Abstract
The SPINK1 gene, encoding the human pancreatic secretory trypsin inhibitor, is one of the major genes involved in predisposition to chronic pancreatitis (CP). In this study we have assessed the potential functional impact of 11 SPINK1 promoter variants by means of both luciferase reporter gene assay and electrophoretic mobility shift assay (EMSA), using human pancreatic COLO-357 cells as an expression system. The 11 promoter variants were found to be separable into three distinct categories on the basis of the reporter gene assay results viz loss-of-function, gain-of-function and functionally neutral. These findings, which were validated by EMSA, concurred with data from previous deletion studies and DNase I footprinting assays. Further, binding sites for two transcription factors, HNF1 and PTF1, were newly identified within the SPINK1 promoter by virtue of their being affected by specific variants. Combining the functional data with epidemiological data (derived by resequencing the SPINK1 promoter region in French, German and Indian CP patients and controls), then allowed us to make meaningful inferences as to each variant's likely contribution to CP. We conclude that only the three promoter variants associated with a loss-of-function (ie, −53C>T, −142T>C and −147A>G) are likely to be disease-predisposing alterations.
Similar content being viewed by others
Introduction
Chronic pancreatitis (CP) is a recurring inflammatory disease of the pancreas. Advances in molecular genetics made over the last 15 years have contributed significantly to our understanding of the pathogenesis of CP. In particular, the identification of gain-of-function missense and copy-number mutations in the cationic trypsinogen (PRSS1; MIM# 276 000) gene,1, 2 and loss-of-function variants in the pancreatic secretory trypsin inhibitor (SPINK1; MIM# 167790) and chymotrypsin C (CTRC; MIM# 601405) genes3, 4, 5 has highlighted the pivotal role of prematurely activated pancreatic trypsin in the aetiology of CP (for a recent review, see Chen and Férec6).
To date, a wide range of different types of SPINK1 mutation have been reported. Of these, the splicing (c.87+1G>A7 and c.194+2T>C3) and frameshift (c.27delC7 and c.98_99insA8) mutations, the gross deletions (c.1-320_c.55+961del1336bp9 and deletion of the entire gene10), and the p.M1T initiation codon mutation3 represent unequivocal loss-of-function mutations. Although the causative variant within the common p.N34S-containing haplotype still remains to be identified,11, 12 the rare SPINK1 missense mutations have all been experimentally shown to be deleterious.13, 14, 15 By contrast, the pathological relevance of the known SPINK1 promoter variants is largely unknown. The underlying reasons for this may be that (i) most of the SPINK1 promoter variants are rare or even unique to particular individuals, (ii) our knowledge of the regulatory elements in the SPINK1 gene promoter is still fairly rudimentary16 and (iii) there are technical difficulties inherent in the functional characterization of promoter variation that invariably need to be overcome.17 In this study we (a) carried out resequencing of the SPINK1 proximal promoter in CP patients and controls from three different populations, (b) performed a thorough functional analysis of the currently known SPINK1 promoter variants and (c) integrated the resulting data in order to assess the pathological relevance of each promoter variant.
Materials and methods
Resequencing the SPINK1 proximal promoter region
The proximal promoter region of the SPINK1 gene was resequenced in both CP patients and controls in three different (French, German and Indian) laboratories (Supplementary Methods). The 213 French patients (all diagnosed with idiopathic chronic pancreatitis (ICP) at 20 years or younger), the 439 Indian patients (292 tropical chronic pancreatitis (TCP), 147 ICP (Supplementary Methods)), and the 418 German ICP patients (Table 1) have been previously described.18, 19 This work was approved by the local ethical review committees of each collaborating institution.
SPINK1 promoter variants included in this study
A total of 11 SPINK1 promoter variants have been reported in the literature. With the exception of the −253T>C variant, which is a common polymorphism,3, 20, 21 all these promoter variants (ie, −7>G, −22C>T, −41G>A, −53C>T, −81C>T, −142T>C, −147A>G, −164G>C, −215G>A and −215G>T; Figure 1a) were included in the analysis reported here. Also included was a novel variant (viz, −170G>A) that was found during the resequencing analysis of the French patients (Table 1). All 11 promoter variants studied were numbered relative to the first nucleotide 5′ to the A of the translational initiation codon, designated –1. NG_008356.1 was used as the SPINK1 reference sequence.
