Abstract
Mitochondrial DNA (mtDNA) depletion syndromes (MDS) are severe autosomal recessive disorders associated with decreased mtDNA copy number in clinically affected tissues. The hepatocerebral form (mtDNA depletion in liver and brain) has been associated with mutations in the POLG, PEO1 (Twinkle), DGUOK and MPV17 genes, the latter encoding a mitochondrial inner membrane protein of unknown function. The aims of this study were to clarify further the clinical, biochemical, cellular and molecular genetic features associated with MDS due to MPV17 gene mutations. We identified 12 pathogenic mutations in the MPV17 gene, of which 11 are novel, in 17 patients from 12 families. All patients manifested liver disease. Poor feeding, hypoglycaemia, raised serum lactate, hypotonia and faltering growth were common presenting features. mtDNA depletion in liver was demonstrated in all seven cases where liver tissue was available. Mosaic mtDNA depletion was found in primary fibroblasts by PicoGreen staining. These results confirm that MPV17 mutations are an important cause of hepatocerebral mtDNA depletion syndrome, and provide the first demonstration of mosaic mtDNA depletion in human MPV17 mutant fibroblast cultures. We found that a severe clinical phenotype was associated with profound tissue-specific mtDNA depletion in liver, and, in some cases, mosaic mtDNA depletion in fibroblasts.
Similar content being viewed by others
INTRODUCTION
During the last decade, an increasing number of nuclear genetic defects have been identified causing mitochondrial dysfunction either through the accumulation of mitochondrial DNA (mtDNA) deletions (multiple deletions) or through a reduction in mtDNA copy number causing mtDNA depletion syndromes (MDS), the latter of which are autosomal recessive diseases characterized by a severe, tissue-specific decrease of mtDNA copy number. At least 12 nuclear genes have been found to be involved in mtDNA maintenance, including POLG, POLG2, PEO1, SLC25A4, TYMP, DGUOK, TK2, SUCLA2, SUCLG1, MPV17, OPA1 and RRM2B genes.1 MDS are caused by recessive defects in proteins directly involved in mtDNA replication or in proteins that affect the availability of deoxyribonucleoside triphosphates for mtDNA synthesis. Clinical presentations of MDS include early-onset hepatocerebral disease overlapping with Alpers-Huttenlocher syndrome (henceforth ‘Alpers’), isolated myopathy, encephalomyopathy and mitochondrial neurogastrointestinal encephalomyopathy syndrome. MDS are classified as myopathic, encephalomyopathic or hepatocerebral forms, of which the latter group has been associated with mutations in POLG, PEO1 (Twinkle), DGUOK and MPV17 genes.2
The human MPV17 gene is located on chromosome 2p21-23, comprising eight exons encoding 176 amino acids. It is expressed in human pancreas, kidney, muscle, liver, lung, placenta, brain and heart. MPV17 encodes a mitochondrial inner membrane protein, which has a so far largely unknown role in mtDNA maintenance. Human MPV17 is the orthologue of the mouse kidney disease gene, Mpv17. Loss of function has been shown to cause hepatocerebral MDS with oxidative phosphorylation failure and mtDNA depletion both in affected individuals and in Mpv17−/− mice.3 To date, MDS caused by MPV17 mutations has been reported in 32 patients with the clinical manifestations including early progressive liver failure, neurological abnormalities, hypoglycaemia and raised blood lactate.4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15 Recently, MPV17 mutations have also been associated with autosomal recessive adult-onset neuropathy and leukoencephalopathy with multiple mtDNA deletions in skeletal muscle.16 Thus, as for POLG, RRM2B and TK2, MPV17 mutations can lead to recessive MDS or recessive multiple mtDNA deletion disorders.
The aim of this study was to clarify further the clinical, biochemical, cellular and molecular features associated with MDS due to MPV17 gene mutations. We report 17 cases from 12 families with 11 novel MPV17 mutations. All the patients presented with early-onset liver disease but only some patients manifested neurological dysfunction. Furthermore, we demonstrate that primary fibroblast cultures from patients with MPV17 mutations may exhibit a mosaic mtDNA depletion.
SUBJECTS AND METHODS
We studied 70 unrelated probands with suspected hepatocerebral MDS who had been referred to Mitochondrial Diagnostic Centres at Oxford, Newcastle or London for clinical assessment, histological, biochemical and/or molecular genetic analyses. This study was approved and performed under the ethical guidelines issued by each institution for clinical studies, with written informed consent obtained for all subjects.
