Abstract
Genealogical analysis has proven a useful method to understand the origins and frequencies of hereditary diseases in many populations. However, this type of analysis has not yet been used for the investigation of drug intolerance among patients suffering from inherited disorders. This study aims to do so, using data from familial hypercholesterolemia (FH) patients receiving high doses of statins. The objective is to measure and compare various genealogical parameters that could shed light on the origins and heritability of muscular intolerance to statins using FH as a model. Analysis was performed on 224 genealogies from 112 FH subjects carrying either the low-density lipoprotein receptor (LDLR) prom_e1 deletion>15 kb (n=28) or c.259T>G (p.Trp87Gly) (n=84) mutations and 112 non-FH controls. Number of ancestors, geographical origins and genetic contribution of founders, inbreeding and kinship coefficients were calculated using the S-Plus-based GENLIB software package. For both mutations, repeated occurrences of the same ancestors are more frequent among the carriers’ genealogies than among the controls’, but no difference was observed between tolerant and intolerant subjects. Founders who may have introduced both mutations in the population appear with approximately the same frequencies in all genealogies. Kinship coefficients are higher among carriers, with no difference according to statins tolerance. Inbreeding coefficients are slightly lower among >15-kb deletion carriers than among c.259 T>G carriers, but the differences between tolerants and intolerants are not significant. These findings suggest that although muscular intolerance to statins shows a family aggregation, it is not transmitted through the same Mendelian pattern as LDLR mutations.
Similar content being viewed by others
INTRODUCTION
Genealogical analysis has proven a useful method to understand the origins and frequencies of hereditary diseases in many populations.1, 2, 3 This type of analysis is based on data that provide genealogical links between individuals in a given population, through their ancestors who were identified during the construction of the genealogies. It can yield valuable information about the structure of the population, such as kinship and inbreeding levels. Genealogical links are also used to estimate the genetic contributions of ancestors to the present-day population, according to various characteristics, such as geographical origin, ethnicity, period of arrival in the population, etc. To our knowledge, genealogical analysis has not yet been used for the investigation of drug intolerance among patients suffering from inherited disorders. This study aims to do so using data from familial hypercholesterolemia (FH) patients. These patients receive high doses of statins, which are the most widely prescribed class of drugs.4
Statins are proven to safely prevent coronary artery disease (CAD) by different mechanisms, including reducing low-density lipoprotein (LDL)-cholesterol levels.4, 5 Although well tolerated, statins are associated with muscular and non-muscular side effects that affect compliance to drug treatment. Muscular intolerance is a common cause of statin discontinuation.6 It follows a continuum from non-specific or atypical myalgias to the full-blown rhabdomyolysis syndrome.7, 8 Although rhabdomyolysis is rare, statin intolerance is a real problem that calls for better knowledge. The more benign muscular symptoms can have important consequences; they limit the clinical and socio-economic benefits that statins usually offer. Moreover, considering the large (and growing) number of patients using statins, the absolute number of individuals complaining of intolerance symptoms is substantial, which makes the associated economic and clinical burden even greater.9
Type of statins, age, gender and daily dose are well-known modulators of the muscular side effects of statins.10, 11 Data also suggest a specific genetic influence in statin intolerance development, although results of these association studies appear inconsistent.12 Apart from rare mutations known to be associated with intrinsic muscle diseases,13 no common gene polymorphism has yet been identified as an undisputed cause of muscular intolerance to statins. However, family aggregation of intolerance to statins is regularly observed by physicians prescribing statins; this observation supports the importance of gene factor transmission.14
FH is a clinical syndrome characterized by high plasma concentrations of LDL-cholesterol and apolipoprotein (apo) B, tendinous xanthomas and an increased risk of premature CAD.15 Most often, FH is caused by mutations in the LDL receptor (LDLR) gene, although it can also be the consequence of genetic variations in APOB, proprotein convertase subtilisin/kexin type 9 or LDLR adaptator protein-1 genes (ARH).16 FH has been widely used as a homogeneous model disease for excessive plasma LDL-cholesterol levels and atherosclerosis. It has played a critical role in drug development and identification of various CAD gene modulators.15 FH patients are among those receiving the highest daily doses of statins. Because the daily dose is an important modulator of statins intolerance, FH patients without signs and symptoms of intolerance to statins could therefore be considered as real statin-tolerant subjects. This makes FH an interesting model to study statin intolerance.
