Abstract
Loss-of-function due to expansion of a CGG repeat located in the 5'UTR of the FMR1 gene is the most frequent cause of fragile X syndrome. Less than 1% of individuals with fragile X syndrome have been reported to have a partial or full deletion or point mutation of the FMR1 gene. However, whether a copy number gain of the FMR1 gene could result in certain clinical phenotypes has not been fully investigated. Here, we report the case of a child who presented with developmental delay starting at 9 months of age, fine motor and speech delay, progressive seizures since 18 months of age and hyperactivity. Molecular workup identified a de novo microduplication in the Xq27.3 region, including the FMR1 gene and the ASFMR1 gene. The expression level of the FMR1 gene in peripheral blood did not differ from that of the controls. In addition, an inherited 363-kb duplication on the chromosome 1q44 region and an inherited deletion of 168 kb on the chromosome 4p15.31 region were detected. It is not clear whether these inherited copy number variations (CNVs) also have a modifying role in the clinical phenotype of this patient.
Similar content being viewed by others
Introduction
Fragile X syndrome (FXS), caused by loss-of-function of the FMR1 gene, has been recognized as the most frequent cause of X-linked mental retardation and autism spectrum disorders.1, 2, 3, 4, 5 FXS is caused by a full mutation, an expansion of CGG trinucleotide repeats (>200 CGG repeats) in the 5'UTR of the FMR1 gene. Individuals harboring an allele with 55–200 CGG repeats are premutation carriers. The FMR1 full mutation causes a broad spectrum of symptoms, including intellectual disabilities, autism spectrum disorders, social anxiety, hyperarousal, inattention, impulsivity and hyperactivity.6, 7 Few cases presenting a clinical phenotype typical of fragile X have been reported to be caused by relative gross duplication in the FMR1 gene.8 Microduplications involving the FMR1 gene alone have not been reported. A prior report in the literature described a family with a 5.1 Mb duplication including both the FMR1 locus and FMR2 and 26 other genes, leading to a heritable syndrome of intellectual disability and short stature with hypogonadism8. Members of this family had a phenotype consisting of mild developmental delay with relative preservation of verbal skills, microcephaly and small hands, feet and testicles. These authors proposed that a functional disomy of FMR1 may contribute to the observed clinical phenotype.
Here, we report the clinical and molecular characterization of a case of a 4-year-old boy with a distinctive phenotype, who has an 86 kb de novo microduplication on Xq27.3 including only the FMR1 gene, the FMR1 antisense RNA gene (AS-FMR1), as well as two paternally inherited genomic alterations: a 363-kb duplication on 1q44 and a 168-kb deletion on 4p15.31.
Clinical description
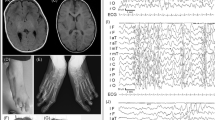
The patient first came to medical attention at around 18 months of age when he was diagnosed with myoclonic seizures, although in retrospect he had fine motor and speech delay since at least 9 months of age. The myoclonic seizures were characterized by episodes of muscle twitching that led to falls and focal activity in both frontal lobes on video EEG. The patient’s clinical symptoms improved after treatment with levetiracetam. A metabolic workup, including measurement of blood lactate, pyruvate and plasma amino acids levels, was normal. An MRI scan of the brain was also normal.
At around 2 years of age, the patient developed behavioral issues, characterized by aggressiveness toward others. He also had breath-holding spells. His neurologist noticed language delay, more expressive than receptive, suggestive of possible oral apraxia. Speech therapy was initiated, and the patient showed considerable improvement. The patient had a follow-up EEG at 3½ years of age, which showed persistent focal seizure activity despite the absence of overt clinical seizure episodes. His speech delay, although milder, persisted. It also became apparent that he was having difficulty attaining new fine motor skills, such as drawing age-appropriate geometric figures.
At 4 years of age, the patient developed absence seizures. These were described as ‘staring spells’ that were increasing in frequency over 2 months. A new video EEG confirmed the presence of absence seizures, with classical generalized 3 Hz spike-and-wave appearance. He had persistent speech and fine motor delay. He also had signs and symptoms suggestive of hyperactivity. A clinical geneticist was consulted at that time.
