Abstract
Patients with terminal deletions of chromosome 6q present with structural brain abnormalities including agenesis of corpus callosum, hydrocephalus, periventricular nodular heterotopia, and cerebellar malformations. The 6q27 region harbors genes that are important for the normal development of brain and delineation of a critical deletion region for structural brain abnormalities may lead to a better genotype–phenotype correlation. We conducted a detailed clinical and molecular characterization of seven unrelated patients with deletions involving chromosome 6q27. All patients had structural brain abnormalities. Using array comparative genomic hybridization, we mapped the size, extent, and genomic content of these deletions. The smallest region of overlap spans 1.7 Mb and contains DLL1, THBS2, PHF10, and C6orf70 (ERMARD) that are plausible candidates for the causation of structural brain abnormalities. Our study reiterates the importance of 6q27 region in normal development of brain and helps identify putative genes in causation of structural brain anomalies.
Similar content being viewed by others
Introduction
Patients with terminal deletions involving chromosome 6q present with a variable phenotype that includes intellectual disability, hypotonia, epilepsy, cardiac defects, retinal abnormalities, ear anomalies, facial dysmorphisms, and malformations of the brain, spinal cord, and vertebrae. Structural brain malformations are consistently observed in these patients and include agenesis of the corpus callosum (ACC), periventricular nodular heterotopia (PNH), polymicrogyria, hydrocephalus, and cerebellar malformations.1, 2, 3, 4, 5, 6, 7, 8, 9, 10 Whereas previous studies have attempted to delineate the critical region responsible for brain malformations in patients with terminal 6q27 deletions, the sensitivity of the methodologies used has prevented the fine-mapping of the critical region.1, 6 A detailed genomic analysis of the subtelomeric chromosome 6q region would help identify the putative genes involved in the causation of brain malformations.
Here, we report the clinical and genomic characterization of seven unrelated patients with nonrecurrent 6q25.3-q27 deletions. These patients presented with various structural abnormalities of the brain including ACC, PNH, cortical atrophy, and hydrocephalus. We determined the size, extent, and genomic content of these deletions using high-resolution array comparative genome hybridization (aCGH), and deciphered a common region of overlap spanning 1.7 Mb. We hypothesize that a subset of genes in the commonly deleted region are dosage-sensitive and that the haploinsufficiency of one or more of these genes interferes with the normal development of nervous system during embryogenesis.
Materials and methods
Human subjects
The research protocol was approved by the Institutional Review Board of Baylor College of Medicine (BCM). Seven subjects with terminal 6q deletions were identified at the Medical Genetics Laboratories (MGL) of BCM by aCGH, fluorescent in situ hybridization (FISH), or high-resolution G-banded chromosome analysis. Clinical information was obtained from their health care provider. Informed consent was obtained from the families of five patients to publish photographs.
Patient 1
A 5-year-old female presented with a history of a repaired thoraco-lumbar meningomyelocele, Arnold Chiari II malformation, obstructive hydrocephalus treated with a ventriculo-peritoneal shunt, neurogenic bladder, vesicoureteral reflux, and epilepsy. Delivery at full term gestation via Cesarean section followed a reportedly uncomplicated pregnancy in a 33-year-old multigravida. Prenatal chromosome analysis had detected a 6q25-q27 deletion. In addition to confirming the hydrocephalus and Arnold Chiari II malformation, MRI of the brain revealed occipital polymicrogyria, dysplastic corpus callosum, malformed tectum, enlarged foramen magnum with herniation of the cerebellum, stenogyria of the occipital lobes, and paucity of supratentorial white matter. She had dysarthria, impaired memory, and learning problems and was enrolled in a preschool special education program. aCGH identified a 10.3 Mb heterozygous deletion at 6q25.3-q27 (Figure 1a). Parental testing could not be performed; however, the deletion is likely to be a de novo event as both parents and six of her siblings are all phenotypically normal. On follow up at the age of 7.5 years, she is entering 2nd grade in regular classes with some individualized instruction.