Molecular genetics
Total genomic DNA was isolated from blood leukocytes, skeletal muscle and liver samples by standard methods. Long-range PCR amplification of 13.8 kb mtDNA was undertaken to screen for mtDNA deletions in the available muscle and liver DNA samples, using primers previously described.17 mtDNA copy number relative to nDNA levels in muscle and/or liver DNA was estimated and compared with age-matched normal controls18, 19 by real-time quantitative PCR as described previously,20 except the nuclear probe was labelled with Vic at the 5′ end and the assays were carried out simultaneously using a PE7500 real-time PCR instrument (Applied Biosystems, Life Technologies, Carlsbad, CA, USA). mtDNA copy number <30% compared with age-matched controls was classed as mtDNA depletion, 30–50% as borderline low, and >50% as normal. The entire coding and flanking intronic regions of MPV17 were amplified by PCR from genomic DNA and sequenced by fluorescent dideoxy sequencing (Applied Biosystems Big Dye Terminator v3.1 kit) and capillary electrophoresis (Applied Biosystems 3730). Results were compared with Genbank reference sequences NM_002437.4 and NG_008075.1, and mutations described in accordance with HGVS nomenclature guidelines. MPV17 exon copy number (exons 1–8) was assessed by MLPA (kit P089-A1; MRC-Holland, Amsterdam, The Netherlands) in patients 16 and 17 (family 12) to confirm the deletion of multiple exons. Where available, parental blood DNA samples were analysed for the familial mutation(s) by sequencing of the appropriate exon, or by MLPA for family 12.
Muscle histology, histochemistry and biochemistry
Tissue samples from muscle and liver were obtained according to standard procedures. Muscle and liver histology and histochemistry were performed by standard methods. The activities of mitochondrial respiratory chain enzyme complexes were determined from skeletal muscle and liver biopsy samples and cultured skin fibroblasts, as previously described.21, 22
Characterization of the cellular mtDNA depletion phenotype by PicoGreen staining of fibroblasts
Cell cultures were analysed using the fluorescent stains PicoGreen (Molecular Probes, Eugene, OR, USA) (dsDNA intercalator) and tetramethylrhodamine methylester (TMRM; reflects mitochondrial membrane potential). Staining with PicoGreen is a simple, semi-quantitative method for visualizing mtDNA in situ within living cells as described.23 mtDNA nucleoids were further quantified using the IN Cell 1000 analyser (GE Healthcare Life Science, Little Chalfont, UK). Control and patients’ fibroblasts were stained with PicoGreen and 30 nM TMRM as above in clear Optimem media (Gibco, Life Technologies). Raw images were acquired and binarized to quantify mtDNA nucleoids per cell. Nucleoids were defined as PicoGreen punctae present in the cytoplasm and colocalizing with mitochondria.
RESULTS
Analysis of the MPV17 gene
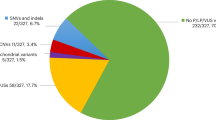
We identified pathogenic mutations in the MPV17 gene in 12 out of 70 probands screened representing 17% of our undiagnosed cohort of children with suspected hepatocerebral MDS. Ten probands harboured homozygous mutations, and two had compound heterozygous mutations. A familial homozygous mutation was also identified in further affected siblings from families 4, 5, 9 and 12, resulting in a total of 17 patients from 12 families (Table 1). Eleven of the 12 mutations we identified are previously unpublished, and as such considered to be novel: c.62T>G (p.Leu21Arg), c.67G>C (p.Ala23Pro), c.107A>C (p.Gln36Pro), c.121C>T (p.Arg41Trp), c.130C>T (p.Gln44*), c.135delA (p.Glu45Aspfs*8), c.191C>G (p.Pro64Arg), c.278A>C (p.Gln93Pro), c.279+1G>T, c.461+1G>C and a deletion including exons 3–8 of MPV17 (Figures 1 and 2).
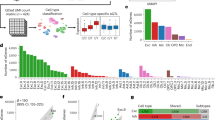
Mutations in the MPV17 gene. Schematic representation of the MPV17 gene, comprising eight exons of which exons 2–8 are coding (coding region coloured in orange); the positions of the 4 α-helical transmembrane spanning domains are indicated by red bars; missense/inframe deletion mutations are shown above the schematic and truncating/splicing mutations are shown below the schematic; novel mutations are indicated in blue, previously reported mutations also identified in this study in green and previously reported mutations not identified in this study in black.
