Abstract
Noninvasive prenatal testing by massive parallel sequencing of maternal plasma DNA has rapidly been adopted as a mainstream method for detection of fetal trisomy 21, 18 and 13. Despite the relative high accuracy of current NIPT testing, a substantial number of false-positive and false-negative test results remain. Here, we present an analysis pipeline, which addresses some of the technical as well as the biologically derived causes of error. Most importantly, it differentiates high z-scores due to fetal trisomies from those due to local maternal CNVs causing false positives. This pipeline was retrospectively validated for trisomy 18 and 21 detection on 296 samples demonstrating a sensitivity and specificity of 100%, and applied prospectively to 1350 pregnant women in the clinical diagnostic setting with a result reported in 99.9% of cases. In addition, values indicative for trisomy were observed two times for chromosome 7 and once each for chromosomes 15 and 16, and once for a segmental trisomy 18. Two of the trisomies were confirmed to be mosaic, one of which contained a uniparental disomy cell line. As placental trisomies pose a risk for low-grade fetal mosaicism as well as uniparental disomy, genome-wide noninvasive aneuploidy detection is improving prenatal management.
Similar content being viewed by others
INTRODUCTION
The presence of circulating cell-free fetal DNA in the maternal plasma of the pregnant woman,1 in combination with recent advances in massively parallel sequencing (MPS) technologies, has made noninvasive prenatal testing (NIPT) of fetal aneuploidy a reality. NIPT reduces the need for invasive sampling and the associated risk of procedure-related pregnancy loss. In 2008, it was demonstrated that noninvasive fetal aneuploidy detection by MPS was feasible.2, 3 Multiple clinical validation studies using either targeted or whole-genome sequencing demonstrated the high sensitivity and specificity of NIPT.4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15 Although most validation studies were predominantly evaluating the clinical validity in pregnancies at increased risk of the most common aneuploidies, it was recently shown that screening all pregnant women has positive predictive values of 45.5% and 40% for detection of trisomies 21 and 18, respectively.16
MPS for aneuploidy detection applies counting statistics to millions of sequencing reads to identify subtle changes in the small percentage of fetal DNA present in the total cell-free DNA isolated from maternal plasma.17, 18 An increase or decrease in the number of normalized sequencing reads, typically converted to a ‘z-score’,18 a ‘normalized chromosome value’,13 genome-wide normalized score19 or by ‘within-sample copy number aberration detector’20 is indicative of aneuploidy for the respective chromosome. Despite the high accuracy of current NIPT testing, a baseline false-positive and false-negative rate remains. Those incorrect results may have both biological and technical causes: (1) maternal CNVs, maternal mosaicism,21 maternal cancers or haematological malignancies,22 and confined placental mosaicism (CPM)23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34 may cause variability of the sequence read count statistics, which could mask or mimic true aneuploidy-related variation, and may, in turn, result in false-positive or false-negative results; (2) the degradation and/or apoptosis of maternal cells following maternal blood sampling would increase the maternal fraction in the plasma and as a consequence reduce the fetal fraction. A low fetal fraction could cause false-negative results and maternal cell degradation may result in low-quality experiments, which can cause false-positive or false-negative results. Here, we present a comprehensive analysis pipeline, which avoids false-positives or false-negatives due to maternal CNVs and improves the robustness of the analysis.
METHODS
Sample preparation and sequencing
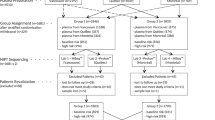
A retrospective validation study was performed with approval of the local ethical committee and informed consent of the pregnant woman. Peripheral blood samples from pregnant women between the gestational ages (g.a.) of 9–24 weeks were collected in either EDTA or Streck tubes. For the following diagnostic NIPT analysis, all maternal blood samples were collected in Streck tubes between 9 and 35 weeks g.a. Women were referred for different reasons, except abnormal US, which is an indication for an invasive test. The plasma was separated on the same day as blood sampling by the standard dual centrifugation method. Briefly, blood samples were centrifuged at 1500 g for 10 min, without brake. Plasma was transferred to either 3x1.5 ml tubes or to 5 ml tubes, which were subject to a second centrifugation at 13 000 g and transferred to sterile tubes for storage at −20 °C before cell-free DNA extraction. Cell-free DNA was extracted using the QIAamp Circulating Nucleic Acid Kit (Qiagen Benelux B.V., Venlo, Netherlands) according to the manufacturer’s recommendations. The final elution volume used was 60 μl. Libraries were prepared using TruSeq ChipSeq Library Preparation Kit (Illumina, San Diego, CA, USA) according to the manufacturer's recommendations with the following modifications: (1) reagent volumes were reduced by half, (2) the gel-free method was used and (3) the final PCR enrichment step used 15 cycles. Every library was analysed on a Fragment Analyser (Advanced Analytical Technologies Inc., Heidelberg, Germany) before sequencing to assess the size distribution as well as the quality of the library. Samples containing high-molecular-weight fragments were classified as not eligible for sequencing. Samples were indexed during library preparation and between 20 and 24 samples were pooled for multiplex sequencing across both lanes of an Illumina HiSeq2500 flow cell. Sequencing was performed on the HiSeq 2500 (Illumina) in fast mode producing 50 bp single-end reads.