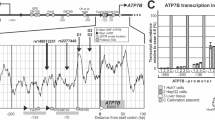
Functional characterization of SPINK1 promoter variants. (a) Partial sequence of the 5′-flanking region and beginning of the coding sequence (underlined) of the human SPINK1 gene. The first nucleotide of the translation initiation codon is numbered +1. The curved arrow indicates the transcription initiation site. The 11 SPINK1 promoter variants under study are positioned in the sequence. Vertical dotted lines delimit the promoter region that had been previously analysed by DNase I footprinting assay; the protein-protected regions I–III16 are shaded. The two downward dotted arrows delimit the fragment that was used to construct the wild-type reporter gene plasmid, pGL3-SPINK1. The double underlined sequences within region IA and region II are the HNF1 and PTF1 recognition sequences characterized in this study. (b) The pGL3-basic vector and pGL3-SPINK1 (pSPINK1) were transfected into the HEK 293 and COLO 357 cells. Firefly luciferase activity was measured and normalised against the co-transfected Renilla luciferase activity. The SPINK1 promoter-driven gene expression is shown as a fold increase compared with that of the pGL3-basic vector. (c) Expression of wild-type (Wt) pGL3-SPINK1 and variant constructs in COLO-357 cells. The effect of each variant on promoter activity is expressed as the ratio of normalised luciferase activity (against co-transfected Renilla activity) to that of the Wt construct. Averaged ratios are from three to six independent experiments. Bars, SD. *P<0.05. **P<0.001. ***P<0.0001.
Construction of reporter plasmids
A DNA fragment spanning −1171 to −1 of the SPINK1 promoter region was PCR amplified from genomic DNA of a wild-type homozygote, using the forward primer 5′-GAGCTCAGAGAAAAGGGACT-3′ containing a SacI restriction site (underlined) and the reverse primer 5′-CCATGGCTGAAGTTCTGCGT-3′ containing an NcoI restriction site (underlined). The PCR product was first subcloned into pcDNA3.1/V5-His-TOPO (Invitrogen, Cergy-Pontoise, France). The SPINK1 promoter insert was then excised from the plasmid mini-prep DNA using SacI and NcoI, and cloned into the SacI/NcoI site of pGL3-basic Reporter Vector (Promega, Charbonnieres, France). The resulting pGL3-SPINK1 wild-type construct, which contains the SPINK1 promoter insert immediately upstream of the translational initiation codon of the luciferase reporter gene, was used to create the 11 SPINK1 promoter variant reporter plasmids by site-directed mutagenesis using the Quick Change Site Directed Mutagenesis Kit (Stratagene, Massy, France). Each construct was sequence checked in order to confirm that no additional mutations had been inadvertently introduced.
Cell culture and transfection
Human pancreatic adenocarcinoma (COLO-357) and human embryonic kidney (HEK 293) cell lines were grown under conditions previously described.22 Transfections were carried out as previously described,22 using 3.8 μg pGL3-basic or pGL3-SPINK1 plus 0.2 μg control pRL-CMV vector.
Analysis of SPINK1 mRNA expression in the COLO-357 cells
Quantitative analysis of SPINK1 mRNA expression in the COLO-357 cells was carried out by quantitative RT-PCR, using the CTRC gene as a control (Supplementary Methods and Supplementary Table S1).
Luciferase reporter gene assay
At 48 h after transfection, cells were washed with phosphate-buffered saline and lysed with 180 μl passive lysis buffer (Promega). After centrifugation, 20 μl supernatant from each sample were added to 100 μl luciferase reagent and 100 μl Renilla reagent, separately. The relative luciferase activity was determined by dividing the luminescence of luciferase activity by that of the co-transfected Renilla. Three to six independent transfections (each using plasmids prepared from different clones) were carried out for each construct and luciferase assays were conducted in triplicate for each transfection. The difference between the luciferase activity of each SPINK1 promoter variant construct and the wild-type construct was then assessed for significance by means of the Student's t-test.
Electrophoretic mobility shift assay (EMSA)
5′-Biotin end-labelled probes and unlabelled specific (SpC), mutant (MuC), irrelevant (IrC) and cross-competitors (CrC; see Table 2 for definition of terms) were purchased from Eurogentec (Angers, France). Oligonucleotide sequences of the labelled probes and unlabelled SpCs and MuCs are provided in the Figures. IrC corresponds to c.136_c.165 of the human SPINK1 gene, whereas the SI CrC corresponds to a 32-bp sequence tract (ie, 5′-TAAGATATTAATAGTTCACAGCTTTGAGAAAT-3′) found in the promoter of the human sucrase-isomaltase (SI) gene that has been experimentally demonstrated to contain a binding site for hepatocyte nuclear factor 1 (HNF1).23 The human PRSS1 CrC (−138_−101 of the gene; NG_001333.2) is homologous to a pancreas-specific transcription factor 1 (PTF1) binding sequence in the rat trypsinogen gene.24
COLO-357 nuclear extracts were prepared using the NE-PER Nuclear and Cytoplasmic Extraction Reagent kit (Pierce, Brebières, France) according to the manufacturer's instructions. EMSA was visualised by means of the LightShift Mobility/Gel Shift Assay kit (Pierce). A volume of 2 μl competitor double-stranded oligonucleotide (0 to 4000 fmoles/μl) was added to a 16 μl mix containing 1 μl poly (dI•dC), 2 μl 10 × binding buffer and 3 μl nuclear extract; and incubated for 5 min at room temperature. Thereafter, 2 μl of labelled probe (10 fmoles/μl) was then added to the mixture and incubated for an additional 20 min. In the case of supershift assays, antibody was added and the reaction was further incubated for 45–60 min. DNA–protein complexes were separated by electrophoresis in a Novex DNA retardation gel (Invitrogen), transferred onto a nitrocellulose membrane, and detected by means of the Chemiluminescent Nucleic Acid Detection Module (Pierce). Anti-human RBP-L (RBPSUHL) and anti-human HNF-1α antibodies were purchased from Interchim (Montluçon, France) and Tebu-bio (Le Perray en Yvelines, France), respectively. The anti-Smad 1/5/8 antibody (Tebu-bio) was used as a control.