The objective of the present study was to measure and compare various genealogical parameters that could shed some light on the origins and heritability of muscular intolerance to statins using FH as a model. Deep-rooted genealogies of tolerant and intolerant French-Canadian carriers of two LDLR mutations from the Saguenay-Lac-St-Jean (SLSJ) population (Quebec, Canada) were analyzed and compared.
MATERIALS AND METHODS
SLSJ population
The SLSJ region is located 200 km north of Quebec city (Figure 1). Settlement in this region began during the second quarter of the nineteenth century.17 During the first 100 years, the population increased rapidly due to high fertility levels.18 Today, the population of SLSJ is estimated at around 273 000.19
Founder effects in the SLSJ population became apparent with the relatively high frequency of some rare inherited disorders, such as FH.1, 20 FH affects approximately one per 500 individuals worldwide,21 but the prevalence is significantly higher in the SLSJ population, where one out of 83 individuals is affected.22 Almost 90% of all cases are the consequence of two LDLR gene mutations: a >15- kb deletion (prom_e1 deletion>15 kb) in the promoter and exon 1 and a missense mutation c.259T>G (p.Trp87Gly, rs121908025:T>G), formerly identified as p.Trp66Gly (W66G) in exon 3.23 (Both variants are described in the Familial Hypercholesterolemia database (http://www.ucl.ac.uk/ldlr/LOVDv.1.1.0/) and UMD Locus-Specific Databases (http://www.umd.be/LDLR/).
Data
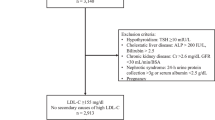
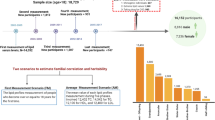
Analysis was performed using extended ascending genealogies from SLSJ FH subjects carrying either the >15-kb deletion or c.259T>G mutation. Considering that the SLSJ population experienced strong founder effects,24, 25, 26 it is assumed that FH patients carrying the same mutation are most probably identical by descent. The c.259T>G mutation was detected by PCR-based restriction fragment analysis,27 whereas the >15-kb deletion was detected by Southern blotting28 (NCBI RefSeq NM_001195799.1). For each mutation, two groups comprising an equal number of carriers (2 × 14 for >15-kb deletion and 2 × 42 for c.259T>G) were formed according to their muscular tolerance to statins (Table 1). Statin tolerance was self-reported by subjects as part of a detailed questionnaire. All subjects were followed up at the Chicoutimi Hospital Lipid Clinic. A written informed consent was obtained from all participants, and all clinical data were de-identified. The Chicoutimi Hospital Ethics Committee approved this project in accordance with the Declaration of Helsinki.
Genealogical material was obtained from the BALSAC population register.29 Genealogies were reconstructed as far as the first immigrants to Quebec encountered in each genealogical branch (most branches go back to the seventeenth century). To compare the genealogical characteristics of carriers with those of the general population, control groups were also formed for each of the four subgroups (>15-kb deletion tolerant, >15-kb deletion intolerant, c.259T>G tolerant and c.259T>G intolerant). Controls were selected randomly among the available data, matched to the carriers’ parents’ dates (±5 years) and places (±10 km) of marriage. As such, LDLR genotype, FH status, lipid-lowering drug use and statin tolerance status were unknown for these controls. However, the comparison of genealogical characteristics between controls and carriers gives a good idea of the degree to which these two groups differ for each characteristic.
Genealogical analyses
All known genealogical links between the ancestors identified in the genealogies were established. Number of ancestors, geographical origins and genetic contribution of founders, inbreeding coefficients and kinship coefficients among subjects and controls were measured using the S-Plus-based GENLIB software package.29
Ancestors were counted as many times as they appeared in each genealogy. This measure gives an account of the variability of occurrences of ancestors in the genealogies. Ancestors who immigrated to Quebec (anywhere in the province) are called the immigrant founders and those who immigrated to SLSJ are called the regional founders.26, 30 As such, an ancestor can be an immigrant and also a regional founder if he/she immigrated directly to the SLSJ region from outside Quebec. The geographical origins of these founders were ascertained using information from the BALSAC database.31 The number of genealogies in which each of these founders appears at least once was calculated. Then, in order to give a more precise account of the variability of the impact of the regional and immigrant founders among each group, the genetic contribution of each founder (Gf) to each group was calculated as follows:32

where
S=all known subjects linked to the founder;
P=all known genealogical paths between the founder and the subject; and
g=number of generations, in each genealogical path, between the founder and the subject.