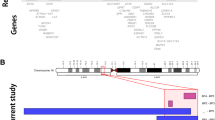
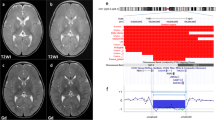
(a) Representative sections of aCGH plots from chromosomes 2–7 are shown for patients 1–7 along with genomic coordinates for the deletions on chromosome 6q. The red signal on the plot shows copy-number loss of 6q25.3-q27. (b) Representative partial karyogram showing a pair of G-banded chromosome 6 (patient 5) with deletion of 6q27. (c) A representative confirmatory metaphase FISH image (patient 5) showing 6q deletion in the distal end of one chromosome 6. The chromosomes are shown in DAPI (Blue), both chromosomes 6 show control signal (green) whereas only one shows the targeted subtelomeric region (red). The full colour version of this figure is available at European Journal of Human Genetics online.
Patient 2
A 42-year-old primigravida woman had an ultrasound at 21 weeks of gestation that showed a small cerebellum. Fetal MRI revealed mild asymmetry between the sizes of lateral ventricles along with a diffusely small cerebellum and cerebellar vermis (Figure 2a and b). G-banded chromosome analysis showed a large 6q25.3-qter deletion. Prenatal aCGH revealed a heterozygous deletion of∼9.3 Mb in 6q26-q27 (Figure 1a). Parental chromosome analysis was normal. The couple opted to terminate the pregnancy.
Representative MRI images showing various brain malformations in patients with 6q25.3-q27 deletions: Fetal MRI in patient 2 at 22 weeks of gestation shows asymmetry in size of lateral ventricles (a) and a diffusely small cerebellum (b). (c) Facial features of patient 5 with frontal bossing and broad nasal bridge. Brain MRI in patient 5 showing T1-weighted axial image with enlargement of the occipital lateral ventricles (colpocephaly) (d) and gray matter subependymal heterotopia lining the lateral ventricle (arrow) in (e). Sagittal midline T1-weighted image in Patient 7 demonstrating a foreshortened and dysplastic corpus callosum (f). Facial characteristics of patient 7 with low-set and posteriorly rotated ears, prominent nasal bridge, and hypertelorism (g).
Patient 3
A newborn female presented with low birth weight and multiple anomalies including: bicuspid aortic valve, imperforate anus with perineal fistula, and partial uterine septum. Additionally, there were segmentation anomalies of the sacrum, dextroconvex scoliosis of the lumbosacral spine, and tethered spinal cord. Craniofacial abnormalities included: macrocephaly, lacrimal duct stenosis, hypertelorism, broad nasal bridge, and thin upper lip. Following repair of the tethered cord at 4 months, she was followed for global developmental delay, hypotonia, and epilepsy. MRI of brain revealed right-sided subependymal nodular gray matter heterotopia and ACC. aCGH showed a 7.6 Mb heterozygous deletion at 6q26-q27 (Figure 1a). Her phenotypically normal parents did not harbor the deletion.
Patient 4
This female child born to a 39-year-old woman with hypertension and history of tobacco and alcohol use during pregnancy was noted to have intra-uterine growth restriction, microcephaly, and dysmorphic features including: prominent forehead, hypertelorism, epicanthal folds, low-set ears, smooth philtrum, micrognathia, and high-arched palate. She also had an atrial septal defect and partial anomalous pulmonary venous return. MRI of the brain detected prominent ventricles and sulci consistent with mild cerebral atrophy and cystic periventricular leukomalacia. When evaluated at the age of 6 years, she had seizures, significant developmental delay, hypotonia, learning difficulties, and behaviors consistent with autism spectrum disorder including flapping of arms, inability to make eye contact, and repetitive movements. The karyotype showed a heterozygous deletion at 6q27. aCGH revealed a 5.99 Mb heterozygous deletion in chromosome 6q27 (Figure 1a). The rearrangement was not detected in the mother whereas the father was unavailable for testing.
Patient 5
An 8-month-old female presented with congenital hydrocephalus and seizures. She was delivered at 36 weeks of gestation by Cesarean section in a 27-year-old female with gestational diabetes mellitus. Her physical examination showed macrocephaly, frontal bossing, and a broad nasal bridge (Figure 2c). The MRI of brain revealed hydrocephalus, dysmorphic ventricles, atrophy of the posterior part of corpus callosum, absent septum pellucidum, bilateral subependymal heterotopia within the lateral ventricles, atrophy of bilateral medial temporal lobes and hippocampi, and hypoplastic olfactory sulci (Figure 2d and e). Electroencephalogram revealed intermittent moderate-to-high voltage slow (delta) activity in bilateral occipital leads. A ventriculo-peritoneal shunt was placed for hydrocephalus. A peripheral blood karyotype showed a heterozygous deletion at 6q27. aCGH revealed a heterozygous deletion of 5.39 Mb within the 6q27 region (Figure 1a–c). The deletion was confirmed to be a de novo rearrangement.