Results
A SPINK1 promoter fragment drives reporter gene expression in human COLO-357 cells
The human COLO-357 cell line was selected for use in the expression assays on account of its pancreatic origin and because it has been reported to express acinar digestive enzymes including trypsin, elastase and chymotrypsin.25 In this study, using quantitative RT-PCR, we demonstrated that the expression level of SPINK1 mRNA is comparable to that of CTRC mRNA in the COLO-357 cell line (Supplementary Table S2). The pGL3-SPINK1 wild-type construct, in which the 1171 bp SPINK1 promoter insert (Figure 1a) was placed immediately upstream of the luciferase reporter gene, induced a significant increase in luciferase activity in COLO 357 cells, but not in HEK 293 cells (Figure 1b).
Determining the functionality of the SPINK1 promoter variants by reporter gene assay
The 11 SPINK1 promoter variants to be tested were introduced into the pGL3-SPINK1 plasmid by site-directed mutagenesis and the resulting constructs were used to transfect COLO-357 cells. Based upon their effect on luciferase expression, they could be allocated into three distinct categories: (i) six variants (−7T>G, −22C>T, −41G>A, −164G>C, −170G>A and −215G>T), henceforth termed functionally neutral, had no appreciable effect on luciferase expression, (ii) three variants (−53C>T, −142T>C and −147A>G) caused between 20 and 46% decreases in luciferase expression (termed loss-of-function); whereas (iii) two variants (−81C>T and −215G>A) resulted in ∼30% increases in luciferase expression (termed gain-of-function) (Figure 1c).
Validating the reporter gene findings by EMSA
Of the six putatively neutral variants, −164G>C, −170G>A and −215G>T were selected for confirmatory EMSA analysis. The labelled wild-type probe containing both −164G and −170G was not found to form any specific complexes when incubated with COLO-357 nuclear extract, as none of the shifted bands were affected by an increasing molar excess of the SpC probe (Supplementary Figure 1). Thus, no evidence was obtained for either the −164G or the −170G allele binding to specific nuclear protein(s). The −215G>T variant will be discussed at the end of this section.
By contrast, the labelled wild-type probes corresponding to the five variants that manifested a functional effect upon reporter gene expression all formed specific protein–DNA complexes when incubated with COLO-357 nuclear extracts (Figures 2, 3, 4, 5b). Thus, for example, the complex (termed S1) formed between the labelled −53C probe (Figure 2a) and protein(s) in the COLO-357 nuclear extracts may be regarded as specific, as its formation was inhibited by the addition of a 25- to 200-fold molar excess of −53C SpC probe but not by a 200-fold molar excess of IrC probe (lanes 3–7, Figure 2b). The labelled −81C probe (Figure 3a) yielded four specific complexes; whereas S2c and S2d were visible with a normal (short) exposure time (lanes 2–6, Figure 3b), S2a and S2b were only visible after longer exposure (lanes 2′–6′, Figure 3b).
Functional characterization of the −53C>T promoter variant. (a) Sequences of the labelled probe, specific competitor (SpC) and mutant competitor (MuC) used for EMSA. Only the sense strand of the double-stranded oligonucleotide is shown. (b) EMSAs performed with labelled probes incubated with COLO-357 nuclear extracts in the absence or presence of SpC, MuC or irrelevant (Ir) competitor. S1 indicates the position of the specific complex detected.
Functional characterization of the −81C>T promoter variant by EMSA and identification of the cognate-binding protein. (a) Sequences of the labelled probe, specific competitor (SpC) and mutant competitor (MuC). The sequence of the labelled probe or SpC corresponds precisely to the DNase I footprinting-mapped region II (see Figure 1a). Underlined are the putative E box and TC box, components of the PTF1 bipartite recognition sequence. The canonical PTF1-binding sequence is shown within the box. (b) EMSAs performed with labelled probes incubated with COLO-357 nuclear extracts in the absence or presence of SpC, MuC or irrelevant (Ir) competitor with two exposure times. S2a, S2b, S2c and S2d indicate the four specific complexes detected. (c) EMSA performed with the human PRSS1 cross-competitor (CrC) containing a putative PTF1-binding sequence. (d) Supershift assay performed with the anti-human RBP-L antibody. A* Indicates anti-Smad 1/5/8 antibody used as control. P* indicates labelled –53C probe (see Figure 2a) used as a control. The dotted arrow in lane 5 indicates the S1 complex formed between the labelled –53C probe and nuclear extracts (see also Figure 2b).