Gf is the sum of the probabilities of transmission of a gene from the founder to the individuals in the population.33 Dividing the result by the number of individuals gives the proportion of the population’s gene pool that is attributable to this founder (the sum of the genetic contributions of all founders is equal to 100% of the gene pool). This measure can be interpreted as a summary of the demographic events that occurred among the descendants of the founder.34
Finally, inbreeding and kinship coefficients among subjects and controls were calculated. These measures provide an account of the importance of shared ancestors in the population. The kinship coefficient between two individuals i and j (Φi,j) was calculated as follows:35

where
A=all known ancestors common to i and j;
P=all known genealogical paths between i and j, through A;
k=number of individuals in P; and
F(A)=inbreeding coefficient for A (ie, A’s parents’ kinship coefficient).
These coefficients were computed for each generation level, up to the fifteenth generation. Mean kinship and inbreeding coefficients were then calculated for each group. A permutation test was used to compare two mean kinship values. The P-values were obtained by performing 5000 permutations. A Wilcoxon’s test was used to compare the mean inbreeding coefficients.
RESULTS
Up to 225 000 ancestors were found in the genealogies, but many of these ancestors appeared more than once (Table 2). Given the structure of the SLSJ population, some remote ancestors can even appear several times in a single genealogy.36, 37 For both mutations, repeated occurrences of the same ancestors were more frequent among the carriers’ genealogies than among the controls’, but no difference was observed between tolerant and intolerant subjects. The mean genealogical depth reached over 10 generations in all the groups, with a maximum depth of 17 generations. At least 90% of the ancestors were identified up to the eighth generation (after the tenth generation, the number of known ancestors declined sharply). As observed for the number of ancestors, numbers of immigrant and regional founders did not vary greatly between the genealogies of tolerant and intolerant carriers of either mutation, except maybe for c.259T>G where the number of immigrant founders was slightly higher among the genealogies of intolerant carriers. Control genealogies of each group showed greater numbers of both types of founders, suggesting a higher genetic diversity than among carriers. In all cases, the vast majority (85%) of immigrant founders came from France, mostly during the seventeenth and eighteenth centuries (Table 3). Regional founders came mainly (79%) from the nearby region of Charlevoix (see Figure 1). Most other regional founders came from the other regions of Quebec. Almost three quarters of the regional founders settled in the SLSJ region during the second half of the nineteenth century.
Most immigrant founders were common to tolerant and intolerant groups among c.259T>G and >15-kb deletion carriers (Figure 2). Of these common immigrant founders, 42 were common to all c.259T>G carriers, and 34 were common to all >15-kb deletion carriers; they all came from France during the seventeenth century. No specific immigrant founder appeared in >6 genealogies of either group of c.259T>G or >15-kb deletion carriers. Because of the fewer number of generations between the regional founders and the subjects, a much lower proportion of regional founders were common to tolerant and intolerant carriers in each group. No regional founder appeared in all genealogies of either group. The maximum number of genealogies in which a common regional founder appeared was six for c.259T>G carriers and three for >15-kb deletion carriers (two founders in each case). No specific regional founder appeared in more than two genealogies. These results are consistent with the demographic structure of the SLSJ population.33
Figure 3 presents the genetic contributions of all common immigrant founders to the tolerant and intolerant groups for both FH mutations (the genetic contributions of specific and regional founders are not shown here, as they represent only a small proportion of carriers). In both cases, results show a strong correlation between the genetic contributions to tolerant and intolerant carriers (>15-kb deletion: r=0.965, P<0.000001; c.259T>G: r=0.996, P<0.000001). One founder had a relatively higher genetic contribution to intolerant >15-kb deletion carriers than tolerant >15-kb deletion carriers, but this founder was an ancestor of only 4 of the 15 intolerant >15-kb deletion subjects.