Location of missense mutations in the MPV17 protein. Schematic representation of the MPV17 protein, which is localized to the inner mitochondrial membrane; the 4 α-helical transmembrane spanning domains are indicated by red rectangles, and the number of the first and last amino acids of each of these domains is annotated in white; novel missense mutations are indicated in blue, previously reported missense mutations also identified in this study in green and previously reported missense/inframe deletion mutations not identified in this study in grey.
Five of the 11 novel mutations are predicted to be protein truncating: c.279+1G>T and c.461+1G>C alter a consensus splice site; c.130C>T is a nonsense mutation (p.Gln44*); c.135delA results in a frameshift (p.Glu45Aspfs*8) and one mutation is a large deletion of exons 3–8. The remaining six novel mutations are missense changes, which are scattered throughout the protein (Figure 2), and alter amino acids, which are highly conserved across vertebrates (Supplementary Figure 1).
Homozygosity for the MPV17 mutation in patients from families 1, 3–5, 7–10 and 12 was confirmed by parental testing, which demonstrated all parents to be heterozygous carriers. Parental samples were not available for family 11. For patients 2 (family 2) and 9 (family 6), at least one parent was found to be a heterozygous carrier in each case, thereby confirming compound heterozygosity.
Molecular genetic studies on mtDNA from muscle and tissue samples
Molecular genetic analyses did not reveal any significant large-scale mtDNA rearrangements in any of the available muscle or liver DNA samples. mtDNA copy number was compared as previously18, 19 with controls matched for age but not ethnicity. Severe mtDNA depletion in liver was demonstrated for five out of seven cases (5–14% of control mtDNA) where liver tissue was available (patients 2, 3, 11, 14 and 16). Patient 9 had clear but less severe mtDNA depletion in liver (21% of controls), whereas mtDNA copy number was borderline low (40% of controls) in the remaining case (patient 4). Mean mtDNA copy number of all seven cases was 16%. In contrast, mtDNA copy number was highly variable across the six cases where muscle tissue was available (10–100% of controls, mean 46%; Table 1).
Clinical and laboratory features of patients with MPV17 mutations
The 17 individuals in our cohort came from 12 families, 6 being male and 11 female, from different ethnic populations including Asian, Middle-Eastern and Caucasian families. All patients manifested with liver disease with abnormal liver function tests. Poor feeding, hypoglycaemia, raised plasma lactate, hypotonia and faltering growth were common preliminary features. In four cases, the growth failure was attributed to poor feeding, two poor weight gain in the presence of adequate food intake and one growth failure. Detailed clinical summaries for each patient are provided in Supplementary File 1.
Laboratory findings revealed hypoglycaemia, and patients with liver impairment had raised plasma levels of bilirubin, transaminases, GGT, ferritin, alpha fetoprotein and coagulopathy. Plasma lactate levels that were initially raised (ranging from 3 mmol/l to 21.4 mmol/l; normal range 0.7–2.1 mmol/l) usually decreased, corresponding to an obvious clinical improvement. CSF lactate values varied from normal to 5.1 mmol/l (normal range 1.1–2.2 mmol/l). Five of the 17 patients had at least one normal blood lactate or one normal CSF lactate. Plasma amino acids were increased including methionine, tyrosine and arginine (15-fold and 10-fold above the upper limit of the normal range, respectively), and a 2–3-fold increase of glutamine, alanine, serine, glycine or threonine.
