said they were motivated by a general curiosity, for example, ‘I’m a completely curious person, I always have been. I like knowing more rather than less’ [Participant 8]. Some said they were motivated to have their genomes sequenced because they were interested in or curious about the science. Within this theme, most stated that it was specifically the field of genetics/genomics that interested them, for example, ‘It’s just fricking cool, man… I’m intrigued by genetics’ [Participant 32]. Two participants were interested in science in general and, for one participant, it was research more broadly. Several participants said that they were motivated by a general curiosity about their genes, for example, ‘Just, I don’t know, getting some insight into genetic makeup I think would be interesting’ [Participant 25].
To contribute to research. Fifteen participants said they were motivated by a desire to contribute to research. This was almost always mentioned at the same time as motivations relating to personal gain, for example, ‘I have two main motivations. One is to learn about myself and my ancestry and any potential medical conditions that could arise in my future life, and the other one, which is just as equally important, is to contribute to scientific knowledge.’ [Participant 14]. Some participants framed their desire to contribute to research in terms of an altruistic desire to help others and the field of medicine, while others framed it as a desire to contribute to the advancement of science.
Interested in their ancestry. Thirteen participants said one of their motivations was an interest in their ancestry, for example, ‘I’m really interested in the ancestry thing. I mean, I really am so I look forward to that piece of it’ [Participant 27].
To provide disease risk information for their children and other family members. Eleven participants said they were motivated by a desire to provide disease risk information for their children and other family members. In most of these cases, participants were motivated by getting information for their existing children, and some of these mentioned that the information would also be useful for their family members generally. A few participants said the information might be useful for their future potential children and family planning.
Self-exploration. A novel theme that came up was self-exploration: seven participants said they were motivated by a general desire to learn more about themselves, for example, ‘I thought this was like the ultimate exploration of me’ [Participant 11].
Novel opportunity. A few participants said they wanted to get their genomes sequenced because it was a novel opportunity, for example, ‘Well, not many people have done this… Very few people have. I feel kinda special. It’s a great conversation piece (laughter)’ [Participant 28].
Useful professionally. One participant who was a medical student stated that part of their motivation to have their genome sequenced was because they thought it would be useful professionally. See Supplementary Table 3 for additional quotes.
Concerns (qualitative)
Over half of participants stated they were not concerned about having their genomes sequenced. Among participants who did have concerns, three overarching themes emerged: (1) Concern about adverse psychological impact of potential results; (2) Concern about implications of potential results for children; and (3) Concern about loss of privacy (see Table 2).
Concern about the adverse psychological impact of potential results. A quarter of participants mentioned they were concerned about the adverse psychological impact of potential results. Of these, several stated they had concern about results regarding untreatable diseases or conditions. In two cases, this was non-specific; in three cases, it was Alzheimer’s disease; and in one case, it was Huntington’s disease (see Supplementary Table 3). A few participants expressed a fear of the unknown, for example, ‘Knowing that you can’t know the results beforehand’ [Participant 8]. One said that they were concerned about breast cancer (BRCA1/2) because of recent media coverage. In most cases, participants said that although they had concerns about the potential psychological impact, they would nonetheless want to know, for example, ‘I just think the benefits of knowing outweigh the risks of being fearful’ [Participant 30].
Concern about implications of potential results for children. The second overarching theme was concern about implications of potential results for children, which emerged in four interviews. Most of these participants had concerns about implications for existing children, for example, ‘…probably mostly about my son. Just knowing his potential to developing a disease...’ [Participant 26]. One participant was concerned about implications for future reproductive decision-making: ‘Only that I would have a rare mutation that I for sure will pass on to my child a hundred percent, and that that would make me not wanna have children.’ [Participant 37].
Concern about loss of privacy. The third theme, raised by three participants, was concern about loss of privacy, for example, ‘I think my only concern would be if somewhere down the line someone got my sequencing data, did some weird analysis on it and published it and it ended up in a newspaper or something, or on CNN. I wouldn’t want that.’ [Participant 9]. See Supplementary Table 3 for additional quotes.
Quantitative results for motivations and concerns
Motivations (quantitative)
In the quantitative, closed-ended questions about why participants were motivated to get their genomes sequenced, 24 (69%) rated ‘Find out what I can do to improve my health,’ and 22 (60%) rated ‘Find out about diseases for which I am at risk,’ as very important motivations to them. Twenty-five (71%) participants rated ‘Curiosity about my genes’ as very important. ‘Participate in research to help others’ was rated as very important by 49% of participants, while 40% rated ‘Interested in my ancestry’ as very important, and 34% rated ‘Want information about risk of health conditions for children and grandchildren’ as very important. See Table 3 for all responses. See Supplementary Table 4 for exploratory factor analysis results.