Figure 4 shows the cumulative kinship coefficients for each group, which were calculated up to the fifteenth generation. These coefficients are significantly higher among carriers of the mutations than among controls (>15-kb deletion tolerant: P=0.0066; >15-kb deletion: intolerant: P=0.0024; c.259T>G tolerant: P=0.0002; c.259T>G intolerant: P=0.0038). The differences between carriers of either mutation according to their tolerance to statins are much smaller and non-significant (>15-kb deletion: P=0.8732; c.259T>G: P=0.4002). Some gaps seem to exist when considering inbreeding levels (Figure 5). For the first five generations of ancestors, inbreeding coefficients were relatively low, and then a sharp increase occurred between the sixth and eleventh generations. At that higher level, parents of almost all individuals shared at least one ancestor. These remote kinship links are explained by early founders of the Quebec population (seventeenth–eighteenth centuries). Except for the first three generations where there is no inbreeding, coefficients were always slightly lower among intolerant carriers, but most differences were not significant (P>0.1200). However, inbreeding coefficients were significantly higher among intolerant c.259T>G carriers than among intolerant >15-kb deletion carriers (P=0.0194 at the thirteenth generation), which may suggest different patterns of demographic behavior for their respective ancestors.
DISCUSSION
Genealogical analysis can prove a useful tool to better understand the origins and frequency of various hereditary disorders in a population.2, 3, 38, 39, 40 It certainly helped to do so in the SLSJ population, where some rare inherited disorders have been previously described.1, 20, 36, 41 The main objective of the present study was to determine whether the genealogical characteristics of individuals affected by intolerance to statins were different from those of unaffected individuals using two FH mutations as models. To the best of our knowledge, this is the first genealogical analysis of drug intolerance expression.
On the whole, results show that statin tolerants and statin intolerants do not differ much. Clearly, both c.259T>G and >15-kb deletion mutations were introduced early in the Quebec population and their initial carriers most probably came from France. Moreover, several regional founders must have carried these mutations in the SLSJ population, as no regional founder was common to >20% of all contemporary carriers. Hence, no particular founder can be associated with tolerant or intolerant carriers of either FH mutation. Immigrant founders who may or may not have introduced these mutations have approximately the same genetic contribution to both groups. Furthermore, some early founders appear in all genealogies, which explains the fact that all SLSJ subjects (carriers, non-carriers, tolerant or intolerant to statins) are kin. Although the inbreeding coefficients of c.259T>G carriers are higher than those of the >15-kb deletion carriers, levels of kinship or inbreeding do not differ significantly between tolerant and intolerant carriers of both mutations.
Although we cannot reject the possibility of heritable factors, our results suggest that genetic susceptibility to muscular tolerance/intolerance to statins may comprise a complex mixture of rare and frequent gene variants that can interact with each other and with several non-genetic factors.9, 13, 42 Hence, if family aggregation of tolerance or intolerance to statins is influenced by genetic variants, it may not be explained by a single Mendelian factor. However, as all intolerant subjects are not affected by exactly the same muscular side effects, another hypothesis could be that the genealogical profiles of intolerant subjects may differ more significantly according to these side effects. The type of statin, which was not the same for all subjects, may be another source of variability in our samples. Hence, further analyses, specific to each type of statin and with sufficiently large numbers of different kinds of intolerant subjects could help confirm or modify our conclusions. Moreover, although our results clearly showed some significant differences between FH carriers and the general population, confirmed statin use and carrier status for controls would also improve the analysis.
References
Scriver C : Human genetics: lessons from Quebec populations. Annu Rev Genomics Hum Genet 2001; 2: 69–101.
Newman DL, Abney M, McPeek MS, Ober C, Cox NJ : The importance of genealogy in determining genetic associations with complex traits. Am J Hum Genet 2001; 69: 1146–1148.
Abney M : Identity-by-descent estimation and mapping of qualitative traits in large, complex pedigrees. Genetics 2008; 179: 1577–1590.
Endo A : The discovery and development of HMG-CoA reductase inhibitors. Atherosclerosis 2004; suppl. 5: 67–80.
Cholesterol Treatment Trialists’ (CTT) Collaboration, Baigent C, Blackwell L et al: Efficacy and safety of more intensive lowering of LDL cholesterol: a meta-analysis of data from 170000 participants in 26 randomised trials. Lancet 2010; 376: 1670–1681.