Patient 6
A female infant born at 33 weeks of gestation presented with obstructive hydrocephalus and underwent placement of a ventriculo-peritoneal shunt. A computed tomography of brain revealed residual gross obstructive hydrocephalus, absence of septum pellucidum, and ACC. aCGH revealed a heterozygous deletion of 1.92 Mb within the 6q27 region (Figure 1a). Parental FISH analysis confirmed the deletion to be a de novo event.
Patient 7
A 30-month-old female presented with developmental delays and dysmorphic craniofacial features including: microcephaly, dolichocephaly, hypertelorism, prominent nasal bridge, and low-set posteriorly rotated ears (Figure 2g). The pregnancy had been complicated by maternal diabetes and HELLP syndrome. MRI of brain performed at 32 months of age showed a foreshortened and dysplastic corpus callosum. (Figure 2f). She had poor weight gain, behavioral issues, and had significant speech delay with a vocabulary of one word at 30 months of age. aCGH revealed a submicroscopic heterozygous deletion of 1.5 Mb within 6q27 region (Figure 1a). The rearrangement was not detected in the mother whereas the father was unavailable for testing.
A detailed comparison of clinical features for all the seven patients and the recently published reports of isolated 6q27 deletions are summarized in Table 1. In addition, a comparison of brain MRI findings in our cohort of patients is summarized in Supplementary Table 1.
aCGH analysis
DNA was extracted from whole blood using the Puregene DNA blood kit (Gentra, Minneapolis, MN, USA) according to the manufacturer’s instructions. aCGH was performed on all seven subjects using the clinical microarray platform routinely used at our institution. The microarrays were designed in the MGL, Houston, TX. The patient samples were interrogated using a custom-designed array with ∼180 000 interrogating oligonucleotides (60mer), manufactured by Agilent Technologies, Inc (Santa Clara, CA, USA). This array contains the ‘best-performing’ oligos selected from Agilent’s online library (eArray;https://earray.chem.agilent.com/earray/) and has been further optimized using empiric data. This array is designed to provide interrogation of all known microdeletion and microduplication syndrome regions, pericentromeric and subtelomeric regions, and computationally predicted NAHR-mediated genomic instability regions flanked by low-copy repeats (LCR) as previously described.11 In addition, ∼1800 known or candidate disease genes have exonic coverage (101 644 probes in 24 319 exons; average of 4.2 probes per exon). The entire genome is covered with an average resolution of 30 kb, excluding LCRs and other repetitive sequences. Further details regarding the design and coverage of the array are available at https://www.bcm.edu/geneticlabs/.12 The procedures for DNA digestion, labeling and hybridization, and data analysis, were performed as previously described.13
FISH analysis
Copy-number changes detected by microarray analysis were verified by FISH using BAC clone RP11-755M8 in patients 3–7 whereas partial karyotype was used to confirm the deletion in patients 1 and 2. The FISH analysis followed a standard clinical cytogenetics laboratory protocol. In addition, to confirm the mode of inheritance, parental samples from these patients were analyzed for 6q rearrangements using the same BAC clones.
Results
Clinical features
The summary of clinical features of 21 patients with variable size deletions between 6q25.3-q27 (seven patients from our cohort and 14 patients previously reported in the literature) are listed in Table 1. Among these patients, 14 were female, 7 were male. In the 12 families with known inheritance patterns, all the 6q deletions arose owing to de novo rearrangements. The clinical features observed in a majority of patients included structural brain abnormalities, learning difficulties, developmental delay, and hypotonia. Excepting a single patient reported by Mosca et al7, all had some form of structural brain abnormalities. Abnormal corpus callosum (62%) was the most common malformation followed by hydrocephalus (27%) and vertebral or spinal cord malformations (22%). There was no characteristic facial gestalt with thin upper lip and ears anomalies being the most common craniofacial dysmorphism.