The data set containing raw sequencing. BAM files obtained from MPS of the validation data set of 296 NIPT samples is available to academic users upon request to the Data Access Committee of KU Leuven via the corresponding author (JRV).
Array CGH, SNP array and FISH
Array CGH (aCGH) analysis was performed as described previously35 using either the 4x 180 K CytoSure Syndrome Plus Leuven Design microarray or the 8x 60 K CytoSure ISCA v2 microarray (AMADID 252721 and 020040, respectively; Oxford Gene Technology (OGT), Oxford, UK). Visualizations of results and data analysis were performed using the CytoSure Interpret Software (OGT) and with Cartagenia BENCH software (Cartagenia NV, Leuven, Belgium).
SNP array copy number profiling and analysis of regions of homozygosity were performed according to standard procedures using the Infinium CytoSNP-850k BeadChip (Illumina). Subsequently, visualizations of SNP array results and data analysis were performed using Nexus 7.5 software (BioDiscovery, Los Angeles, CA, USA). Results were classified with Cartagenia BENCH software.
Fluorescent in situ hybridization (FISH) was performed following standard procedures.
Data analysis
The 50 bp sequence reads were aligned to the reference genome using the Burrows–Wheeler aligner,36 deduplicated with Picard tools (http://picard.sourceforge.net/) and realigned and recalibrated with GATK.37 To reduce between sample variability, reads were removed that (1) match (suboptimally) at multiple locations, (2) contain mismatches or gaps in the alignment or (3) start in blacklisted regions taken from an in-house curated list of common polymorphic CNVs, collapsed repeats,38 DAC blacklisted regions generated for the ENCODE project39 and the undefined portion of the reference genome (ie, the N’s).
The aligned and filtered reads were counted as described,40 except that normalization was performed with respect to the autosomes only and the calculation was extended beyond chromosomes 13 and 18. We partitioned all autosomes into 50-kb bins, corrected the sequence counts with LOESS regression according to the bin GC content and divided the result by the total sum of autosomal sequence counts to obtain per-bin GC-corrected genomic representations (GR). These were finally aggregated per autosome, and per 5 Mb windows consisting of 100 subsequent 50 kb bins, where the 5 Mb windows are sliding by 50 kb.
A reference set of 100 samples (50 males (46,XY), 50 females (46,XX) pregnancies) was used to determine GR values expected in normal samples both at the chromosome and the 5 Mb bin level. From the mean and standard deviation of GR in this reference set, z-scores for autosomes and 5 Mb bins were calculated for all samples. We defined three extra parameters on top of the traditional chromosome-wide z-score. First, the zz-score is calculated as the standard score of the z-score of a given autosome in comparison with the z-scores of remaining autosomes. Second, the BM (bin median) is calculated from the median of z-scores measured per 5 Mb bin in the autosome of interest. Third, the OM (other median) is the median of the absolute value of the 5 Mb z-scores over the remaining bins.
Trisomy (or monosomy) is then associated with autosomes where: (1) the chromosome-level z-score is high (low), that is, the chromosome has attracted more (less) reads than expected; (2) the zz-score is high (low), that is, the remaining autosomes look normal such that the chromosome of interest stands out; (3) BM is high (low), that is, the increased (decreased) number of reads cannot be attributed to a local event; and (4) OM is low, that is, at the 5 Mb bin level the remaining autosomes also do not show signs of aneuploidy.
Cutoffs for these four parameters were derived from the commonly reported z-score≥3 setting, fixed before starting the validation study described below, and combined into the following decision rule:
-
normal if (−2.5<z-score<2.5) and (−2.5<zz-score<2.5) and (−1<BM<1),
-
trisomy if (z-score≥3) and (zz-score≥3) and (BM≥1.5) and (OM<1),
-
monosomy if (z-score≤−3) and (zz-score≤−3) and (BM≤−1.5) and (OM<1),
-
no trisomy if not monosomy and (z-score<2.5) and (zz-score<2.5) and (BM<1), and moreover (z-score≤−2.5) or (zz-score≤−2.5) or (BM≤−1),
-
no monosomy if not trisomy and (z-score>−2.5) and (zz-score>−2.5) and (BM>−1), and moreover (z-score≥2.5) or (zz-score≥2.5) or (BM≥1) and
-
undetermined in all other cases.
The decision rule incorporates inconclusive zones (no trisomy, no monosomy, undetermined) where confidence is considered too low to distinguish accurately between euploid and aneuploidy, as well as to flag cases where individual parameters are discordant or borderline.
To determine possible reasons for discordant or inconclusive parameters (eg, localized events such as segmental aneuploidy or maternal/fetal CNVs, maternal malignancies or technical noise (see Results and discussion for examples), the visual representation can aid to determine the cause. In those cases where z-score, zz-score or BM fall into an inconclusive range, and there is no support for localized events, no definitive call can be made and a repeat sample is requested (for chromosomes 13, 18 and 21), although the other chromosomes of interest can be reported as normal (eg, normal 21, normal 18, inconclusive for 13). For all other autosomes, aneuploidies are only reported where all parameters are in the abnormal range, and repeat samples are not requested where z, zz and/or BM are inconclusive.
The automated genotype classification achieved with the above rule is systematically applied to all autosomes and reported in combination with a plot of the 5 Mb bin z-scores across the chromosome.