Age at presentation ranged from birth to 5 years. Five patients (4, 9, 13, 14 and 17) have survived with current ages varying from 5 months to 11.5 years (mean age 4.4 years). The median age at death was 13 months (range from 3 months to 4.3 years). Two patients (6 and 9) underwent liver transplantation. Patient 6 received a liver transplant at the age of 9 months, but died due to neurological and renal manifestations at the age of 2.5 years. Conversely, patient 9 received a liver transplant at the age of 3 years and is currently 11.5 years old manifesting with progressive demyelinating peripheral neuropathy, hypoparathyroidism and severe growth hormone deficiency. Where studied, liver ultrasound showed an enlarged echogenic liver (patients 1, 7 and 12), a distended gall bladder with biliary sludge (patients 1 and 8), nodular/heterogeneous liver parenchyma (patients 4, 5 and 8) and ascites in the abdomen and pelvis (patients 5 and 12). Brain MRI scans revealed global cerebral and cerebellar atrophy (patient 6), brain infarction (patients 6 and 7) and white matter changes consistent with metabolic leucodystrophy in five patients (patients 1, 4, 5, 6 and 10) from our cohort. Neuropathy was a feature of family 4, with corneal scarring likely due to sensory deficit, as described in Navajo familial neurogenic arthropathy.24
Histopathological features
The histological assessment of a diagnostic muscle biopsy was performed in six patients. Histopathological changes varied from normal findings (patients 2, 4 and 5) to fatty infiltration (patients 7 and 9), to a generalized fibre atrophy with small angular fibres (patient 16).
Liver histology was available from 13 of the 17 patients. Histopathological features included: fatty infiltration (patients 2, 4, 6, 7, 9, 10, 13, 14 and 16), fibrosis/cirrhosis (patients 2, 4, 5, 10 and 14), severe panlobular loss of hepatocytes and stromal collapse (patient 3), giant cell hepatitis with haemorrhagic necrosis (patient 6), non-specific changes (patient 11), cholestasis (patients 14 and 16), patchy hepatocellular oncocytosis and single-cell necrosis (patient 15), abundant mitochondria with mild pleomorphism (patient 9), marked distension of hepatocytes and few periportal glycogen deposits (patient 10) and mild iron deposition in Kupffer cells (patient 13).
Biochemical and histochemical analyses
Table 1 Muscle tissue, liver tissue and/or skin fibroblasts were available for the assessment of mitochondrial respiratory chain function from 7 out of 12 probands. Respiratory chain analysis in muscle at or soon after initial clinical presentation suggested a combined deficiency of respiratory chain complex activities in four patients (7, 9, 10 and 15), whereas the enzyme activities were normal in muscle samples from patients 2, 3 and 4 as well as in liver from patient 4 (who had the highest mtDNA copy number) and fibroblasts from patient 3. Unfortunately, biochemical analyses of liver from patients 3, 11 and 15 with mosaic mtDNA depletion in fibroblasts were not available. Sequential COX-SDH histochemistry revealed a severe, near-global COX deficiency in the liver from patient 2. Initial muscle histology at the age of 28 months from patient 9 was within normal limits, concurrent respiratory enzyme assays showing a borderline complex I deficiency. A repeat muscle biopsy from patient 9 at 3 years revealed slightly decreased respiratory chain activities of both complex I and complex IV (cytochrome c oxidase; 82% of control values). Complex IV activity was 27% of control in the explanted liver tissue. These findings are also summarized in Table 1.
Characterization of the cellular mtDNA depletion phenotype by PicoGreen staining of fibroblasts
Fibroblast cultures were available from 8 of the 17 patients in our cohort. mtDNA was visualized in these cultures using PicoGreen fluorescence microscopy and the membrane potential-dependent mitochondrial probe, TMRM. PicoGreen staining provides a quantitative measure of mtDNA content in a cell, and is more sensitive to small localized depletion than real-time PCR. mtDNA is arranged into punctate DNA/protein structures, termed nucleoids in vivo, consisting of several mtDNA genomes complexed to nucleoid proteins. Mosaic mtDNA depletion was clearly observed in fibroblasts from four MPV17 patients (patients 3, 11, 15 and 17; Figures 3c and e and Supplementary Information 2). In two patients, intermediate mtDNA depletion was observed with fibroblast mtDNA staining being generally paler (patients 4 and 8) whereas PicoGreen staining was normal in fibroblasts from patients 2 and 9. Cells showing the most marked mtDNA depletion exhibited a decreased mitochondrial membrane potential following TMRM staining. This was reflected by real-time PCR (Supplementary Information 2), which showed that the average mtDNA copy number in the cell lines with mosaic depletion was 70% (range 35–100%) of expected compared with 89% (range 43–150%) in the other patient lines (four controls, mean 100%, range 70–136%). Real-time PCR was supported by quantitation of nucleoid numbers using IN Cell 1000 analyser (Supplementary Information 2).