Molecular cytogenetic analysis
Here, we report our cohort of seven patients with nonrecurrent 6q25.3-q27 deletions of variable sizes. The 6q genomic rearrangements in all the patients were identified by aCGH, chromosome analysis, or FISH (Figure 1a–c). The genomic characterization of the deletions revealed nonrecurrent rearrangements ranging in size from 1.7–10.8 Mb. The smallest region of overlap (SRO) in our cohort of patients spanned 1.7 Mb between the g.169,148,957_170,899,992 (GRCh37/Hg19 build). The SRO encompasses a number of protein-coding genes (THBS2, DLL1, WDR27, PHF10, TCTE3, C6orf70(ERMARD), C6orf120, FAM120B, PSMB1, TBP, and PDCD2), two long nonprotein coding RNA genes (LINC00242 and LINC00574), one miscellaneous RNA (LOC154449), and one microRNA (MIR45441). Amongst these genes THBS2, PHF10, and DLL1 are putative candidates for causation of structural brain malformations. A recent study also suggested a potential role for C6orf70 present within this SRO contributing to PNH, thus making this another candidate gene for brain malformations (Figure 3). Since LCRs or segmental duplications can stimulate nonrecurrent events,14 we performed in silico analysis of the deletion breakpoints in all our patients. Our analysis did not reveal segmental duplications near the deletion breakpoints in any of our patients.
Size, extent, and genomic content of deletions of 6q27, and mapping of critical region for the brain malformations. Upper panel depicts the ideogram of chromosome 6 with genomic coordinates; middle panel represents the deletions in our cohort (blue bars) and published literature (gray bars). Note that the scale has been expanded to depict the three smallest deletions (P7, P21, and P18). All patients except P11 had structural brain abnormalities. The SRO for brain anomalies in our cohort maps from THBS2 to PDCD2 (orange shade). THBS2, PHF10, DLL1, and C6orf70 are putative candidate genes for causation of structural brain malformations as underlined.
Discussion
Several reports of monosomy for the terminal portion of the chromosome 6q either owing to unbalanced translocations or isolated deletions have been previously described.1, 2, 3, 4, 5, 6, 7, 8, 9, 15, 16 Though the clinical phenotypes associated with this deletion are quite variable, reports in unrelated subjects from independent investigators have shown that ACC, PNH, polymicrogyria, hydrocephalus, and cerebellar malformations are the consistently observed features. A recent analysis of a comprehensive map of loci for ACC from 374 patients revealed that chromosome 6q27 was one of the few loci wherein six or more subjects with ACC have been reported.10 Additionally, polymicrogyria, cerebellar, and other unspecified structural brain abnormalities were also mapped to this genomic region. These data suggest that the subtelomeric region of chromosome 6q harbors genes that have a critical role in the normal development of the brain. Delineation of a deletion region that is critical for the brain abnormalities may help in the counseling of patients.
Using a reverse genomic approach, in the cohort of seven patients we identified a SRO of 1.7 Mb that contains plausible candidate genes, THBS2, PHF10, DLL1, and C6orf70 (Figure 3). These genes have important functions in morphogenesis of the nervous system during embryogenesis. Two amongst these candidate genes, DLL1 and THBS2 are important ligands for the activation of Notch-3 signaling cascade. The Notch signaling pathway is an evolutionarily conserved, intercellular signaling cascade that has an essential role in homeostatic function and development of multiple organ systems.17 The Notch signaling pathway is critical for neural proliferation, migration, and differentiation during embryonic development of the nervous system.18
DLL1 is a human homolog of the Notch Delta ligand and a member of the delta/serrate/jagged family. It is involved in various biological processes regulating cell adhesion, cell–cell communication and cell-fate determination. DLL1 is expressed in the paraxial mesoderm, which correlates with somitogenesis in the nervous system.19 The Notch ligand DLL1 is important for Notch receptor processing, regulation of neural stem cell proliferation, survival, self-renewal, and differentiation during embryogenesis.20 Homozygous deletion of Dll1 in murine embryos results in severe patterning defects in the para-axial mesoderm and the nervous system.21 Haploinsufficiency of DLL1 in mice results in a complex metabolic phenotype with reduced body weight, altered fat to lean ratio, and hypertension but not developmental anomalies of the brain.22 However, recently a 2.2 MB region on 6q27 between the g.168,690,655_170,899,992 was mapped to be a critical region for the causation of holoprosencephaly in humans.23 A three base-pair deletion in exon 9 of DLL1 was also observed in a subject with semilobar holoprosencephaly.23 In addition, a de novo insertion of a single base-pair altering the reading frame has been noted in a single patient with autism spectrum disorder. These data show that haploinsufficiency of DLL1 may lead to both structural and functional abnormalities of the brain.