A final sample level quality score (QS) was determined by calculating the standard deviation of all the autosome z-scores following removal of the highest and the lowest scoring chromosomes. Samples with QS exceeding 2 were considered to be poor quality and repeat sampling was requested, and QS between 1.5 and 2 were of intermediate quality.
The sex was determined by verifying the presence of Y-chromosome-specific reads. Because of similarities between X and Y chromosomes, a number of reads in female pregnancies is systematically mapping to the Y chromosome. Twenty 50 kb bins unique to the Y chromosome were identified. Samples with ≧3 Y-specific bins containing >1 read were classified as male pregnancies. In cases where only a single Y-specific bin was found with only a single read, the sample was classified as a female pregnancy. In all other cases, the sex was left undetermined.
The fetal fraction was determined for all the male pregnancies by measuring the difference at the 50 kb bin level in the median number of reads mapping to the autosomes and the median number of reads on the X chromosome, divided by the median number of reads per bin on the autosomes multiplied by two.41
RESULTS
Optimized data analysis pipeline
The traditional approach to identify aneuploidies is to calculate a z-score that quantifies the deviation of the number of sequence reads at the chromosome level from the number expected in normal, that is, euploid, samples.17, 18 We have defined two new parameters, resulting in higher accuracy for aneuploidy detection by zooming out and in from the chromosome level (Figure 1 and Supplementary Figure 1a). (1) Zooming out to the genome level, we use the z-scores of the remaining, supposedly euploid, chromosomes to calculate a sample-specific mean and standard deviation of z-scores for ‘euploid’ chromosomes. With these values, we calculate a meta z-score (zz-score) that quantifies the deviation of the chromosome z-score from the z-score of ‘euploid’ chromosomes within the same sample. (2) Zooming in to overlapping 5 Mb subchromosomal bins, we break down the chromosome-level z-score by recalculating it per bin. The median z-score across these bins (BM) discriminates between (a) aneuploidies, where consistently high or low z-scores across all 5 Mb bins are reflected in higher or lower BM values, and (b) events that span less than half of the chromosome and leave the BM largely unaffected. Notably, (small) maternal CNVs are present at a high percentage and can skew the scores significantly. Visualization of the events in (b), achieved by plotting the per-bin z-scores across the chromosome, further assist in the clinical interpretation of cases where a high chromosome-level z-score is corrected by a low BM value. Such cases are flagged as inconclusive by the above decision rule.
An example report of a trisomy 21 sample. Details are shown for chromosomes 13, 18 and 21, and for each chromosomes z, zz and BM scores are indicated. When aneuploidy is present, the OM is shown for comparison with BM score. Dot plots of z- vs zz-score (Y and X axis, respectively) are shown with undetermined zones marked (2.5–3) with grey lines. Z-scores of 5 Mb sliding windows are plotted across the chromosome to visualize the aberrations, dotted lines represent ±1.5 z-scores and the areas above and below these cutoff values are coloured in green and red, respectively. Violin plots of BM and OM with their respective median shown with a single dot represent the distribution of all the bin-level z-scores.
The introduction of those statistical measures allows us to address the major causes of errors. First, maternal CNVs would skew the z statistics leading to false-positive and false-negative aneuploidy calls, as demonstrated for four different NIPT results shown in Figure 2. Actually, with z-scores alone, seven false-positive calls for trisomy 13 (4 cases), 18 (2 cases) and 21 (1 case) would be made in 1646 samples (4 in the diagnostic and 3 in the validation data set) (see Figure 3).
Four examples of maternal CNVs that alter counting statistics causing potential false-positive results. For every sample, the NIPT (top) result and the aCGH (bottom) result is shown: (a) 2.55 Mb deletion on chromosome 22q11.21 (chr21.hg19: g.18894820–21440515del); (b) 1.38 Mb duplication on chromosome 13q33.1–q33.2 (chr13.hg19: g.104397732–1057779818dup); (c) 1.73 Mb duplication on chromosome 13q31.3 (chr13.hg19: g.90201888–91936194dup); (d) 1.03 Mb duplication on chromosome 21q11.2–q21.1 (chr21.hg19: g.15485038–16515907dup). While duplications and deletions affect both z- and zz-score, the BM remains in a normal range. The sliding plots help visualize the localization of the aberration as well as the effect.
Z-score vs zz-score (left), and BM vs zz-score (right) for chromosomes 21 and 18 results from all validation study samples. This highlights how the combined analysis reduces potential false-positive and false-negative cases. Triangles and boxes represent trisomies 18 and 21, respectively. Full diamond shape is the mosaic trisomy 18 case, whereas the hollow diamond shapes represent the undetermined trisomy 18 cases. Hollow circles represent the normal population, and full circles represent the cases classified as undetermined by the automated pipeline. The shaded lines on the X and Y axis mark the inconclusive zones between the cutoff values of 2.5–3 for both z- and zz-score, and 1–1.5 for the BM.
The low BM in these four examples, one on chromosome 22, resulting in a negative z-score, two on chromosome 13 and one on chromosome 21, resulting in z-scores >3, is indicative for the presence of a maternal CNV. Indeed, subsequent array analysis of the maternal white blood cell DNA confirmed that these local deviant z-scores were of maternal origin. Second, technical artefacts, maternal mosaicism, maternal cancers or haematological malignancies, as well as placental disorders, may cause variability of the sequence read count statistics, which could mask or mimic true aneuploidy-related variation, and may, in turn, result in false-positive or false-negative results. Such genome-wide variation would result in low BM but high OM. An example is shown in Supplementary Figure 1b. The presence of high genome-wide variation also leads to a high QS, requiring a repeat sample. When the same genome-wide variation is observed upon resampling, a biological cause of this variation can be suspected.