Concerns (quantitative)
As Table 3 shows, 66% of participants had at least one concern about participating in the study and receiving their personal results from WGS. Of these, 20 (57%) participants endorsed concerns related to potential privacy issues about their data. Six of the 10 participants who provided additional detail regarding their privacy concerns cited concerns about insurance (see Table 3, footnote 4). Ten (29%) endorsed ‘Concerns related to not knowing how I will feel about my results,’ and eight (23%) endorsed ‘Concerns related to learning about my disease risk from whole genome sequencing.’
Intentions to receive personal WGS results
As Table 4 shows, 33 (94%) participants reported that they would like to receive all personal WGS results available. All except one of the participants endorsed ‘yes probably / yes definitely’ for all nine of the specific results categories they were asked about. There was some variation in the proportion who endorsed ‘yes definitely,’ however, with 100% participants endorsing ‘yes definitely’ for pharmacogenetics, compared with 77% who endorsed ‘yes definitely’ for a diagnosis of a rare genetic disorder. Seventy-four percent reported that they would want to know their genetic risk for a disease that they could not do anything to prevent.
Participants’ choices regarding WGS data, specimen storage and data sharing
Most participants (32/35, 91.4%), indicated that they wanted their WGS data stored in a way that was linked to their identity. All 32 participants who gave permission for their WGS data to be stored also gave permission for their data to be used by Mount Sinai investigators in future research studies directly related to the present research. However, only 21 (60.0%) gave permission for their data to be used by investigators at other institutions even if the purposes were directly related to the present research, and the same proportion gave permission for investigators at Mount Sinai to use their data for purposes not directly related to the present research. Most participants (31/35, 88.6%) said that they would like to receive their raw WGS data. Thirteen (37.1%) participants gave consent for their WGS data to be shared with the Database of Genotypes and Phenotypes (dbGaP). Supplementary Table 5 also shows participants’ informed consent choices regarding storage and sharing of their blood specimens.
Discussion
In this study, we found that the most common motivations for ostensibly healthy people to seek personal genome sequencing were curiosity and a desire to obtain personal health-related information. These findings are consistent with previous survey research in which individuals who obtained genotyping were motivated by their own curiosity and to find out their disease risk to improve their health.13, 14 Similarly, in the ClinSeq project a major motivation for participants seeking to obtain WES was a desire to learn more about genetic factors that contribute to one’s own health risk.15
A similar proportion of HealthSeq and ClinSeq15 participants reported being motivated by a desire to contribute to research, 43% and 44%, respectively (both assessed qualitatively). However, ClinSeq participants were either motivated by a desire to obtain personal genomic information about themselves or by a desire to contribute to research, but not by both.15 In contrast, these motivations overlapped in HealthSeq participants. This could be due to contextual differences: return-of-results is integral to HealthSeq but only a secondary goal in the ClinSeq project.
We found that interest in ancestry was another important motivating factor for pursuing personal genome sequencing: 37% of participants spontaneously mentioned this in the qualitative interview and 40% endorsed this as a ‘very important’ reason on the questionnaire. The Coriell Personalized Medicine Collaborative (CPMC) and ClinSeq studies do not include non-health-related information and so do not capture this motivation. The public interest in ancestry information, as demonstrated in our cohort and by the 100 000s of customers who have purchased direct-to-consumer genetic ancestry services,26, 27 could provide a useful mechanism for encouraging people to also seek out and obtain useful personal health-related genomic information. The inclusion of ancestry information could perhaps broaden public interest in and acceptance of WGS beyond the early adopters.
Six of the 10 participants who provided additional details about their privacy concerns in the questionnaire specifically cited insurance-related concerns. This was the most common type of privacy concern reported. We hypothesize that insurance discrimination is a common concern among potential genomic research participants and that concerns about insurance may discourage participation in genomic research studies (as is suggested in the preliminary results from the MedSeq study in which 25% of prospective participants who declined to continue in the study cited a fear of insurance discrimination28). Incorporating additional questions to better elucidate participants’ specific privacy concerns would enhance future study instruments.
We observed differences in reported privacy concerns between the qualitative (9% of participants) and quantitative methodologies (57%). This difference could reflect different prompting.29 The qualitative interview is essentially ‘unprompted’, that is, people are asked to generate and therefore ‘recall’ their own concerns. The quantitative survey is a ‘prompted’ approach, that is, people are asked to respond to and therefore ‘recognize’ a list of concerns. The latter has led to higher rates of endorsement in other health-related contexts.29 It is also possible that the protocol contributed to this difference: the qualitative interviews were conducted immediately after participants had been taken through the consent form, which included a significant emphasis on privacy and confidentiality, and so they may have felt that their privacy concerns had been aired and did not need to be raised in the interview. In contrast, when completing the questionnaire, participants may have felt that endorsing privacy concerns was reasonable given the concerns they had felt coming into the study.