On physical examination, the patient’s growth parameters were within the normal range. The patient was hyperactive, running around constantly in his hospital room, and was unable to focus on a single task for more than a few seconds. He had bilateral fifth-finger clinodactyly. His strength, sensation and coordination were normal. The patient spoke very few words, and these were difficult to understand, but he had good speech comprehension. The patient was unable to draw a stick figure or a circle.
Molecular findings
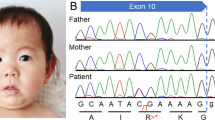
Molecular testing for fragile X syndrome and an oligonucleotide CGH microarray were carried out to investigate possible underlying genetic mechanisms. Southern blot and PCR analysis for fragile X syndrome were carried out as previously described.9, 10 The Southern blot displayed a normal male pattern and PCR analysis showed the presence of a 29 CGG repeat allele. Oligonucleotide array CGH (aCGH) was performed using a custom-designed NimbleGen 135K array developed by Signature Genomics (Spokane, WA, USA) (SignatureChipOS, version 2). The probes on the array have an average spacing of one probe every 35 kb throughout the genome and one probe every 10 kb in regions with known clinical significance. The aCGH revealed an approximately 86-kb copy number gain in chromosome Xq27.3 between nucleotides 146 783 933 and 146 868 568, which included 11 consecutive markers that were duplicated, with adjacent probe gaps of 9.89 and 2.27 kb (nucleotide coordinates were based on the UCSC Genome Browser (hg18, March 2006, NCBI build 36.1) (Figure 1).
The duplication only encompasses the FMR1 and the FMR1 antisense RNA gene (AS-FMR1). In addition to the above duplicated region, an approximately 363.16-kb copy number gain in the 1q44 region (nucleotides 245 377 586–245 556 179) and an approximately 168.86-kb copy number loss in the 4p15.31 region (nucleotides 22 897 572–23 066 434) were also detected.
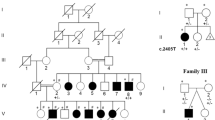
To further characterize the copy number changes found in this patient, parental studies were carried out. The copy number changes in the 1q44 and 4p15 regions were detected in the patient’s father, who does not have mental retardation, seizures or speech delay. The copy number gain in Xq27.3 was determined to be a de novo event, as it was not detected in the mother.
FMR1 mRNA expression levels were measured in peripheral blood leukocytes by quantitative RT-PCR as previously described,11 and were found to be within the normal range (1.09± 0.16).
Discussion
Mutations in the FMR1 gene have been associated with various phenotypes. These include fragile X syndrome, in which loss-of-function in FMR112 leads to intellectual disability.13 The most common mechanism leading to fragile X syndrome is a CGG trinucleotide repeat expansion in the promoter region of the FMR1 gene leading to methylation of the promoter of the gene, with subsequent silencing of transcription and the absence of the encoded product FMRP,14 which is important for synaptic plasticity and synaptic protein synthesis.15, 16 However, other loss-of-function mutations, including deletions, missense mutations and splice-site mutations, have been reported.17, 18, 19, 20
The case described here has an 86-kb microduplication that only encompasses the FMR1 and ASFMR1 genes. The ASFMR1 gene promoter is located within intron 2 of the FMR1 gene.21 Interestingly, the expression of ASFMR1, similar to that of FMR1, is elevated in fragile X premutation carriers and decreased in individuals with full mutation alleles.21 To date, translation of the ASFMR1 gene has not been reported, and whether it has a pathogenic role is not known.
Parental microarray analysis indicates that the microduplication in the FMR1 gene region in our patient is a de novo event. The gain in the 1q44 and the loss in the 4p15 regions were both inherited from a healthy father. Both copy number gains and losses of a larger chromosome 1q44 region encompassing genomic coordinates 245 141 053 to 246 395 458 had been reported in normal controls.22 The 1q44 region contains three genes (ZNF124, ZNF496 and VN1R5). ZNF124 is a transcription factor that inhibits apoptotic death in hematopoietic and leukemia cell lines23 and has been identified as part of large pathogenic deletions in 1q44,24 although patients who have a normal copy number of ZNF124 may have the same phenotype as those with a deletion. ZNF496 is a transcription cofactor of NSD1. Children with Sotos syndrome have mutations in NSD1, some of which prevent it from interacting with ZNF496.25 A third gene in our patient’s 1q44 gain, VN1R5, is thought to be a pseudogene.26 None of these genes have been reported as causative of human disease. The 4p15 loss has been reported as a normal copy number variant in the Database of Genomic Variants (http://projects.tcag.ca/variation). There are no genes in this region.