To estimate the influence of number of reads on the accuracy of the test, we performed an in silico analysis. An NIPT sequencing data set of a normal pregnancy (46,XX) was mixed with a data set from a trisomic male fetus (47,XY,+21), and was repeated using a second trisomy 21 male case. Sequencing reads were mixed in silico at the FASTQ level, mixing, respectively, 2, 3, 4, 5, 6, 7, 8 and 9 million randomly selected reads (Supplementary Figure 2) and targeting calculated fetal fractions from 2% to 8%, with 0.1% increments. The results of this in silico experiment demonstrate that false negatives for chromosome 21 were absent with a fetal fraction above 3.3% and >7 million analysed sequencing reads. On this basis, 7 million analysed reads were selected as the minimum required read number. This equates to a minimum of ~10 million raw reads based on our analysis pipeline.
Validation
A retrospective validation study was performed to test the accuracy of the developed analysis pipeline. The study cohort consisted of 296 pregnant women who were initially referred for invasive prenatal testing. Chorionic villus sampling or amniocentesis followed by conventional karyotyping, FISH or genomic microarray analysis (aCGH) revealed 17 cases positive for trisomy 21, 9 for trisomy 18 and 2 for trisomy 13. In addition to these aneuploidies, one mosaic fetal trisomy 18, 16 and 13 as well as one placental mosaic trisomy 13 were diagnosed.
A total of 310 NIPT experiments were performed as eight samples were repeated, and three samples were repeated in triplicate, to assess the reproducibility and repeatability of the analysis pipeline. Repeat samples gave comparable results regarding (i) general sample quality (QS), (ii) aneuploidy detection (parameters z-score, zz-score, OM and BM) and graphical plot of 5 Mb bin z-scores across all chromosomes (Supplementary Figure 3). An example patient report is shown in Supplementary Figure 4.
Out of 296 samples, a single sample was excluded after library preparation owing to the presence of maternal peripheral blood DNA contamination (most likely due to blood sampling in EDTA tube and a delay in plasma separation). The remaining 295 samples were sequenced and analysed using the pipeline described above, and the parameters (QS, z-score, zz-score, OM and BM) assessed for every chromosome. Four samples did not meet the QS criteria (QS>2) and were excluded from further analysis, whereas four samples were of intermediate quality (1.5<QS<2). Of the 287 samples that passed these quality criteria, 24 samples had inconclusive parameter value(s) for chromosome 13, 18 and/or 21. However, when considering the respective z-, zz-, OM and BM scores in combination with a visual analysis of the chromosomal z-scores across 5 Mb bin plots, a correct result could be determined in 16 of those 24 cases. In 8 cases, including 2 trisomy 18 cases, a repeat sample would be requested. Consequently, 279 samples were correctly classified (including all 17 trisomy 21 cases and 7 of the 9 trisomy 18 cases).
In addition to correct classification of 17 full trisomy 21 cases and 7 full trisomy 18 cases, the NIPT analysis pipeline identified two fetal trisomy 13 cases, one fetal mosaic trisomy 18 and one placental mosaic trisomy 13 case. However, it failed to detect the sample with trisomy 16 and one fetal mosaic trisomy 13. Furthermore, fetal sex was correctly identified in all pregnancies.
Taken together, this NIPT analysis pipeline is robust, reproducible and repeatable for sex determination and the detection of trisomy 21 and trisomy 18.
Diagnostic results
The validated analysis pipeline was applied prospectively to 1350 diagnostic samples from women with a gestational age between 9 and 35 weeks. A total of 60% of women were referred because of an increased risk for trisomy 21; 24% of women had an increased trisomy risk as assessed by the combined test; 33% had an increased risk of trisomy because of maternal age (36 years and older); and 3% because of prior familial history of trisomy 21. Forty percent (40%) of patients were classified as low risk and referred because of maternal anxiety.
Repeat sampling was requested in 23 because of high standard deviation of the NIPT results (QS>1.5). For two no second maternal blood sample was received. In 14 out of 21 repeated samples, the standard deviation was below 1.5 (QS<1.5). A normal result could be issued in all but two of these 21 repeated samples. Additionally, nine samples showed an undetermined result for either chromosome 21 (n=5) or chromosome 18 (n=4), and for six of those a repeat sample was obtained, which gave a normal result. Upon second sampling, the fetal fraction (in males) was on average 1% higher compared with first sampling. Hence, a result could be issued in 97.6% (1318/1350) at first sampling and 99.9% (1343/1345) following repeat sampling.