Functional characterization of the −147A>G and −142T>C promoter variants by EMSA and identification of the cognate-binding protein. (a) Sequences of the labelled probe, specific competitor (SpC) and mutant competitors (MuCs). Underlined is a predicted HNF1 recognition sequence, of which consensus sequence is shown within the box. (b) EMSAs performed with labelled probes incubated with COLO-357 nuclear extracts in the presence or absence of SpC, MuCs or irrelevant (Ir) competitor. S3 indicates the specific complex detected. (c) EMSA performed with the human SI cross-competitor (CrC) that contains an experimentally confirmed HNF1-binding site. (d) Supershift assay carried out with the anti-human HNF1α antibody. S* denotes control antibody. In b, c and d, only the informative data are shown.
Functional characterization of the −215G>A and −215G>T promoter variants by EMSA. (a) Sequences of the labelled probe, specific competitor (SpC) and mutant competitors (MuC). (b) EMSAs performed with labelled probes incubated with COLO-357 nuclear extracts in the presence or absence of SpC, MuCs or irrelevant (Ir) competitor. S4 indicates the specific complex detected. In b, only the informative data are shown.
For each variant with a functional effect, we then compared the ability of the MuC and SpC probes to compete with the labelled probe for protein binding under the same conditions. In the cases of −53C>T (lanes 10–11, Figure 2b), −142T>C and −147A>G (lanes 10–12, Figure 4b), the MuC probes were much less efficient at inhibiting specific complex formation than the corresponding SpC probes. This indicates that all these variants have a markedly lower affinity for the cognate binding protein than for the wild-type allele.
Conversely, in the cases of −81C>T (lanes 10, 11 and 10′, 11′; Figure 3b) and −215G>A (lanes 11–12, Figure 5b), the MuC probes were found to be more efficient at inhibiting specific complex formation than the corresponding SpC probes. This indicates that in both cases, the variant alleles have a higher affinity for their cognate binding protein(s) than their wild-type counterparts.
The functionally neutral variant, −215G>T, affects the same nucleotide as the abovementioned functional −215G>A variant. However, the lack of any effect of the −215G>T variant on gene expression is readily explicable in terms of its similar binding affinity for the cognate binding protein to that displayed by its wild-type counterpart (lanes 11 and 13, Figure 5b).
Identification of two cis-acting regulatory elements in the SPINK1 gene promoter
To obtain further insight into the mechanisms underlying those variants with a demonstrable functional effect, we screened the −260 to −1 region of the SPINK1 5′-regulatory region in silico for transcription factor binding sites using MATCH (public version 1.0; http://www.gene-regulation.com/cgi-bin/pub/programs/match/bin/match.cgi?), with the parameters ‘group of matrices: vertebrates’ and ‘cut-off selection for matrix group: to minimise false positives’. The only hit obtained was a putative HNF1-binding sequence, 5′-GTCAATCATAAC-3′ (Figure 1a), located within the previously defined region IA,16 which harbours the −142T>C and −147A>G variants. In addition, for each of the 11 promoter variants, a 21-bp sequence tract, containing the variant plus 10-bp flanking sequences on either side, was screened using the same parameters, but no hits were obtained. In short, the only relevant finding to emerge from this search was that both the −142T>C and −147A>G variants were predicted to disrupt the putative HNF1-binding sequence. This was subsequently confirmed in vitro by cross-competition and supershift assays; specific complex (S3) formation between the labelled −142T/−147A probe and proteins in the COLO-357 nuclear extract was inhibited by the human SI CrC (Figure 4c) and supershifted by the anti-human HNF1α antibody (Figure 4d).