Zhang H, Plutzky J, Skentzos S et al: Discontinuation of statins in routine care settings: a cohort study. Ann Intern Med 2013; 158: 526–534.
Pasternak RC, Smith SC, Bairey-Merz CN et al: ACC/AHA/NHLBI Clinical Advisory on the Use and Safety of Statins. Circulation 2002; 106: 1024–1028.
Thompson PD, Clarkson P, Karas RH : Statin-associated myopathy. JAMA 2003; 289: 1681–1690.
Mancini GB, Baker S, Bergeron J et al: Diagnosis, prevention, and management of statin adverse effects and intolerance proceedings of a Canadian Working Group Consensus Conference. Can J Cardiol 2001; 27: 635–662.
Wilke RA, Lin DW, Roden DM et al: Identifying genetic risk factors for serious adverse drug reactions: current progress and challenges. Nat Rev Drug Discov 2007; 6: 904–916.
Vuorio A, Docherty KF, Humphries SE, Kuoppala J, Kovanen PT : Statin treatment of children with familial hypercholesterolemia—trying to balance incomplete evidence of long-term safety and clinical accountability: are we approaching a consensus? Atherosclerosis 2013; 226: 315–320.
Brunham LR, Lansberg PJ, Zhang L et al: Differential effect of the rs4149056 variant in SLCO1B1 on myopathy associated with simvastatin and atorvastatin. Pharmacogenomics J 2012; 12: 233–237.
Vladutiu GD : Genetic predisposition to statin myopathy. Curr Opin Rheumatol 2008; 20: 648–655.
Francesca Notarangelo M, Marziliano N, Antonietta Demola M et al: Genetic predisposition to atorvastatin-induced myopathy: a case report. J Clin Pharm Ther 2012; 37: 604–606.
Goldstein JL, Hobbs HH, Brown MS : Familial hypercholesterolemia; in Scriver CR, Beaudet AL, Sly WS, Valle D (eds): The Metabolic and Molecular Basis of Inherited Disease. New York, NY, USA: McGraw-Hill, 1987.
Hopkins PN, Toth PP, Ballantyne CM, Rader DJ : National Lipid Association Expert Panel on Familial Hypercholesterolemia: familial hypercholesterolemias: prevalence, genetics, diagnosis and screening recommendations from the National Lipid Association Expert Panel on Familial Hypercholesterolemia. J Clin Lipidol 2011; 5: S9–17.
Roy R, Bouchard G, Declos M : La première génération de Saguenayens: provenance, apparentement, enracinement. Cah Que Demogr 1988; 17: 113–133.
Bouchard G : Quelques Arpents d'Amérique. Population, économie, famille au Saguenay, 1838–1971. Montreal: Boreal, 1996.
Institut de la statistique du Québec http://www.stat.gouv.qc.ca/donstat/societe/demographie/dons_regnl/regional/ra_total.htm Accessed 27 March 2013.
Laberge A-M, Michaud J, Richter A et al: Population history and its impact on medical genetics in Quebec. Clin Genet 2005; 68: 287–301.
Liyanage KE, Burnett JR, Hooper AJ, van Bockxmeer FM : Familial hypercholesterolemia: epidemiology, Neolithic origins and modern geographic distribution. Crit Rev Clin Lab Sci 2011; 48: 1–18.
Gaudet D, Tremblay G, Perron P, Gagné C, Ouadahi Y, Moorjani S : L'hypercholestérolémie familiale dans l'est du Québec: un problème de santé publique? L'expérience de la clinique des maladies lipidiques de Chicoutimi. Union Med Can 1995; 124: 54–60.
Vohl M-C, Moorjani S, Roy M et al: Geographic distribution of French Canadian low-density lipoprotein receptor gene mutations in the Province of Quebec. Clin Genet 1997; 52: 1–6.
Heyer E : One founder/one gene hypothesis in a new expanding population: Saguenay (Quebec, Canada). Hum Biol 1999; 71: 99–109.
Tremblay M, Arsenault J, Heyer E : The transmission probabilities of founder genes in five regional populations of Quebec. Population-E 2003; 58: 361–380.