A large 5.1-Mb interstitial duplication in the Xq27.3q28 region containing 28 genes, including FMR1 and FMR2, has been reported in a French family.8 Three adult men in this family had a syndrome consisting of short stature, microcephaly, small hands and feet, low testicular volume, as well as facial features including deep-set eyes, bulbous nasal tip and thin lips. The individuals had a history of intrauterine growth restriction, failure to thrive in infancy, undescended testes requiring orchidopexy, walking after the age of 16 months and mild learning difficulties, with preserved verbal skills. The expression level of FMR1 was not measured in the French family; it is unclear whether the phenotype described was due to increased FMR1 transcript levels. X-inactivation studies suggested that female carriers preferentially inactivated the X chromosome with the interstitial duplication.
Our patient’s phenotype differs from the one described in the French family. The patient does not have any distinctive facial features; he did not have cryptorchidism, small testicular volume, nor small hands or feet. The larger duplication observed in these cases suggests that these distinctive traits identified in the French family may be caused by duplication of genes other than FMR1. Our patient has a relative milder phenotype, but he has additional features such as speech delay, fine motor delay and hyperactivity, which were not documented in the French family. However, it is difficult to compare these traits in a 4-year-old with those of the adults in the French case series.
Around 13–16% of individuals with fragile X develop seizures during their lifetime,27 which suggests a role for the FMR1 gene in epilepsy. Decreased inhibitory signaling from GABA-A receptors28 and overactivity of the metabotropic glutamate receptor (mGluR) with resulting altered AMPA receptor activity29 have been reported as pathogenic mechanisms for increased risk of seizures in fragile X syndrome. Animal models of fragile X syndrome with heterozygous knockout of the mGluR5 receptor suggest phenotypic rescue of many of the features of fragile X, including increased risk for seizures.30 These studies demonstrate a clear role for the FMR1 gene in seizures. However, whether the observed increased gene dosage of the FMR1 could also lead to the seizure activity seen in our patient is not clear.
Although the lack of FMRP causes fragile X syndrome, an increase in FMR1 mRNA, leading to RNA toxicity, results in disease phenotypes such as fragile X-associated tremor/ataxia syndrome (FXTAS)31 and premature ovarian failure.32 This may indicate the dosage-sensitive character of the FMR1 gene. However, we cannot state that excess mRNA is the cause of our patient’s phenotype, as the levels of FMR1 expression were found to be normal. This does not rule out a pathogenic microduplication, especially if it is a de novo event in which the only gene involved is known to cause disease. In addition, blood mRNA expression levels do not necessarily reflect those in the brain. CGG-binding protein sequestration has been reported to depend on the number of CGG repeats and may occur despite normal mRNA levels.33 The level of sequestration may be insufficient to cause a disease but may modulate the penetrance of a phenotype due to another genetic alteration.
It is possible that the microduplication represents a second hit that triggers the phenotype in the presence of another genomic alteration, such as the duplication on chromosome 1 or the deletion on chromosome 4 detected in this case. Indeed, copy number variation burden has been reported to positively correlate with the severity of childhood disability.34
Finally, the patient we report may have other genetic or environmental exposures to neurotoxicants that could contribute to his phenotype. The patient in the present report has a phenotype with features such as speech delay or hyperactivity, which change with time, may be age dependent and are difficult to accurately measure. Despite these limitations, we believe that it is important to share this case report with the medical genetics community.
Conclusion
A microduplication in Xq27.3 encompassing exclusively the FMR1 and ASFMR1 genes may be causative of motor and speech delay, focal seizures, absence seizures and hyperactivity, which are clinical traits frequently observed in individuals with an altered function of the FMR1 gene. Other genetic mechanisms may underlie the clinical phenotype presented here.