Based upon our current knowledge of promoter elements regulating expression of secretory digestive enzyme genes, we also identified, by visual inspection, a putative PTF1 bipartite sequence within region II (Figure 3a). Further, the electrophoretic profile of the four specific complexes formed between the labelled −81C probe and COLO-357 nuclear extracts (Figure 3b) was found to be consistent with that of PTF1 and its subunits.26 Hence, we surmised that S2a, S2b, S2c and S2d might correspond to the PTF1 heterotrimer, the PTF1a (also known as P48, a class B basic helix-loop-helix (bHLH) protein)/class A bHLH protein heterodimer, RBP-J (a human homologue of the Drosophila gene Suppressor of Hairless), and RBP-L (a paralogue of RBP-J), respectively. To test this hypothesis, we performed cross-competition analysis; a 400-fold molar excess of the human PRSS1 CrC probe significantly inhibited S2d formation but increased S2a formation (Figure 3c). It would thus seem that the CrC probe preferentially bound free RBP-L, thereby leaving more labelled −81C probe available for PTF1 heterotrimer binding. We also carried out a supershift assay using anti-human RBP-L antibody, which significantly reduced the intensity of the S2d complex (lane 4, Figure 3d). By contrast, the S2d-specific complex was not supershifted by a control antibody (lane 3, Figure 3d) nor was a control-specific complex supershifted by the anti-human RBP-L antibody (lanes 5 and 6, Figure 3d). Taken together, these latter findings constituted convincing evidence to support our contention that region II contains a PTF1-binding sequence.
Discussion
The foremost requirement for the appropriate functional analysis of promoter variants is the use of a physiologically relevant cell line as an expression system. Although it is known that the SPINK1 gene impacts upon the CP phenotype through its function in human pancreatic acinar cells, there are currently no such human cell lines available for in vitro experimentation. Consequently, we selected the human pancreatic adenocarcinoma cell line COLO-357 as an alternative on the basis that it is known to express pancreatic acinar enzymes such as trypsin, elastase and chymotrypsin.25 Here we have further demonstrated, by means of quantitative RT-PCR analysis, that the COLO-357 cells also express SPINK1. Indeed, the human COLO 357 cell line displayed significantly increased reporter gene expression upon transfection with the pGL3-SPINK1 wild-type construct (Figure 1b). Consequently, all further experiments were performed in COLO-357 cells. This notwithstanding, it should be pointed out that the expression level of SPINK1, as well as that of the digestive enzymes in the COLO-357 cells, is certainly much lower than that in their principal site of physiological expression, human pancreatic acinar cells. Hence, to avoid any misleading conclusions emanating from the use of COLO-357 cells as an expression system, we sought other means to validate the resulting functional data, as described below.
Cross-correlating the results of reporter gene and EMSA analyses with data from previous deletion mutagenesis and DNase I footprinting studies
We first attempted to see whether our functional data concurred with findings from previous deletion mutagenesis and DNase I footprinting studies.16 Of the 11 SPINK1 promoter variants functionally analysed here, nine occurred within the −1 to −179 region previously analysed by DNase I footprinting. A perfect correlation was noted between deduced functionality and location for these nine variants; all four functional variants occurring within this region, −53C>T, −81C>T, −142T>C and −147A>G, were located within protein-protected regions, whereas all five putatively neutral variants (viz, −7T>G, −22C>T, −41G>A, −164G>C and −170G>A) were not (Figure 1a). Consistent with this finding, EMSA revealed that although the wild-type probes corresponding to the four functional variants formed specific DNA–protein complexes when incubated with COLO-357 nuclear extract, the wild-type probes corresponding to the neutral −164G>C and −170G>A (−7T>G, −22C>T, and −41G>A were not tested) variants did not. In the following discussion, we shall focus specifically on the putative functional variants.
The −53C>T variant occurred within the DNase I footprinting-mapped region III (Figure 1a). As position −60 represents the major transcriptional initiation site,27 region III is likely to encompass the core SPINK1 promoter that serves to position RNA polymerase II at the correct site for transcriptional initiation to occur. Therefore, the negative effect of the −53C>T variant on gene expression (Figure 1c) is explicable simply in terms of the disruption of a positive regulatory element within the core promoter.
The −142T>C and −147A>G variants occurred within region IA, a previously defined positive regulatory element within the SPINK1 promoter.16 In this study, we have provided definitive evidence that these two variants caused a reduction in gene expression by disrupting the SPINK1 HNF1-binding site, a site which is well conserved among mammalian species (Supplementary Figure 2). HNF1 binds DNA as a homo or heterodimer of two peptides, HNF1α and HNF1β, both members of the homeobox family of transcription factors. Although HNF1 is known to be abundantly expressed in the liver, intestine and both exocrine and endocrine pancreatic cells,28 no HNF1-binding site has so far been described in any gene encoding a secretory digestive enzyme.
That both the −81C>T and −215G>A alleles cause a gain-of-function (Figure 1c) and have a higher affinity for their cognate binding protein(s) than their wild-type counterparts (Figures 3b and 5b), suggested that they should effect positive regulatory elements. Consistent with this prediction, the −81C>T variant occurred within the positive regulatory element in region II (Figure 1a); analysis of previously reported deletion mutagenesis data16 revealed that variant −215G>A occurs within a positively acting region spanning −245 to −180. In addition, we were able to provide compelling evidence that the −81C>T variant occurred within a PTF1-binding site in the SPINK1 gene. PTF1 has a pivotal role in the development of the pancreas and in the maintenance of the differentiated state of the adult exocrine pancreas.26, 29
In summary, the consistency observed between findings derived from two entirely independent sources is strongly suggestive of the validity of our conclusions.