PicoGreen staining of fibroblast cultures. Healthy control fibroblasts exhibit typical bright punctate nucleoid staining when stained with the fluorescent DNA dye PicoGreen (a). Co-staining these cells with the potential sensitive mitochondrial stain, TMRM demonstrated a normal polarized mitochondrial network in all cells (b). Fibroblasts from patient 3 mostly exhibit typical bright nucleoids staining, but some cells (arrow) lack puncta (green in on-line version) because they are depleted of mtDNA, termed mosaic mtDNA depletion (c). Co-staining of these cells with TMRM demonstrates that some of the mtDNA-depleted cells have a depolarized mitochondrial network (d). Fibroblasts grown from patient 17 umbilical cord also show mosaic mtDNA depletion (e), most cells having normal TMRM signal (f) but one depolarized cell is shown (arrow) consistent with mosaic mtDNA depletion. Scale bars=50 μm.
DISCUSSION
We have performed clinical, biochemical, immunocytochemical and molecular genetic studies of 17 patients with MDS associated with MPV17 mutations, demonstrating a loose relationship between the clinical phenotype and mutational genotype, with patients with the most severe mtDNA depletion in liver tending to present and die at an earlier age. We have also demonstrated a mosaic mtDNA depletion in fibroblasts from some of the more severely affected patients.
Previously, MPV17 mutations have been reported in 32 patients with the hepatocerebral form of MDS in which the most common clinical manifestations included early progressive liver failure, hypoglycaemia and raised plasma lactate concentrations.4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15 Some of these patients manifested with neuromuscular abnormalities including hypotonia, nystagmus, encephalopathy, developmental delay/mental retardation, microcephaly, seizures, myoclonus, myopathy, ataxia and/or peripheral neuropathy (Navajo neurohepatopathy). White matter changes and abnormalities within the reticular formation of the lower brain stem and within the reticulospinal tracts at the cervicocranial junction on MR imaging have been reported.13, 15
Clinical findings and differential diagnoses
All our 17 patients presented with liver disease; other common features included raised serum lactate concentrations, hypotonia and faltering growth. Rarer features included endocrine abnormalities (hypoparathyroidism in patient 9, pituitary dysfunction in patient 12 and hypothyroidism in patient 15), retinal pigmentation (patients 10 and 13), corneal scarring24 (patient 5) and progressive demyelinating peripheral neuropathy (patients 4 and 9).7, 11 In addition, three patients were found with minor dysmorphic features (patients 12, 15 and 16), which have not been described previously. Thus, the clinical findings in our cohort of MDS patients with MPV17 mutations are broadly similar to those previously reported.
The infantile phenotypes of hepatocerebral forms of MDS associated with mutations in POLG, PEO1 (Twinkle), DGUOK or MPV17 have similarities, presenting with liver failure along with various neurological and endocrinological manifestations. Significant hypoglycaemia, that is glucose levels that are more difficult to maintain with oral supplementation than would be expected from the degree of liver failure, was commonly present in patients with MPV17 mutations as previously reported.4 We also observed raised levels of tyrosine in blood and/or urine in three patients, as can occur in patients with DGUOK mutations.24 We have also noted a tendency for the liver failure to precede severe neurological problems, whereas in our experience the sequence tends to be the other way round in patients with classical Alpers syndrome due to POLG mutations: in only 1 out of 24 of our series did liver dysfunction clearly precede neurological involvement.20 For this reason, patient 6 underwent a liver transplant at 15 months of age, but died 15 months later, following neurological and renal deterioration. In summary, presentations of patients with MPV17 mutations were most like those of patients with DGUOK-associated MDS, who typically have neonatal liver failure and hypotonia.25
MPV17 mutations
Twelve different mutations were identified in our cohort of 17 patients (12 probands), including 11 novel mutations (6 missense, 5 truncating) in the MPV17 gene (Figure 1). This adds significantly to the 21 previously reported mutations.4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16 The seven missense mutations (six novel and one previously reported) identified in our cohort are scattered throughout the protein (Figure 2), with no significant clustering. This is in contrast to El-Hattab et al,11 who reported clustering of missense mutations in the region of the putative protein kinase C phosphorylation site.