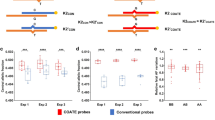
Of those 1350 patients, 11 were positive for trisomy 21 and 4 for trisomy 18 (Figures 4a and b). All trisomy 21 and trisomy 18 cases were confirmed by either FISH or aCGH following invasive prenatal testing. One sample positive for trisomy 18 by NIPT proved to be mosaic as 77% of the chorionic villus cells showed three FISH signals with a centromere 18 probe. aCGH analysis performed on DNA extracted from amniotic fluid cells was normal, demonstrating that the presence of the trisomic line was (mainly) confined to the placenta. In addition to the traditional trisomies 21 and 18, the genome-wide analysis revealed two cases of trisomy 7, one case of trisomy 15 and one case of trisomy 16 (Figures 4c–e). The trisomy 15 was mosaic in about 20% of fetal cells as shown by SNP array analysis on DNA isolated from uncultured amniotic fluid cells. A subsequent MLPA test targeted to the 15q11–q13 locus showed a maternal heterodisomy in the Prader– Willi/Angelman critical region. The trisomy 16 was confirmed to be mosaic (7% of cells) by FISH using the centromere 16 probe on amniotic fluid cells. One predicted trisomy 7 could not be confirmed by FISH following amniocentesis, this being suggestive for CPM. The second positive trisomy 7 result could not be further investigated as the patient did not wish to undergo invasive testing.
Dot plot of zz-scores (Y axis) vs BM (X axis) for all diagnostic study samples for chromosomes 21, 18, 7, 15 and 16. Blue circles represent the normal samples and black circles represent the cases classified as undetermined from automated pipeline. The red boxes represent the cases classified as trisomy for the respective chromosome.
One patient was referred for invasive sampling owing to fetal ultrasound abnormalities. NIPT was performed concomitantly with array CGH analysis, both of which detected a partial trisomy 18, which is 29 Mb in size (Figure 5). This indicates the importance of the novel parameters we introduce of zz and BM, in combination with the genome-wide chromosome plots.
DISCUSSION
NIPT is reducing the need for invasive prenatal genetic diagnosis. Nevertheless, when applied to all pregnant women, the positive predictive value for trisomy 21 detection is currently at best about 50%.16 Hence, one in every two women with a positive NIPT has a normal diploid fetus. The analysis pipeline presented has the potential to reduce this false-positive rate and hence the number of unnecessary invasive tests.
We analysed the z-scores genome wide in 5 Mb intervals, which provides a genome-wide visualization of all potential fetal abnormalities. Similar, although slightly different in approach, genome-wide scanning of binned z-scores has been applied before, to demonstrate the feasibility to noninvasively detect fetal subchromosomal imbalances by deep sequencing,42, 43 as well as by ‘shallow-depth’ sequencing.44 Whereas those studies have focused on the ability to detect noninvasively ever smaller fetal copy number variants with the aim to reduce the need for invasive sampling and chromosomal microarray analysis, we have focused on using this additional information to improve the quality of the aneuploidy detection. When applying only chromosomal z-score test statistics, aberrant test results may result from either the presence of maternal CNVs or by low quality of the DNA samples. First, the presence of maternal CNVs can skew z-scores such that they become indicative for fetal trisomy or monosomy. By visualizing the bin median, a maternal CNV markedly influencing the z-score can readily be identified. It is tempting to speculate that the high false-positive rate for trisomy 13 in large-scale NIPT studies results from the higher incidence of rare but benign CNVs in the population. A second potential source of error is genome-wide variation. If there is large genome-wide variation in the z-scores, the chromosome-specific z-score is less reliable. Genome-wide variation due to degradation of maternal lymphocytes may result in deviant test statistics (data not shown). Other biological determinants in the maternal plasma, such as free-floating tumour DNA, might also lead to erroneous z-scores. By introducing the zz-score, this genome-wide variation is accounted for, which in turn improves the accuracy of aneuploidy detection. We obtained a 100% sensitivity and specificity for trisomy 21 and 18 detection (95% CI: trisomy 21 – sensitivity 80.33–100% and specificity 98.59–100%; trisomy 18 – sensitivity 58.93–100% and specificity 98.64–100%), which is in line with other studies. For 97.6% of the diagnostic samples, an NIPT result was obtained immediately and for 99.9% when including a second sampling. The second sampling resulted in lower standard deviations in all but one case. It seems plausible that improved quality is resulting from increased fetal fractions at later gestational age. Interestingly, in 7 out of 13 repeat samples, which were due to high QS, despite lower standard deviation, similar z-score variation profiles were observed across all chromosomes when compared with the original sample. This suggests a biological rather than a technical cause for the repeatedly observed variation.
Over the past 3 years, NIPT has been extensively validated for the detection of the most frequent viable aneuploidies, trisomy 13, 18 and 21. However, the presence of other chromosomal aneuploidies has, in general, been neglected in those validation studies, and they are not reported by commercial NIPT providers. The ability to detect noninvasively other fetal aneuploidies has already been demonstrated.5, 29 In this study, trisomies other than 13, 18 and 21 were identified by NIPT in 4/1350 samples or 0.3% of all pregnancies. Trisomies were identified for chromosomes 7, 15 and 16. In three, an amniocentesis was performed followed by aCGH or SNP array, and MLPA or FISH analysis of the amniotic fluid cells. In two, the presumed chromosomal trisomies were shown to be low-grade mosaic disomic/trisomic in the amniotic fluid. As full trisomies would not be viable, it may not be surprising that the fetuses are mosaic. Nevertheless, it is striking that such low-grade fetal mosaicism can readily be identified by noninvasive testing. It therefore seems likely that the majority of trophoblast cells would be trisomic and that the low-grade mosaicism would be confined to the fetus. One trisomy 7 could not be confirmed upon invasive testing. Further follow-up analysis of placental material at birth may assist in confirming this hypothesis. CPM is estimated to occur in 1–2% of pregnancies.26, 27 Discordant results between fetal karyotyping and NIPT by maternal plasma sequencing has been demonstrated in a case of uniparental disomy 21 due to trisomic rescue.32 Hence, although our analysis pipeline may increase the accuracy of NIPT, the main cause for discrepancy between the noninvasive and the invasive test, however, is likely the high incidence of placental mosaicism. As a consequence, careful follow-up by either invasive testing or ultrasound examination will remain essential in the clinical management of those aneuploidies.