Assessing the pathological relevance of SPINK1 promoter variants through the combined consideration of functional and epidemiological data
We then attempted to ascertain whether the results of the promoter variant functional assays correlated with the available epidemiological data. If we first consider the three putatively neutral polymorphisms, the −7T>G and −164G>C variants have been shown to occur at comparable allele frequencies (∼5–6%) in both patients and controls in the Brazilian population,30 whereas the −41G>A variant has been reported in both controls31 and patients32 in two US studies. By sequencing 151 healthy subjects originating from Cameroon, we identified the –41G>A variant in 44/151 (29.1%) individuals (including three homozygotes) indicating that –41G>A represents a common West-African variant (H Witt et al, unpublished data). In vitro functional analysis consistently showed that none of these three polymorphisms affect reporter gene expression (Figure 1c).
Let us now turn to the three variants (ie, −142T>C, −147A>G and −215G>A) that occurred more frequently in patients than in controls in at least two different populations (Table 1). The pathological relevance of the first two of these variants is strongly supported by the observation that they both disrupt the HNF1-binding site (Figure 4), thereby reducing SPINK1 gene expression (Figure 1c). At first sight, it would seem as if the gain-of-function exhibited by −215G>A (Figure 1c) is incompatible with the variant's presumed pathological role (Table 1). However, −215G>A is in complete linkage disequilibrium with c.194+2T>C (IVS3+2T>C), a SPINK1 mutation that has been shown experimentally to cause exon 3 skipping both in vivo33 and in vitro.11 Clearly, any influence of the −215G>A variant on SPINK1 gene expression would be completely masked in vivo by the dramatic effect of the cis-linked c.194+2T>C mutation on the mRNA splicing phenotype.
In similar vein two other variants, of which normal or elevated function initially seemed to be at odds with their presumed pathological role turned out to be explicable in terms of the presence of other allelic or non-allelic mutations in the patients concerned. First, the gain-of-function −81C>T (Figure 1c), so far reported in only a single patient,32 occurred in an individual who also harboured the CFTR (MIM# 602 421) p.F508del mutation on one allele and the CFTR 5T allele of the intron 8 polyT tract on the other. Compound heterozygotes for CFTR mutations comprising a highly deleterious mutation on one allele (such as p.F508del) and a less severe mutation on the other allele (such as intron 8 5T) have long been known to display an increased risk of ICP (reviewed by Chen and Férec6). Thus, somewhat ironically, the SPINK1 −81C>T variant may actually constitute a protective allele, the effect of which would have been counteracted by the CFTR alleles in this particular case. The second apparently anomalous case is the −215G>T variant, which was only detected in three Indian patients (Table 1). Taken together, the reporter gene and EMSA data argue against the functionality of this variant (Figures 1c and 5b), a conclusion supported by the finding that all three subjects harbouring the −215G>T variant also possessed the SPINK1 allele with the N34S-containing haplotype, the most frequent CP-predisposing factor, in trans.21 In stark contrast to these cases, neither of the loss-of-function variants, −142T>C and −147A>G, were known to reside either in cis or in trans to any reported CP-associated mutations in the SPINK1, PRSS1, CTRC or CFTR genes.
Thus, the above lines of evidence are generally supportive of the functional data. In other words, the functional data obtained from the COLO-357 cells proved to be highly informative with regard to the pathogenesis of pancreatitis that is presumed to initiate in the acinar cells of the pancreas. This makes the assessment of the pathogenic role of the remaining three promoter variants relatively straightforward. −53C>T is a rare variant detected in 2/493 patients and 1/379 controls in Germany (Table 1). As it is located within the indispensable core promoter and serves to reduce gene expression (Figure 1c), −53C>T is likely to be disease predisposing. Consistent with this conclusion, −53C>T was not known to lie either in cis or in trans to any reported CP-associated mutations in the SPINK1, PRSS1, CTRC or CFTR genes. The −22C>T variant was found in 2/40020 and 1/19031 control chromosomes but was detected in only 1/381 pancreatitis patients,32 not inconsistent with its apparent neutral effect on reporter gene expression. Finally, the functionally neutral −170G>A variant was only found in a single French patient (Table 1), which might simply be a chance finding.