The majority of patients in our cohort presented in the first few weeks of life and died before 2 years of age (although three patients currently aged 5–14 months are living and so their clinical course cannot be predicted), and these patients had either truncating or missense mutations in MPV17 that presumably led to complete loss of functional protein. However, patients from families 4 and 6 (patients 4, 5, 6 and 9) appear to have a milder disease with better prognosis. These patients are/were either homozygous or compound heterozygous for missense mutations, namely p.Arg41Trp, p.Pro64Arg and p.Pro98Leu, suggesting that these mutant MPV17 proteins retain some residual function. This is similar to the findings of Karadimas et al,5 who reported patients homozygous for p.Arg50Gln or compound heterozygous for p.Gly94Arg and p.Pro98Leu, with a relatively mild form of hepatocerebral MDS, namely Navajo neurohepatopathy. It is interesting to note that both p.Arg41Trp and p.Arg50Gln are located in the same protein loop in the intermembrane space (Figure 2).
As a result of identifying pathogenic MPV17 mutations, prenatal diagnosis and/or preimplantation genetic diagnosis is now available to these 12 families. Indeed, the parents of patient 16 opted to have amniocentesis and MPV17 analysis in two subsequent pregnancies, who were accurately identified as an unaffected carrier and an affected girl (patient 17).
Mitochondrial respiratory chain analyses and mtDNA depletion in tissue samples
Respiratory chain deficiency and mtDNA depletion are usually present in the muscle of MDS patients, but not in all patients. In cases with isolated liver involvement, respiratory chain enzyme activity and mtDNA copy number have been normal in muscle and decreased only in liver tissue. For example, many patients with autosomal recessive mutations in the POLG gene causing Alpers syndrome (intractable epilepsy, hepatopathy) have normal respiratory chain enzyme activity and mtDNA copy number in skeletal muscle.20, 26 The characterization of tissues from previously described patients with MPV17 mutations has revealed a severe mtDNA depletion that was most pronounced in liver, followed by a less severe, but still significant depletion in skeletal muscle.6, 15 In our cohort, respiratory chain analysis in muscle revealed a combined deficiency of respiratory chain complexes in four out of seven muscle biopsy samples available for biochemical studies, and mtDNA depletion was identified in three out of eight muscle samples with borderline low copy number in two samples. Therefore, muscle from three out of eight patients showed no evidence of mtDNA depletion. Interestingly, there was no clear correlation between the level of mtDNA depletion in muscle, disease severity or mutation location.
As predicted, respiratory chain enzyme activities and mtDNA copy number in liver tissue correlated more closely with disease severity. Liver was available for mtDNA copy number analysis in seven patients. Severe mtDNA depletion in liver (<20% compared with age-matched controls) was associated with early onset and severe disease course (Table 1). In addition, when measured in both tissues (five patients), the mtDNA content was significantly less in the liver than in the muscle (P=0.04, one-tailed paired sample T-test). Both of these features are also apparent in patients with Alpers syndrome due to POLG mutations.20 Although liver respiratory chain enzyme activities or histochemistry were only available for three patients, this also showed some correlation with disease severity, as the only sample with normal activity was from patient 4 with the mildest disease.
PicoGreen stainings of fibroblasts
We identified mosaic mtDNA depletion in fibroblasts by PicoGreen staining in three cases (patients 3, 11 and 15) with an early-onset liver disease (birth–2.5 months), two of whom had severe mtDNA depletion in liver (5% in patient 3 and 11% in patient 11); fibroblasts grown from the umbilical cord of patient 17 (now 5 months old and deteriorating rapidly with liver failure, and whose older sibling, patient 16, was severely affected and had severe liver mtDNA depletion) also showed mosaic mtDNA depletion. In four other patients (2, 4, 8 and 9) from whom fibroblasts were available, there was no clear evidence of mosaic mtDNA depletion even after extensive passaging, and these patients had more variable onset (birth–5 years) and mtDNA copy number in liver (14–40% of controls). Muscle mtDNA copy number was highly variable both in patients with mosaic mtDNA depletion (25–100% of controls) and in patients without mosaic mtDNA depletion (46–80% of controls). Thus, mosaic mtDNA depletion was more apparent in patients with severe clinical phenotype and low mtDNA content in liver, but did not appear to correlate with muscle mtDNA content. A similar association between the cellular phenotype and severity of the clinical phenotype has previously been seen in patients with the most severe POLG mutations.20, 23 Mosaic mtDNA depletion has also been detected in serum-deprived mouse embryonic fibroblasts lacking MPV17 protein, using antiDNA antibodies,3 but has not previously been observed in patients with MPV17 mutations. Finding mosaic depletion in fibroblasts, post mortem may provide a useful clue to aetiology in cases where fibroblasts are the only material available for diagnosis.27
CONCLUSION
Our data confirm previous reports describing MPV17 mutations as an important cause of mtDNA maintenance disorders, specifically hepatocerebral MDS. We describe 17 patients with 11 novel MPV17 mutations and provide the first description of recessive MPV17 mutations associated with mosaic mtDNA depletion in fibroblasts. We present evidence that MPV17 mutations lead to tissue selective impairment of mtDNA replication and to a mosaic defect pattern in fibroblasts, which is associated at least in some patients with the severity of clinical phenotype.