References
Hagerman PJ : The fragile X prevalence paradox. J Med Gen 2008; 45: 498–499.
Rogers SJ, Wehner EA, Hagerman R : The behavioral phenotype in fragile X: symptoms of autism in very young children with fragile X syndrome, idiopathic autism, and other developmental disorders. J Dev Behav Pediatr 2001; 22: 409–417.
Demark JL, Feldman MA, Holden JJ : Behavioral relationship between autism and fragile X syndrome. Am J Ment Retard 2003; 108: 314–326.
Kaufmann WE, Cortell R, Kau AS et al. Autism spectrum disorder in fragile X syndrome: communication, social interaction, and specific behaviors. Am J Med Genet A 2004; 129A: 225–234.
Hatton DD, Sideris J, Skinner M et al. Autistic behavior in children with fragile X syndrome: prevalence, stability, and the impact of FMRP. Am J Med Genet A 2006; 140A: 1804–1813.
Hagerman RJ, Hagerman PJ : The fragile X premutation: into the phenotypic fold. Curr Opin Genet Dev 2002; 12: 278–283.
Reiss AL, Dant CC : The behavioral neurogenetics of fragile X syndrome: analyzing gene-brain-behavior relationships in child developmental psychopathologies. Dev Psychopathol 2003; 15: 927–968.
Rio M, Malan V, Boissel S et al. Familial interstitial Xq27.3q28 duplication encompassing the FMR1 gene but not the MECP2 gene causes a new syndromic mental retardation condition. Eur J Hum Genet 2010; 18: 285–290.
Verkerk AJ, Pieretti M, Sutcliffe JS et al. Identification of a gene (FMR-1) containing a CGG repeat coincident with a breakpoint cluster region exhibiting length variation in fragile X syndrome. Cell 1991; 65: 905–914.
Fu YH, Kuhl DP, Pizzuti A et al. Variation of the CGG repeat at the fragile X site results in genetic instability: resolution of the Sherman paradox. Cell 1991; 67: 1047–1058.
Tassone F, Hagerman RJ, Taylor AK, Gane LW, Godfrey TE, Hagerman PJ : Elevated levels of FMR1 messenger RNA in carrier males: A new mechanism of involvement in the fragile X syndrome. Am J Hum Genet 2000; 66: 6–15.
Verkerk AJ, De Graaff E, De Boulle K et al. Alternative splicing in the fragile X gene FMR1. Hum Mol Genet 1993; 2: 399–404.
Dykens EM, Hodapp H, Ort S, Finucane B, Shapiro L, Leckman J : The trajectory of cognitive development in males with fragile X syndrome. J Am Acad Child Adolesc Psychiatry 1989; 28: 422–426.
Pietrobono R, Tabolacci E, Zalfa F et al. Molecular dissection of the events leading to inactivation of the FMR1 gene. Hum Mol Genet 2005; 14: 267–277.
Huber KM, Gallagher SM, Warren ST, Bear MF : Altered synaptic plasticity in a mouse model of fragile X mental retardation. Proc Natl Acad Sci U S A 2002; 99: 7746–7750.
Antar LN, Dictenberg JB, Plociniak M, Afroz R, Bassell GJ : Localization of FMRP-associated mRNA granules and requirement of microtubules for activity-dependent trafficking in hippocampal neurons. Genes Brain Behav 2005; 4: 350–359.
Hammond LS, Macias MM, Tarleton JC, Shashidhar Pai G : Fragile X syndrome and deletions in FMR1: new case and review of the literature. Am J Med Genet 1997; 72: 430–434.
De Boulle K, Verkerk AJ, Reyniers E et al. A point mutation in the FMR-1 gene associated with fragile X mental retardation. Nat Genet 1993; 3: 31–35.
Lugenbeel KA, Peier AM, Carson NL, Chudley AE, Nelson DL : Intragenic loss of function mutations demonstrate the primary role of FMR1 in fragile X syndrome. Nat Genet 1995; 10: 483–485.
Collins SC, Bray SM, Suhl JA et al. Identification of novel FMR1 variants by massively parallel sequencing in developmentally delayed males. Am J Med Genet Part A 2010; 152: 2512–2520.