Very few participants mentioned privacy concerns during the interview, but approximately a quarter of participants expressed concerns about the psychological impact of WGS findings. This may be because how personal data will be protected and shared is to a large extent ‘knowable,’ controllable and predictable; in contrast, it is impossible to precisely predict exactly what types of personal results will be obtained from WGS and how distressing any given individual may find those results. The protocol order may also have been an influencing factor: participants completed the T1-pre interview after informed consent, which had a greater emphasis on privacy, but before the genetic counselling session, which had a greater emphasis on psychosocial considerations. Given these considerations, it is perhaps not surprising that more participants discussed their concerns regarding the potential psychological impact of results rather than regarding privacy.
Regardless of these concerns, the majority of participants indicated that they wanted to receive all or most of the personal results arising from WGS. In contrast, only a third consented for their genome data to be shared with dbGaP. This highlights an important difference between our population and participants in the Personal Genome Project (PGP),1, 8 in which open access to and sharing of genomic data is integral to the study and participants enter the study with that knowledge. Research participants’ views on data sharing are influenced by multiple interrelated factors, including risks, such as privacy, benefits, such as helping others, trust and perceptions of sensitivity and controllability of genomic data.24 Our measures were not designed to specifically assess data-sharing preferences, however, the trend of more sharing under more restricted conditions, for example, within ISMMS and for related projects only, is consistent with factors reported to influence participants’ views on sharing. The more restrictive conditions increase participant control, keep the data within the known and presumably trusted institution, and reduce the chances for misuse. Given the importance of data sharing to advancing genomics research, an in-depth study of the attitudes of HealthSeq participants towards data sharing is important follow-on work.
A limitation of this study was the small sample size. In addition, although we attempted to recruit individuals from a range of socio-demographic backgrounds, the participants were predominantly White non-Hispanic with high incomes and high levels of educational attainment. Thus, the findings may not generalize to the general population. The makeup of the HealthSeq participants may partially reflect the characteristics of early adopters who are interested in obtaining and using new genomic technologies for personal use. It may also reflect issues of access. For instance, we considered advertising the study in a community-based clinic, but after consultation with the clinic staff ultimately did not do so because of concerns about the access to and potential costs of follow-up care. In order for WES/WGS not to exacerbate existing health disparities, the challenging issue of access to follow-up care as well as to the technology itself will need to be actively addressed.
A strength was the mixed-methods approach that enabled us to compare our quantitative survey results with previous research,13 and to generate a list of motivations and concerns that participants expressed unprompted in qualitative interviews. This list, which includes the novel motivations ‘self-exploration’ and ‘novel opportunity’, will be a valuable resource for developing measures for use in future studies of personal genome sequencing.
In conclusion, desire for personal genomic information does not automatically overlap with a desire to share that information publicly. Early adopters of personal genome sequencing are unsurprisingly enthusiastic but nonetheless have concerns about how the results will affect them emotionally, and about protecting their privacy, and may not wish to share their data in dbGaP let alone more widely.
References
Church GM : The personal genome project. Mol Syst Biol 2005; 1: 2005.
Ashley EA, Butte AJ, Wheeler MT et al: Clinical assessment incorporating a personal genome. Lancet 2010; 375: 1525–1535.
Gonzalez-Garay ML, McGuire AL, Pereira S, Caskey CT : Personalized genomic disease risk of volunteers. Proc Natl Acad Sci USA 2013; 110: 16957–16962.
Vassy JL, Lautenbach DM, McLaughlin HM et al: The MedSeq Project: a randomized trial of integrating whole genome sequencing into clinical medicine. Trials 2014; 15: 1–12.
Yang Y, Muzny DM, Xia F et al: Molecular findings among patients referred for clinical whole-exome sequencing. JAMA 2014; 312: 1870–1879.
Jacob HJ, Abrams K, Bick DP et al: Genomics in clinical practice: lessons from the front lines. Sci Transl Med 2013; 5: 194cm5.
Biesecker LG, Green RC : Diagnostic clinical genome and exome sequencing. N Engl J Med 2014; 371: 1170.
Angrist M : Eyes wide open: the personal genome project, citizen science and veracity in informed consent. Per Med 2006; 6: 691–699.
Goldberger JJ, Buxton AE : Personalized medicine vs guideline-based medicine. JAMA 2013; 309: 2559–2560.