Conclusions
Despite the general difficulties inherent in promoter analysis17 and the lack of any human pancreatic acinar cell lines in this specific case, we succeeded in assessing the pathogenic relevance of the 11 currently known SPINK1 promoter variants. On the basis of both functional and epidemiological data, we concluded that only three variants (ie, −53C>T, −142T>C and −147A>G) could reasonably be defined as CP-predisposing factors. This conclusion does not, however, exclude the participation of other genetic and environmental factors in causing the disease. We may nevertheless reasonably pose the question as to why a 20–40% decrease in SPINK1 gene transcription (see Figure 1c) might increase the risk for CP when transcription is upregulated 1000-fold under inflammatory conditions.34 The answer may be that only the former contributes to disease initiation, whereas the latter simply represents a late stage phenomenon. Finally, the identification of binding sites for two transcription factors, HNF1 and PTF1 within the SPINK1 promoter by virtue of their being affected by specific variants would be further strengthened by HNF1 and PTF1 knockdown experiments such as the use of HNF1 and PTF1 siRNAs.
References
Whitcomb DC, Gorry MC, Preston RA et al: Hereditary pancreatitis is caused by a mutation in the cationic trypsinogen gene. Nat Genet 1996; 14: 141–145.
Le Maréchal C, Masson E, Chen JM et al: Hereditary pancreatitis caused by triplication of the trypsinogen locus. Nat Genet 2006; 38: 1372–1374.
Witt H, Luck W, Hennies HC et al: Mutations in the gene encoding the serine protease inhibitor, Kazal type 1 are associated with chronic pancreatitis. Nat Genet 2000; 25: 213–216.
Rosendahl J, Witt H, Szmola R et al: Chymotrypsin C (CTRC) variants that diminish activity or secretion are associated with chronic pancreatitis. Nat Genet 2008; 40: 78–82.
Masson E, Chen JM, Scotet V, Le Maréchal C, Férec C : Association of rare chymotrypsinogen C (CTRC) gene variations in patients with idiopathic chronic pancreatitis. Hum Genet 2008; 123: 83–91.
Chen JM, Férec C : Chronic pancreatitis: genetics and pathogenesis. Annu Rev Genomics Hum Genet 2009; 10: 63–87.
Le Maréchal C, Chen JM, Le Gall C et al: Two novel severe mutations in the pancreatic secretory trypsin inhibitor gene (SPINK1) cause familial and/or hereditary pancreatitis. Hum Mutat 2004; 23: 205.
Gaia E, Salacone P, Gallo M et al: Germline mutations in CFTR and PSTI genes in chronic pancreatitis patients. Dig Dis Sci 2002; 47: 2416–2421.
Masson E, Le Maréchal C, Chen JM, Frebourg T, Lerebours E, Férec C : Detection of a large genomic deletion in the pancreatic secretory trypsin inhibitor (SPINK1) gene. Eur J Hum Genet 2006; 14: 1204–1208.
Masson E, Le Maréchal C, Levy P et al: Co-inheritance of a novel deletion of the entire SPINK1 gene with a CFTR missense mutation (L997F) in a family with chronic pancreatitis. Mol Genet Metab 2007; 92: 168–175.
Kereszturi E, Király O, Sahin-Tóth M : Minigene analysis of intronic variants in common SPINK1 haplotypes associated with chronic pancreatitis. Gut 2009; 58: 545–549.
Chen JM, Férec C : The true culprit within the SPINK1 p.N34S-containing haplotype is still at large. Gut 2009; 58: 478–480.
Király O, Boulling A, Witt H et al: Signal peptide variants that impair secretion of pancreatic secretory trypsin inhibitor (SPINK1) cause autosomal dominant hereditary pancreatitis. Hum Mutat 2007; 28: 469–476.
Boulling A, Le Maréchal C, Trouvé P, Raguénès O, Chen JM, Férec C : Functional analysis of pancreatitis-associated missense mutations in the pancreatic secretory trypsin inhibitor (SPINK1) gene. Eur J Hum Genet 2007; 15: 936–942.
Király O, Wartmann T, Sahin-Tóth M : Missense mutations in pancreatic secretory trypsin inhibitor (SPINK1) cause intracellular retention and degradation. Gut 2007; 56: 1433–1438.
Yasuda T, Ohmachi Y, Katsuki M et al: Identification of novel pancreas-specific regulatory sequences in the promoter region of human pancreatic secretory trypsin inhibitor gene. J Biol Chem 1998; 273: 34413–34421.
de Vooght KM, van Solinge WW : Gene promoter analysis in molecular diagnostics: do or don’t? Expert Rev Mol Diagn 2009; 9: 403–405.
Masson E, Paliwal S, Bhaskar S et al: Genetic analysis of the glycoprotein 2 gene in patients with chronic pancreatitis. Pancreas 2010; 39: 353–358.
Witt H, Rosendahl J, te Morsche RH et al: Mutational analysis of the gene encoding the zymogen granule membrane glycoprotein 2 (GP2) in patients with chronic pancreatitis. Pancreas 2010; 39: 188–192.
Chen JM, Mercier B, Audrézet MP, Férec C : Mutational analysis of the human pancreatic secretory trypsin inhibitor (PSTI) gene in hereditary and sporadic chronic pancreatitis. J Med Genet 2000; 37: 67–69.