THBS2 is a member of the thrombospondin family. It is a matricellular glycoprotein that has a role in bone growth, cell adhesion, extracellular matrix modeling, inflammatory responses, developmental, and pathological angiogenesis.24, 25 It is highly expressed in the cerebral cortex and cerebellum. Disruption of THBS2 in mice results in a complex phenotype characterized mainly by abnormalities in fibroblasts, connective tissues, blood vessels,26 extracellular matrix deposition, and leakage of the blood-brain-barrier.27 Interestingly, a recent study revealed that matrix protein THBS2, is expressed in vascular smooth muscle cells and capable of altering Notch-3 signaling pathway independent of its canonical ligands.28
In addition to DLL1 and THBS2, a third candidate gene PHF10/(BAF45a), encoding a zing finger domain protein present in the SRO is essential for self-renewal of the multipotent neural stem cells and neuronal differentiation. The development of the vertebrate nervous system occurs at mitotic exit, when cells lose multipotency and begin to develop stable connections. This transition involves a switch in ATP-dependent chromatin-remodeling mechanisms that coincides with terminal differentiation of post-mitotic neurons. The switch involves exchange of subunits consisting of PHF10 within Swi/Snf-like neural-progenitor-specific BAF (npBAF) complexes for the homologous subunits within neuron-specific BAF complexes in post-mitotic neurons. The subunits of the npBAF complex are essential for neural-progenitor proliferation, and mice with reduced dosage for the genes encoding these subunits have defects in neural-tube closure similar to those in human spina bifida.29
A more recent study by Conti et al30 showed that a 1.2 Mb SRO common to 12 patients in the 6q27 region contains plausible genes THBS2, PHF10, TCTE3, DLL1, WDR27, and C6orf70. Of the 12 patients reported in the study, 9 patients had PNH, in addition to other brain malformations. PNH is caused owing to defective neuronal migration that results in nodules around the ventricles. Using whole exome sequencing, one patient with isolated bilateral PNH and epilepsy was found to harbor a de novo missense mutation that affects both the isoforms of C6orf70 corresponding to p.lle250Asn (c.752T>A) in the long isoform and p.lle207Asn (c.622T>A) in the short isoform. Moreover, in utero silencing of C6orf70 altered neuronal migration. These data suggest a potential role for C6orf70 in the development of PNH. As C6orf70 is also present within the 1.7 Mb SRO common to patients 1–7 (Supplementary Figure 1), it is possible that haploinsufficiency of C6orf70 could be contributing to structural brain malformations including PNH as observed in patients 3 and 5 reported here.
The smallest reported deletion in this region is a terminal deletion of 400 kb in a subject (P18 in Figure 3) with intellectual disability, hydrocephalus, and ACC.1 This region contains only three protein-coding genes, PSMB1, TBP, and PDCD2, encoding a proteasome subunit, a TATA box binding protein, and a nuclear protein involved in programmed cell death, respectively, and exhibit significant conserved synteny.31, 32 TBP encodes for TATA-binding protein that associates with TBP-associated factors to form transcription factor IID that is critical for initiation of transcription. Many transcriptional regulators are involved in neurodevelopmental disorders33 and it is possible that the haploinsufficiency of TBP interferes with the normal development of the brain. It is to be noted that the characterization of this small deletion was performed by the use of FISH and the exact location of the centromeric breakpoint could not be delineated. The centromeric breakpoint of the deletion was within 100 kb of DLL1. Genomic deletions can contribute to phenotypes not only by modifying the expression levels of genes within the aneuploid segments but also that of neighboring genes.34 It is hence possible that DLL1 expression may have been altered in this subject with the smallest reported deletion on 6q27.
Although, there is significant phenotypic overlap of common clinical features such as structural brain abnormalities, ear anomalies, developmental delay, mild dysmorphic features, learning difficulties, and hypotonia in all the patients, there is also some degree of phenotypic variability. Various corpus callosum abnormalities including ACC were noted in patients 1, 3, 5, 6, and 7. Patient 7 with the smallest deletion of 1.7 Mb in the 6q27 region has a dysplastic corpus callosum. This suggests that the SRO for structural brain abnormalities observed in all the patients described here, may also include the minimal critical interval mediating corpus callosum abnormalities, as elaborated in the report by O’Driscoll.10 In summary, chromosome 6q27 harbors genes that are critical for the normal development of the brain and it is likely that more than one gene is responsible for causation of the structural brain abnormalities.