Sanderson SC : Genome sequencing for healthy individuals. Trends Genet 2013; 29: 556–558.
Caulfield T, Evans J, McGuire A et al: Reflections on the cost of ‘low-cost’ whole genome sequencing: framing the health policy debate. PLoS Biol 2013; 11: e1001699.
Bloss CS, Ornowski L, Silver E, Cargill M et al: Consumer perceptions of direct-to-consumer personalized genomic risk assessments. Genet Med 2010; 12: 556–566.
Gollust SE, Gordon ES, Zayac C et al: Motivations and perceptions of early adopters of personalized genomics: perspectives from research participants. Public Health Genomics 2012; 15: 22–30.
Vayena E, Gourna E, Streuli J, Hafen E, Prainsack B : Experiences of early users of direct-to-consumer genomics in Switzerland: an exploratory study. Public Health Genomics 2012; 15: 352–362.
Facio FM, Brooks S, Loewenstein J, Green S, Biesecker LG, Biesecker BB : Motivators for participation in a whole-genome sequencing study: implications for translational genomics research. Eur J Hum Genet 2011; 19: 1213–1217.
Facio FM, Sapp JC, Linn A, Biesecker LG : Approaches to informed consent for hypothesis-testing and hypothesis-generating clinical genomics research. BMC Med Genomics 2012; 5: 45.
Facio FM, Eidem H, Fisher T et al: Intentions to receive individual results from whole-genome sequencing among participants in the ClinSeq study. Eur J Hum Genet 2013; 21: 261–265.
Jamal L, Sapp JC, Lewis K et al: Research participants’ attitudes towards the confidentiality of genomic sequence information. Eur J Hum Genet 2014; 22: 964–968.
Kaufman D, Murphy J, Scott J, Hudson K : Subjects matter: a survey of public opinions about a large genetic cohort study. Genet Med 2008; 10: 831–839.
Bollinger JM, Scott J, Dvoskin R, Kaufman D : Public preferences regarding the return of individual genetic research results: findings from a qualitative focus group study. Genet Med 2012; 14: 451–457.
Trinidad SB, Fullerton SM, Bares JM, Jarvik GP, Larson EB, Burke W : Genomic research and wide data sharing: views of prospective participants. Genet Med 2010; 12: 486–495.
Haga SB, O’Daniel J : Public perspectives regarding data-sharing practices in genomics research. Public Health Genomics 2011; 14: 319–324.
Oliver JM, Slashinski MJ, Wang T, Kelly PA, Hilsenbeck SG, McGuire AL : Balancing the risks and benefits of genomic data sharing: genome research participants’ perspectives. Public Health Genomics 2012; 15: 106–114.
Shabani M, Bezuidenhout L, Borry P : Attitudes of research participants and the general public towards genomic data sharing: a systematic literature review. Expert Rev Mol Diagn 2014; 14: 1053–1065.
Braun V, Clarke V : Using thematic analysis in psychology. Qual Res Psychol 2006; 3: 77–101.
Su Y, Howard HC, Borry P : Users’ motivations to purchase direct-to-consumer genome-wide testing: an exploratory study of personal stories. J Community Genet 2011; 2: 135–146.
Wagner JK, Cooper JD, Sterling R, Royal CD : Tilting at windmills no longer: a data-driven discussion of DTC DNA ancestry tests. Genet Med 2012; 14: 586–593.
Green RC, Lautenbach D, McGuire AL : GINA, genetic discrimination, and genomic medicine. N Engl J Med 2015; 372: 397–399.
Waller J, McCaffery K, Wardle J : Measuring cancer knowledge: comparing prompted and unprompted recall. Br J Psychol 2004; 95: 219–234.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on European Journal of Human Genetics website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Sanderson, S., Linderman, M., Suckiel, S. et al. Motivations, concerns and preferences of personal genome sequencing research participants: Baseline findings from the HealthSeq project. Eur J Hum Genet 24, 14–20 (2016). https://doi.org/10.1038/ejhg.2015.118
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2015.118
This article is cited by
-
Balancing the safeguarding of privacy and data sharing: perceptions of genomic professionals on patient genomic data ownership in Australia
European Journal of Human Genetics (2023)
-
Willingness to participate in various nontherapeutic cancer research activities among urban and rural African American and Latinx healthy volunteers
Cancer Causes & Control (2022)
-
The return of individual genomic results to research participants: design and pilot study of Tohoku Medical Megabank Project
Journal of Human Genetics (2022)
-
Cascade health service use in family members following genetic testing in children: a scoping literature review
European Journal of Human Genetics (2021)
-
Parents, their children, whole exome sequencing and unsolicited findings: growing towards the child’s future autonomy
European Journal of Human Genetics (2021)