The detection of additional trisomies other than the common autosomal trisomies 13, 18 and 21 has important clinical implications. First, all those fetuses are at high risk for uniparental disomies, resulting from the loss of one chromosome following a trisomic conception. Uniparental disomies of chromosomes 7 and 15 are known to cause developmental disorders, respectively, Silver Russel syndrome and Prader–Willi or Angelman syndrome. In our series, we demonstrate that trisomy 15 identified by NIPT revealed the presence of a uniparental disomy in the fetus. Uniparental isodisomies for other chromosomes are at increased risk of developing recessive disorders. Hence, UPD testing following an invasive sampling is advisable in the follow-up of those mosaic trisomies. Second, for all those chromosomal trisomies, low-grade mosaicism can be viable and have been detected in patients with birth defects.
Genome-wide aneuploidy analysis enables not only the detection of whole chromosomal aneuploidies but also allows the detection of segmental chromosomal imbalances. A number of case reports, as well as retrospective and prospective validation studies of novel algorithms, have recently been reported on the ability to detect segmental imbalances and even submicroscopic CNVs.5, 13, 20, 29, 43, 44, 45, 46, 47, 48 Lau et al29, 47 reported on the use of the FCAPS pipeline detecting a fetal duplication of 18p, which is 14 Mb in size. Here, we show the detection of a segmental trisomy 18, which is 29 Mb in size, demonstrating the power of this genome-wide profiling approach.
The clinical interpretation in the context of the respective pregnancy and the associated US anomalies (if any) remains important. Owing to CPM (and other factors, both technical and biological), chromosomal anomalies may be identified from the analysis of cell-free DNA, which do not represent the true fetal genotype. Extending the screen to all chromosomal aneuploidies pertain a risk of overdiagnosis with a higher number of false positives leading to unnecessary invasive testing. This risk is especially increased because a proper validation study to determine the false-positive and false-negative rate for those other aneuploidies is virtually impossible. In the absence of US anomalies most, if not all, aneuploidies other than trisomy 21 represent mosaicism and an invasive genetic test is not a good reference as it does not necessarily sample the same cells as those giving rise to the cell-free fetal DNA in the maternal plasma. However, as two out of three potential trisomies with a follow-up invasive genetic analysis were confirmed true positives by the presence of a trisomy in a fraction of the amniotic fluid cells, this would suggest a high sensitivity, which warrants follow-up invasive testing. The detection of other trisomies in 0.3% of all pregnancies, a figure which is within the range of miscarriages known to result from other trisomies, suggests a high specificity as well. It remains, however, questionable as to whether an invasive test should be recommended for these patients. It has been suggested that the investigation of trisomy 13 detected upon NIPT, for example, in low-risk patients and in the absence of US anomalies may do more harm than good.49 A similar note of caution can be made for all other (segmental) aneuploidies.
References
Lo YM, Corbetta N, Chamberlain PF et al: Presence of fetal DNA in maternal plasma and serum. Lancet 1997; 350: 485–487.
Chiu RW, Chan KC, Gao Y et al: Noninvasive prenatal diagnosis of fetal chromosomal aneuploidy by massively parallel genomic sequencing of DNA in maternal plasma. Proc Natl Acad Sci USA 2008; 105: 20458–20463.
Fan HC, Blumenfeld YJ, Chitkara U, Hudgins L, Quake SR : Noninvasive diagnosis of fetal aneuploidy by shotgun sequencing DNA from maternal blood. Proc Natl Acad Sci USA 2008; 105: 16266–16271.
Ashoor G, Syngelaki A, Wagner M, Birdir C, Nicolaides KH : Chromosome-selective sequencing of maternal plasma cell-free DNA for first-trimester detection of trisomy 21 and trisomy 18. Am J Obstet Gynecol 2012; 206: 322–325.
Bianchi DW, Platt LD, Goldberg JD, Abuhamad AZ, Sehnert AJ, Rava RP : Genome-wide fetal aneuploidy detection by maternal plasma DNA sequencing. Obstet Gynecol 2012; 119: 890–901.
Chiu RW, Akolekar R, Zheng YW et al: Non-invasive prenatal assessment of trisomy 21 by multiplexed maternal plasma DNA sequencing: large scale validity study. BMJ 2011; 342: c7401.
Dan S, Wang W, Ren J et al: Clinical application of massively parallel sequencing-based prenatal noninvasive fetal trisomy test for trisomies 21 and 18 in 11,105 pregnancies with mixed risk factors. Prenat Diagn 2012; 32: 1225–1232.
Ehrich M, Deciu C, Zwiefelhofer T et al: Noninvasive detection of fetal trisomy 21 by sequencing of DNA in maternal blood: a study in a clinical setting. Am J Obstet Gynecol 2011; 204: 205–211.