References
Eash D, Waggoner D, Chung J, Stevenson D, Martin CL : Calibration of 6q subtelomere deletions to define genotype/phenotype correlations. Clin Genet 2005; 67: 396–403.
Bertini V, De Vito G, Costa R, Simi P, Valetto A : Isolated 6q terminal deletions: an emerging new syndrome. Am J Med Genet A 2006; 140: 74–81.
Sherr EH, Owen R, Albertson DG et al: Genomic microarray analysis identifies candidate loci in patients with corpus callosum anomalies. Neurology 2005; 65: 1496–1498.
Rooms L, Reyniers E, Scheers S et al: TBP as a candidate gene for mental retardation in patients with subtelomeric 6q deletions. Eur J Hum Genet 2006; 14: 1090–1096.
Elia M, Striano P, Fichera M et al: 6q terminal deletion syndrome associated with a distinctive EEG and clinical pattern: a report of five cases. Epilepsia 2006; 47: 830–838.
Striano P, Malacarne M, Cavani S et al: Clinical phenotype and molecular characterization of 6q terminal deletion syndrome: five new cases. Am J Med Genet A 2006; 140: 1944–1949.
Mosca AL, Callier P, Masurel-Paulet A et al: Cytogenetic and array-CGH characterization of a 6q27 deletion in a patient with developmental delay and features of Ehlers-Danlos syndrome. Am J Med Genet A 2010; 152A: 1314–1317.
Gerber JC, Neuhann TM, Tyshchenko N, Smitka M, Hackmann K : Expanding the clinical and neuroradiological phenotype of 6q27 microdeletion: olfactory bulb aplasia and anosmia. Am J Med Genet A 2011; 155A: 1981–1986.
Rigon C, Salviati L, Mandarano R, Dona M, Clementi M : 6q27 subtelomeric deletions: is there a specific phenotype? Am J Med Genet A 2011; 155A: 1213–1214.
O'Driscoll MC, Black GC, Clayton-Smith J, Sherr EH, Dobyns WB : Identification of genomic loci contributing to agenesis of the corpus callosum. Am J Med Genet A 2010; 152A: 2145–2159.
El-Hattab AW, Smolarek TA, Walker ME et al: Redefined genomic architecture in 15q24 directed by patient deletion/duplication breakpoint mapping. Hum Genet 2009; 126: 589–602.
Boone PM, Bacino CA, Shaw CA et al: Detection of clinically relevant exonic copy-number changes by array CGH. Hum Mutat 2010; 31: 1326–1342.
Ou Z, Kang SH, Shaw CA et al: Bacterial artificial chromosome-emulation oligonucleotide arrays for targeted clinical array-comparative genomic hybridization analyses. Genet Med 2008; 10: 278–289.
Gu W, Zhang F, Lupski JR : Mechanisms for human genomic rearrangements. PathoGenet 2008; 1: 4.
Shen-Schwarz S, Hill LM, Surti U, Marchese S : Deletion of terminal portion of 6q: report of a case with unusual malformations. Am J Med Genet 1989; 32: 81–86.
Birnbacher R, Chudoba I, Pirc-Danoewinata H et al: Microdissection and reverse painting reveals a microdeletion 6(q26qter) in a de novo r(6) chromosome. Ann Genet 2001; 44: 13–18.
Artavanis-Tsakonas S, Rand MD, Lake RJ : Notch signaling: cell fate control and signal integration in development. Science 1999; 284: 770–776.
Lui JH, Hansen DV, Kriegstein AR : Development and evolution of the human neocortex. Cell 2011; 146: 18–36.
Bettenhausen B, Hrabe de Angelis M, Simon D, Guenet JL, Gossler A : Transient and restricted expression during mouse embryogenesis of Dll1, a murine gene closely related to Drosophila Delta. Development 1995; 121: 2407–2418.
Lathia JD, Mattson MP, Cheng A : Notch: from neural development to neurological disorders. J Neurochem 2008; 107: 1471–1481.