Ladd PD, Smith LE, Rabaia NA et al. An antisense transcript spanning the CGG repeat region of FMR1 is upregulated in premutation carriers but silenced in full mutation individuals. Hum Mol Genet 2007; 16: 3174–3187.
de Smith AJ, Tsalenko A, Sampas N, Array CGH : analysis of copy number variation identifies 1284 new genes variant in healthy white males: implications for association studies of complex diseases. Hum Mol Genet 2007; 16: 2783–2794.
Kuramoto K, Uesaka T, Kimura A, Kobayashi M, Watanabe H, Katoh O : ZK7, a novel zinc finger gene, is induced by vascular endothelial growth factor and inhibits apoptotic death in hematopoietic cells. Cancer Res 2000; 60: 425–430.
Caliebe A, Kroes HY, van der Smagt JJ et al. Four patients with speech delay, seizures and variable corpus callosum thickness sharing a 0.440Mb deletion in region 1q44 containing the HNRPU gene. Eur J Med Genet 2010; 53: 179–185.
Pasillas MP, Shah M, Kamps M, NSD1 PHD : Domains bind methylated H3K4 and H3K9 using interactions disrupted by point mutations in human sotos syndrome. NSD1 PHD domains bind methylated H3K4 and H3K9 using interactions disrupted by point mutations in human Sotos syndrome. Hum Mutat 2011; 32: 292–298.
Young JM, Massa HF, Hsu L, Trask BJ : Extreme variability among mammalian V1R gene families. Genome Res 2010; 20: 10–18.
Hagerman RJ, Berry-Kravis E, Kaufmann WE : Advances in the treatment of fragile X syndrome. Pediatrics 2009; 123: 378–390.
D'Hulst C, Kooy RF : The GABA-A receptor: a novel target for treatment of fragile X? Trends Neurosci 2007; 30: 425–431.
Li J, Pelletier MR, Perez Velazquez JL, Carlen PL : Reduced cortical synaptic plasticity and GluR1 expression associated with fragile X mental retardation protein deficiency. Mol Cell Neurosci 2002; 19: 138–151.
Dölen G, Osterweil E, Rao BS et al. Correction of fragile X syndrome in mice. Neuron 2007; 56: 955–962.
Hagerman RJ, Leehey M, Heinrichs W et al. Intention tremor, parkinsonism, and generalized brain atrophy in male carriers of fragile. X. Neurology 2001; 57: 127–130.
Allingham-Hawkins DJ, Babul-Hirji R et al. Fragile X premutation is a significant risk factor for premature ovarian failure: the International Collaborative POF in Fragile X study-preliminary data. Am J Med Genet 1999; 83: 322–325.
Sellier C, Rau F, Liu Y et al. Sam68 sequestration and partial loss of function are associated with splicing alterations in FXTAS patients. EMBO J 2010; 29: 1248–1261.
Girirajan S, Brkanac Z, Coe BP et al. Relative burden of large CNVs on a range of neurodevelopmental phenotypes. PLoS Genet 2011; 7: e1002334.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Vengoechea, J., Parikh, A., Zhang, S. et al. De novo microduplication of the FMR1 gene in a patient with developmental delay, epilepsy and hyperactivity. Eur J Hum Genet 20, 1197–1200 (2012). https://doi.org/10.1038/ejhg.2012.78
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2012.78
Keywords
This article is cited by
-
Long contiguous stretches of homozygosity detected by chromosomal microarrays (CMA) in patients with neurodevelopmental disorders in the South of Brazil
BMC Medical Genomics (2019)
-
Mapping the landscape of tandem repeat variability by targeted long read single molecule sequencing in familial X-linked intellectual disability
BMC Medical Genomics (2018)
-
X-chromosomale Intelligenzminderung
Medizinische Genetik (2018)
-
Intragenic FMR1 disease-causing variants: a significant mutational mechanism leading to Fragile-X syndrome
European Journal of Human Genetics (2017)
-
Genomic analysis identifies candidate pathogenic variants in 9 of 18 patients with unexplained West syndrome
Human Genetics (2015)