References
Copeland WC : Inherited mitochondrial diseases of DNA replication. Annu Rev Med 2008; 59: 131–146.
Spinazzola A, Invernizzi F, Carrara F et al: Clinical and molecular features of mitochondrial DNA depletion syndromes. J Inherit Metab Dis 2009; 32: 143–158.
Viscomi C, Spinazzola A, Maggioni M et al: Early-onset liver mtDNA depletion and late-onset proteinuric nephropathy in Mpv17 knockout mice. Hum Mol Genet 2009; 18: 12–26.
Spinazzola A, Viscomi C, Fernandez-Vizarra E et al: MPV17 encodes an inner mitochondrial membrane protein and is mutated in infantile hepatic mitochondrial DNA depletion. Nat Genet 2006; 38: 570–575.
Karadimas CL, Vu TH, Holve SA et al: Navajo neurohepatopathy is caused by a mutation in the MPV17 gene. Am J Hum Genet 2006; 79: 544–548.
Wong LJ, Brunetti-Pierri N, Zhang Q et al: Mutations in the MPV17 gene are responsible for rapidly progressive liver failure in infancy. Hepatology 2007; 46: 1218–1227.
Navarro-Sastre A, Martín-Hernández E, Campos Y et al: Lethal hepatopathy and leukodystrophy caused by a novel mutation in MPV17 gene: description of an alternative MPV17 spliced form. Mol Genet Metab 2008; 94: 234–239.
Spinazzola A, Santer R, Akman OH et al: Hepatocerebral form of mitochondrial DNA depletion syndrome: novel MPV17 mutations. Arch Neurol 2008; 65: 1108–1113.
Kaji S, Murayama K, Nagata I et al: Fluctuating liver functions in siblings with MPV17 mutations and possible improvement associated with dietary and pharmaceutical treatments targeting respiratory chain complex II. Mol Genet Metab 2009; 97: 292–296.
Parini R, Furlan F, Notarangelo L et al: Glucose metabolism and diet-based prevention of liver dysfunction in MPV17 mutant patients. J Hepatol 2009; 50: 2105–2110.
El-Hattab AW, Li FY, Schmitt E, Zhang S, Craigen WJ, Wong LJ : MPV17-associated hepatocerebral mitochondrial DNA depletion syndrome: new patients and novel mutations. Mol Genet Metab 2010; 99: 300–308.
Navarro-Sastre A, García-Silva MT, Martín-Hernández E, Lluch M, Briones P, Ribes A : Functional splicing assay supporting that c.70+5G>A mutation in the MPV17 gene is disease causing. J Inherit Metab Dis 2010; 33: doi:10.1007/s10545-010-9155-x.
Merkle AN, Nascene DR, McKinney AM : MR imaging findings in the reticular formation in siblings with MPV17-related mitochondrial depletion syndrome. AJNR Am J Neuroradiol 2012; 33: E34–E51.
AlSaman A, Tomoum H, Invernizzi F, Zeviani M : Hepatocerebral form of mitochondrial DNA depletion syndrome due to mutation in MPV17 gene. Saudi J Gastroenterol 2012; 18: 285–289.
El-Hattab AW, Scaglia F, Craigen WJ, Wong LJC : MPV17-Related Hepatocerebral Mitochondrial DNA Depletion Syndrome; in Pagon RA, Bird TD, Dolan CR, Stephens K, Adam MP (eds): GeneReviews™ [Internet]. Seattle (WA): University of Washington 1993–2012.
Blakely EL, Butterworth A, Hadden RD et al: MPV17 mutation causes neuropathy and leukoencephalopathy with multiple mtDNA deletions in muscle. Neuromuscul Disord 2012; 22: 587–591.