Nicolaides KH, Syngelaki A, Ashoor G, Birdir C, Touzet G : Noninvasive prenatal testing for fetal trisomies in a routinely screened first-trimester population. Am J Obstet Gynecol 2012; 207: 374–376.
Norton ME, Brar H, Weiss J et al: Non-Invasive Chromosomal Evaluation (NICE) Study: results of a multicenter prospective cohort study for detection of fetal trisomy 21 and trisomy 18. Am J Obstet Gynecol 2012; 207: 137–138.
Palomaki GE, Deciu C, Kloza EM et al: DNA sequencing of maternal plasma reliably identifies trisomy 18 and trisomy 13 as well as Down syndrome: an international collaborative study. Genet Med 2012; 14: 296–305.
Palomaki GE, Kloza EM, Lambert-Messerlian GM et al: DNA sequencing of maternal plasma to detect Down syndrome: an international clinical validation study. Genet Med 2011; 13: 913–920.
Sehnert AJ, Rhees B, Comstock D et al: Optimal detection of fetal chromosomal abnormalities by massively parallel DNA sequencing of cell-free fetal DNA from maternal blood. Clin Chem 2011; 57: 1042–1049.
Sparks AB, Struble CA, Wang ET, Song K, Oliphant A : Noninvasive prenatal detection and selective analysis of cell-free DNA obtained from maternal blood: evaluation for trisomy 21 and trisomy 18. Am J Obstet Gynecol 2012; 206: 319.
Sparks AB, Wang ET, Struble CA et al: Selective analysis of cell-free DNA in maternal blood for evaluation of fetal trisomy. Prenat Diagn 2012; 32: 3–9.
Bianchi DW, Parker RL, Wentworth J et al: DNA sequencing versus standard prenatal aneuploidy screening. N Engl J Med 2014; 370: 799–808.
Chiu RW, Cantor CR, Lo YM : Non-invasive prenatal diagnosis by single molecule counting technologies. Trends Genet 2009; 25: 324–331.
Fan HC, Quake SR : Sensitivity of noninvasive prenatal detection of fetal aneuploidy from maternal plasma using shotgun sequencing is limited only by counting statistics. PLoS One 2010; 5: e10439.
Yeang CH, Ma GC, Hsu HW et al: Genome-wide normalized score: a novel algorithm to detect fetal trisomy 21 during non-invasive prenatal testing. Ultrasound Obstet Gynecol 2014; 44: 25–30.
Straver R, Sistermans EA, Holstege H, Visser A, Oudejans CB, Reinders MJ : WISECONDOR: detection of fetal aberrations from shallow sequencing maternal plasma based on a within-sample comparison scheme. Nucleic Acids Res 2014; 42: e31.
Yao H, Zhang L, Zhang H et al: Noninvasive prenatal genetic testing for fetal aneuploidy detects maternal trisomy X. Prenat Diagn 2012; 32: 1114–1116.
Osborne CM, Hardisty E, Devers P et al: Discordant noninvasive prenatal testing results in a patient subsequently diagnosed with metastatic disease. Prenat Diagn 2013; 33: 609–611.
Choi H, Lau TK, Jiang FM et al: Fetal aneuploidy screening by maternal plasma DNA sequencing: 'false positive' due to confined placental mosaicism. Prenat Diagn 2013; 33: 198–200.
Futch T, Spinosa J, Bhatt S, de FE, Rava RP, Sehnert AJ : Initial clinical laboratory experience in noninvasive prenatal testing for fetal aneuploidy from maternal plasma DNA samples. Prenat Diagn 2013; 33: 569–574.
Gao Y, Stejskal D, Jiang F, Wang W : False-negative trisomy 18 non-invasive prenatal test result due to 48,XXX,+18 placental mosaicism. Ultrasound Obstet Gynecol 2014; 43: 477–478.
Hahnemann JM, Vejerslev LO : Accuracy of cytogenetic findings on chorionic villus sampling (CVS) – diagnostic consequences of CVS mosaicism and non-mosaic discrepancy in centres contributing to EUCROMIC 1986-1992. Prenat Diagn 1997; 17: 801–820.
Hahnemann JM, Vejerslev LO : European collaborative research on mosaicism in CVS (EUCROMIC) – fetal and extrafetal cell lineages in 192 gestations with CVS mosaicism involving single autosomal trisomy. Am J Med Genet 1997; 70: 179–187.
Hall AL, Drendel HM, Verbrugge JL et al: Positive cell-free fetal DNA testing for trisomy 13 reveals confined placental mosaicism. Genet Med 2013; 15: 729–732.
Lau TK, Jiang FM, Stevenson RJ et al: Secondary findings from non-invasive prenatal testing for common fetal aneuploidies by whole genome sequencing as a clinical service. Prenat Diagn 2013; 33: 602–608.
Mao J, Wang T, Wang BJ et al: Confined placental origin of the circulating cell free fetal DNA revealed by a discordant non-invasive prenatal test result in a trisomy 18 pregnancy. Clin Chim Acta 2014; 433C: 190–193.
Mennuti MT, Cherry AM, Morrissette JJ, Dugoff L : Is it time to sound an alarm about false-positive cell-free DNA testing for fetal aneuploidy? Am J Obstet Gynecol 2013; 209: 415–419.