Chandak GR, Idris MM, Reddy DN, Bhaskar S, Sriram PV, Singh L : Mutations in the pancreatic secretory trypsin inhibitor gene (PSTI/SPINK1) rather than the cationic trypsinogen gene (PRSS1) are significantly associated with tropical calcific pancreatitis. J Med Genet 2002; 39: 347–351.
Boulling A, Le Gac G, Dujardin G, Chen JM, Férec C : The c.1275A>G putative chronic pancreatitis-associated synonymous polymorphism in the glycoprotein 2 (GP2) gene decreases exon 9 inclusion. Mol Genet Metab 2010; 99: 319–324.
Rodolosse A, Carriere V, Rousset M, Lacasa M : Two HNF-1 binding sites govern the glucose repression of the human sucrase-isomaltase promoter. Biochem J 1998; 336 (part 1): 115–123.
Cockell M, Stevenson BJ, Strubin M, Hagenbuchle O, Wellauer PK : Identification of a cell-specific DNA-binding activity that interacts with a transcriptional activator of genes expressed in the acinar pancreas. Mol Cell Biol 1989; 9: 2464–2476.
Morgan RT, Woods LK, Moore GE, Quinn LA, McGavran L, Gordon SG : Human cell line (COLO 357) of metastatic pancreatic adenocarcinoma. Int J Cancer 1980; 25: 591–598.
Beres TM, Masui T, Swift GH, Shi L, Henke RM, MacDonald RJ : PTF1 is an organ-specific and Notch-independent basic helix-loop-helix complex containing the mammalian Suppressor of Hairless (RBP-J) or its paralogue, RBP-L. Mol Cell Biol 2006; 26: 117–130.
Horii A, Kobayashi T, Tomita N et al: Primary structure of human pancreatic secretory trypsin inhibitor (PSTI) gene. Biochem Biophys Res Commun 1987; 149: 635–641.
Miquerol L, Lopez S, Cartier N, Tulliez M, Raymondjean M, Kahn A : Expression of the L-type pyruvate kinase gene and the hepatocyte nuclear factor 4 transcription factor in exocrine and endocrine pancreas. J Biol Chem 1994; 269: 8944–8951.
Masui T, Swift GH, Hale MA, Meredith DM, Johnson JE, Macdonald RJ : Transcriptional autoregulation controls pancreatic Ptf1a expression during development and adulthood. Mol Cell Biol 2008; 28: 5458–5468.
Bernardino AL, Guarita DR, Mott CB et al: CFTR, PRSS1 and SPINK1 mutations in the development of pancreatitis in Brazilian patients. JOP 2003; 4: 169–177.
Pfutzer RH, Barmada MM, Brunskill AP et al: SPINK1/PSTI polymorphisms act as disease modifiers in familial and idiopathic chronic pancreatitis. Gastroenterology 2000; 119: 615–623.
Keiles S, Kammesheidt A : Identification of CFTR, PRSS1, and SPINK1 mutations in 381 patients with pancreatitis. Pancreas 2006; 33: 221–227.
Kume K, Masamune A, Kikuta K, Shimosegawa T : [-215G>A; IVS3+2T>C] mutation in the SPINK1 gene causes exon 3 skipping and loss of the trypsin binding site. Gut 2006; 55: 1214.
Khalid A, Finkelstein S, Thompson B et al: A 93 year old man with the PRSS1 R122H mutation, low SPINK1 expression, and no pancreatitis: insights into phenotypic non-penetrance. Gut 2006; 55: 728–731.
Acknowledgements
We are grateful to Kathrin Schneider (University of Heidelberg, Germany) for providing the human COLO-357 cell line. This work was supported by the INSERM (Institut National de la Santé et de la Recherche Médicale), the FRM (Fondation pour la Recherche Médicale), the APCH (Association des Pancréatites Chroniques Héréditaires), the Programme Hospitalier de Recherche Clinique (grant PHRC R 08–04), and the Association de Transfusion Sanguine et de Biogénétique Gaetan Saleun, France; the Deutsche Forschungsgemeinschaft (DFG) (Wi 2036/2-1 and 2–2), Germany; and the Council of Scientific and Industrial Research (CSIR), Ministry of Science and Technology, Government of India, India.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on European Journal of Human Genetics website
Supplementary information
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Boulling, A., Witt, H., Chandak, G. et al. Assessing the pathological relevance of SPINK1 promoter variants. Eur J Hum Genet 19, 1066–1073 (2011). https://doi.org/10.1038/ejhg.2011.79
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2011.79
Keywords
This article is cited by
-
Toward a clinical diagnostic pipeline for SPINK1 intronic variants
Human Genomics (2019)
-
Genetic Risk in Chronic Pancreatitis: The Trypsin-Dependent Pathway
Digestive Diseases and Sciences (2017)