Li YY, Hengstenberg C, Maisch B : Whole mitochondrial genome amplification reveals basal level multiple deletions in mtDNA of patients with dilated cardiomyopathy. Biochem Biophys Res Commun 1995; 210: 211–218.
Poulton J, Sewry C, Potter CG et al: Variation in mitochondrial DNA levels in muscle from normal controls. Is depletion of mtDNA in patients with mitochondrial myopathy a distinct clinical syndrome? J Inherit Metab Dis 1995; 18: 4–20.
Morten KJ, Ashley N, Wijburg F et al: Liver mtDNA content increases during development: a comparison of methods and the importance of age- and tissue-specific controls for the diagnosis of mtDNA depletion. Mitochondrion 2007; 7: 386–395.
Ashley N, O'Rourke A, Smith C et al: Depletion of mitochondrial DNA in fibroblast cultures from patients with POLG1 mutations is a consequence of catalytic mutations. Hum Mol Genet 2008; 17: 2496–2506.
Kirby DM, Thorburn DR, Turnbull DM, Taylor RW : Biochemical assays of respiratory chain complex activity. Methods Cell Biol 2007; 80: 93–119.
Hargreaves IP, Heales SJ, Land JM : Mitochondrial respiratory chain defects are not accompanied by an increase in the activities of lactate dehydrogenase or manganese superoxide dismutase in paediatric skeletal muscle biopsies. J Inherit Metab Dis 1999; 22: 925–931.
Ashley N, Harris D, Poulton J : Detection of mitochondrial DNA depletion in living human cells using PicoGreen staining. Exp Cell Res 2005; 303: 432–446.
Johnsen SD, Johnson PC, Stein SR : Familial sensory autonomic neuropathy with arthropathy in Navajo children. Neurology 1993; 43: 1120–1125.
Dimmock DP, Zhang Q, Dionisi-Vici C et al: Clinical and molecular features of mitochondrial DNA depletion due to mutations in deoxyguanosine kinase. Hum Mutat 2008; 29: 330–331.
Nguyen KV, Ostergaard E, Holst Ravn S et al: POLG mutations in Alpers syndrome. Neurology 2005; 65: 1493–1495.
Rahman S, Poulton J : Diagnosis of mitochondrial DNA depletion syndromes. Arch Dis Child 2009; 94: 3–5.
Acknowledgements
We thank the patients and their families for participating. This study was supported by grants from the MRC, the Wellcome Trust and the Angus Memorial Mitochondrial Fund (JP), the Foundation for Paediatric Research (JU), the Sigrid Juselius Foundation (JU), and the Academy of Finland (JU, Decision number: 138566). RWT is supported by a Wellcome Trust Strategic Award (906919). SR is supported by Great Ormond Street Hospital Children’s Charity. We also thank the UK NHS Specialist Commissioners who fund the ‘Rare Mitochondrial Disorders of Adults and Children’ Diagnostic Service (http://www.mitochondrialncg.nhs.uk) in Oxford, London and Newcastle upon Tyne.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on European Journal of Human Genetics website
Rights and permissions
This work is licensed under a Creative Commons Attribution 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by/3.0/
About this article
Cite this article
Uusimaa, J., Evans, J., Smith, C. et al. Clinical, biochemical, cellular and molecular characterization of mitochondrial DNA depletion syndrome due to novel mutations in the MPV17 gene. Eur J Hum Genet 22, 184–191 (2014). https://doi.org/10.1038/ejhg.2013.112
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2013.112
Keywords
This article is cited by
-
Genetic testing for mitochondrial disease: the United Kingdom best practice guidelines
European Journal of Human Genetics (2023)
-
Clinical and molecular basis of hepatocerebral mitochondrial DNA depletion syndrome in Japan: evaluation of outcomes after liver transplantation
Orphanet Journal of Rare Diseases (2020)
-
Clinical and molecular characterization of three patients with Hepatocerebral form of mitochondrial DNA depletion syndrome: a case series
BMC Medical Genetics (2019)
-
A monoclonal antibody raised against bacterially expressed MPV17 sequences shows peroxisomal, endosomal and lysosomal localisation in U2OS cells
BMC Research Notes (2016)
-
Mitochondrial dysfunction in liver failure requiring transplantation
Journal of Inherited Metabolic Disease (2016)