Lavoie EM, Tremblay M, Houde L, Vézina H : Demogenetic study of three populations within a region with strong founder effects. Community Genet 2005; 8: 152–160.
Vohl M-C, Couture P, Moorjani S et al: Rapid restriction fragment analysis for screening four point mutations of the low-density lipoprotein receptor gene in French Canadians. Hum Mutat 1995; 6: 243–246.
Ma YH, Betard C, Roy M, Davignon J, Kessling AM : Identification of a second French Canadian LDL receptor gene deletion and development of a rapid method to detect both deletions. Clin Genet 1989; 36: 219–228.
BALSAC Project: www.uqac.ca/balsac Accessed 27 March 2013.
Bherer C, Labuda D, Roy-Gagnon MH, Houde L, Tremblay M, Vézina H : Admixed ancestry and stratification of Quebec regional populations. Am J Phys Anthropol 2011; 144: 432–441.
Lavoie ÈM, Jomphe M, Tremblay M, Vézina H : Sources et procédures de reconstitutions généalogiques: RETRO, CARGO, BALSAC. Document no I-C-240, Projet BALSAC, Université du Québec à Chicoutimi, Chicoutimi, Canada 2011.
O’Brien E, Kerber RA, Jorde LB, Rogers AR : Founder effect: assessment of variation in genetic contributions among founders. Hum Biol 1994; 66: 185–204.
Heyer E : Genetic consequences of differential demographic behaviour in the Saguenay region, Quebec. Am J Phys Anthropol 1995; 98: l–11.
Heyer E, Tremblay M : Variability of the genetic contribution of Quebec population founders associated to some deleterious genes. Am J Hum Genet 1995; 56: 970–978.
Thompson EA : Pedigree Analysis in Human Genetics. Baltimore, MD, USA: Johns Hopkins University Press, 1986, p 25.
Heyer É, Tremblay M, Desjardins B : Seventeenth-century European origins of hereditary diseases in the Saguenay population (Quebec, Canada). Hum Biol 1997; 69: 209–225.
Vézina H, Tremblay M, Houde L : Mesures de l’apparentement biologique au Saguenay-Lac-St-Jean (Québec, Canada) à partir de reconstitutions généalogiques. Ann Demogr Hist 2004; 2: 67–84.
Heyer É, Zietkiewicz E, Rochowski A, Yotova V, Puymirat J, Labuda D : Phylogenetic and familial estimates of mitochondrial substitution rates: study of control region mutations in deep-rooting pedigrees. Am J Hum Genet 2001; 69: 1113–1126.
Helgason A, Yngvadóttir B, Hrafnkelsson B, Gulcher J, Stefánsson K : An Icelandic example of the impact of population structure on association studies. Nat Genet 2005; 37: 90–95.
Niazi TN, Cannon-Albright LA, Couldwell WT : Utah Population Database: a tool to study the hereditary element of nonsyndromic neurosurgical diseases. Neurosurg Focus 2010; 28: E1.
Brais B, Desjardins B, Labuda D, St-Hilaire M, Tremblay M, Vézina H : The genetics of French Canadians; in Cavalli-Sforza L, Feldman M (eds): Human Population Genetics: Evolution and Variation. The Biomedical & Life Sciences Collection. London, UK: Henry Stewart Talks Ltd, 2007.
Fiegenbaum M, da Silveira FR, Van der Sand CR et al: The role of common variants of ABCB1, CYP3A4, and CYP3A5 genes in lipid-lowering efficacy and safety of simvastatin treatment. Clin Pharmacol Ther 2005; 78: 551–558.
Acknowledgements
We thank Ève-Marie Lavoie and Michèle Jomphe (BALSAC Project) for their technical assistance and the three anonymous reviewers for their useful comments. This study was supported by the ECOGENE-21 project (CIHR TEAM Grant no. CTP-82941) and the Social Sciences and Humanities Research Council of Canada (Grant no. 410-2009–2292).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Tremblay, M., Bouhali, T., Gaudet, D. et al. Genealogical analysis as a new approach for the investigation of drug intolerance heritability. Eur J Hum Genet 22, 916–922 (2014). https://doi.org/10.1038/ejhg.2013.270
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2013.270
Keywords
This article is cited by
-
GENLIB: an R package for the analysis of genealogical data
BMC Bioinformatics (2015)