Hrabe de Angelis M, McIntyre J 2nd, Gossler A : Maintenance of somite borders in mice requires the Delta homologue DII1. Nature 1997; 386: 717–721.
Rubio-Aliaga I, Przemeck GK, Fuchs H et al: Dll1 haploinsufficiency in adult mice leads to a complex phenotype affecting metabolic and immunological processes. PloS one 2009; 4: e6054.
Dupe V, Rochard L, Mercier S et al: NOTCH, a new signaling pathway implicated in holoprosencephaly. Hum Mol Genet 2011; 20: 1122–1131.
Armstrong LC, Bornstein P : Thrombospondins 1 and 2 function as inhibitors of angiogenesis. Matrix Biol 2003; 22: 63–71.
Adams JC : Thrombospondins: multifunctional regulators of cell interactions. Ann Rev Cell Dev Biol 2001; 17: 25–51.
Kyriakides TR, Leach KJ, Hoffman AS, Ratner BD, Bornstein P : Mice that lack the angiogenesis inhibitor, thrombospondin 2, mount an altered foreign body reaction characterized by increased vascularity. Proc Natl Acad Sci USA 1999; 96: 4449–4454.
Tian W, Sawyer A, Kocaoglu FB, Kyriakides TR : Astrocyte-derived thrombospondin-2 is critical for the repair of the blood-brain barrier. Am J Pathol 2011; 179: 860–868.
Meng H, Zhang X, Hankenson KD, Wang MM : Thrombospondin 2 potentiates notch3/jagged1 signaling. J Biol Chem 2009; 284: 7866–7874.
Yoo AS, Staahl BT, Chen L, Crabtree GR : MicroRNA-mediated switching of chromatin-remodelling complexes in neural development. Nature 2009; 460: 642–646.
Conti V, Carabalona A, Pallesi-Pocachard E et al: Periventricular heterotopia in 6q terminal deletion syndrome: role of the C6orf70 gene. Brain 2013; 136 (Pt 11): 3378–3394.
Trachtulec Z, Forejt J : Synteny of orthologous genes conserved in mammals, snake, fly, nematode, and fission yeast. Mamm Genome 2001; 12: 227–231.
Trachtulec Z, Vlcek C, Mihola O, Forejt J : Comparative analysis of the PDCD2-TBP-PSMB1 region in vertebrates. Gene 2004; 335: 151–157.
Hong EJ, West AE, Greenberg ME : Transcriptional control of cognitive development. Curr Opin Neurobiol 2005; 15: 21–28.
Merla G, Howald C, Henrichsen CN et al: Submicroscopic deletion in patients with Williams-Beuren syndrome influences expression levels of the nonhemizygous flanking genes. Am J Hum Genet 2006; 79: 332–341.
Acknowledgements
We thank all the patients and their family members for their cooperation and participation in this research study. We also thank all the research coordinators at BCM-MGL for their assistance in obtaining clinical information. This work was supported in part by fellowship grants by the Clinical Scientist Development Award from The Doris Duke Charitable Foundation (SNSC), DK081735-01A1, NIH /NIGMS T32 contract grant number GM07526 (AE).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on European Journal of Human Genetics website
Rights and permissions
About this article
Cite this article
Peddibhotla, S., Nagamani, S., Erez, A. et al. Delineation of candidate genes responsible for structural brain abnormalities in patients with terminal deletions of chromosome 6q27. Eur J Hum Genet 23, 54–60 (2015). https://doi.org/10.1038/ejhg.2014.51
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2014.51
This article is cited by
-
The phenotypic spectrum of terminal 6q deletions based on a large cohort derived from social media and literature: a prominent role for DLL1
Orphanet Journal of Rare Diseases (2023)
-
Differences in whole-brain metabolism are associated with the expression of genes related to neurovascular unit integrity and synaptic plasticity in temporal lobe epilepsy
European Journal of Nuclear Medicine and Molecular Imaging (2023)
-
Multiple genomic copy number variants associated with periventricular nodular heterotopia indicate extreme genetic heterogeneity
European Journal of Human Genetics (2019)
-
Williams–Beuren syndrome in Mexican patients confirmed by FISH and assessed by aCGH
Journal of Genetics (2019)
-
A feasible diagnostic approach for the translocation carrier from the indication of products of conception
Molecular Cytogenetics (2018)