Pan M, Li FT, Li Y et al: Discordant results between fetal karyotyping and non-invasive prenatal testing by maternal plasma sequencing in a case of uniparental disomy 21 due to trisomic rescue. Prenat Diagn 2013; 33: 598–601.
Srebniak MI, Diderich KE, Noomen P, Dijkman A, de VF, Opstal DV : Abnormal NIPT results concordant with the karyotype of the cytotrophoblast, but not reflecting the abnormal fetal karyotype. Ultrasound Obstet Gynecol. 2014; 44: 109–111.
Wang Y, Zhu J, Chen Y et al: Two cases of placental T21 mosaicism: challenging the detection limits of non-invasive prenatal testing. Prenat Diagn 2013; 33: 1207–1210.
Brady PD, Delle CB, Christenhusz G et al: A prospective study of the clinical utility of prenatal chromosomal microarray analysis in fetuses with ultrasound abnormalities and an exploration of a framework for reporting unclassified variants and risk factors. Genet Med 2013; 16: 469–476.
Li H, Durbin R : Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 2009; 25: 1754–1760.
McKenna A, Hanna M, Banks E et al: The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 2010; 20: 1297–1303.
Pickrell JK, Gaffney DJ, Gilad Y, Pritchard JK : False positive peaks in ChIP-seq and other sequencing-based functional assays caused by unannotated high copy number regions. Bioinformatics 2011; 27: 2144–2146.
Bernstein BE, Birney E, Dunham I, Green ED, Gunter C, Snyder M : An integrated encyclopedia of DNA elements in the human genome. Nature 2012; 489: 57–74.
Chen EZ, Chiu RW, Sun H et al: Noninvasive prenatal diagnosis of fetal trisomy 18 and trisomy 13 by maternal plasma DNA sequencing. PLoS One 2011; 6: e21791.
Rava RP, Srinivasan A, Sehnert AJ, Bianchi DW : Circulating fetal cell-free DNA fractions differ in autosomal aneuploidies and monosomy X. Clin Chem 2014; 60: 243–250.
Srinivasan A, Bianchi DW, Huang H, Sehnert AJ, Rava RP : Noninvasive detection of fetal subchromosome abnormalities via deep sequencing of maternal plasma. Am J Hum Genet. 2013; 92: 167–176.
Yu SC, Jiang P, Choy KW et al: Noninvasive prenatal molecular karyotyping from maternal plasma. PLoS One 2013; 8: e60968.
Chen S, Lau TK, Zhang C et al: A method for noninvasive detection of fetal large deletions/duplications by low coverage massively parallel sequencing. Prenat Diagn 2013; 33: 584–590.
Peters D, Chu T, Yatsenko SA et al: Noninvasive prenatal diagnosis of a fetal microdeletion syndrome. N Engl J Med 2011; 365: 1847–1848.
Jensen TJ, Dzakula Z, Deciu C, van den Boom D, Ehrich M : Detection of microdeletion 22q11.2 in a fetus by next-generation sequencing of maternal plasma. Clin Chem 2012; 58: 1148–1151.
Lau TK, Cheung SW, Lo PS et al: Non-invasive prenatal testing for fetal chromosomal abnormalities by low-coverage whole-genome sequencing of maternal plasma DNA: review of 1982 consecutive cases in a single center. Ultrasound Obstet Gynecol 2014; 43: 254–264.
Chu T, Yeniterzi S, Rajkovic A et al: High resolution non-invasive detection of a fetal microdeletion using the GCREM algorithm. Prenat Diagn 2014; 34: 469–477.
Verweij EJ, de Boer MA, Oepkes D : Non-invasive prenatal testing for trisomy 13; more harm than good? Ultrasound Obstet Gynecol. 2014; 44: 112–114.
Acknowledgements
This work has been made possible by University of Leuven (KU Leuven) SymBioSys (PFV/10/016 and GOA/12/015 to JRV and KD) and by funding from the Belgian Science Policy Office Interuniversity Attraction Poles (BELSPO-IAP) programme through the project IAP P7/43-BeMGI. HVE and KD are ‘Clinical Investigators’ of the Fonds voor Wetenschappelijk Onderzoek Vlaanderen (FWO). We thank Elly Pijkels and Kristien Philippe for their contribution to the validation study. We are grateful to all women who consented to participate in our validation study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on European Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Bayindir, B., Dehaspe, L., Brison, N. et al. Noninvasive prenatal testing using a novel analysis pipeline to screen for all autosomal fetal aneuploidies improves pregnancy management. Eur J Hum Genet 23, 1286–1293 (2015). https://doi.org/10.1038/ejhg.2014.282
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2014.282
This article is cited by
-
Population screening for 15q11-q13 duplications: corroboration of the difference in impact between maternally and paternally inherited alleles
European Journal of Human Genetics (2024)
-
Prenatal diagnosis of a 46,XY karyotype female fetus with an SRY-associated gonadal dysgenesis, conceived through an intracytoplasmic sperm injection: a case report
BMC Pregnancy and Childbirth (2022)
-
Rare autosomal trisomies detected by non-invasive prenatal testing: an overview of current knowledge
European Journal of Human Genetics (2022)
-
Machine learning-based detection of immune-mediated diseases from genome-wide cell-free DNA sequencing datasets
npj Genomic Medicine (2022)
-
Estimation of cell-free fetal DNA fraction from maternal plasma based on linkage disequilibrium information
npj Genomic Medicine (2021)