Abstract
In several laboratories, genome-wide array analysis has been implemented as the first tier diagnostic test for the identification of copy number changes in patients with mental retardation and/or congenital anomalies. The identification of a pathogenic copy number variant (CNV) is not only important to make a proper diagnosis but also to enable the accurate estimation of the recurrence risk to family members. Upon the identification of a de novo interstitial loss or gain, the risk recurrence is considered very low. However, this risk is 50% if one of the parents is carrier of a balanced insertional translocation (IT). The apparently de novo imbalance in a patient is then the consequence of the unbalanced transmission of a derivative chromosome involved in an IT. To determine the frequency with which insertional balanced translocations would be the origin of submicroscopic imbalances, we investigated the potential presence of an IT in a consecutive series of 477 interstitial CNVs, in which the parental origin has been tested by FISH, among 14 293 patients with developmental abnormalities referred for array. We demonstrate that ITs underlie ∼2.1% of the apparently de novo, interstitial CNVs, indicating that submicroscopic ITs are at least sixfold more frequent than cytogenetically visible ITs. This risk estimate should be taken into account during counseling, and warrant parental and proband FISH testing wherever possible in patients with an apparently de novo, interstitial aberration.
Similar content being viewed by others
Introduction
Structural chromosomal rearrangements result from chromosomal breakage, followed by reconstitution in an abnormal combination. They are defined as balanced if the chromosomes have the normal chromosomal complement, or unbalanced if there is additional or missing material. Balanced, reciprocal translocations are relatively common, and were originally estimated to occur in ∼1:500 newborns,1 but more recent studies provided estimate of about 1:200.2 They require a two-break event and involve the exchange of chromosome segments between two, usually non-homologous chromosomes. Although such translocations are usually harmless for the carrier, they are associated with reduced fertility and an increased risk of unbalanced gametes and abnormal progeny. Therefore, when an unbalanced translocation is identified in a patient, it is essential to test the parents for the presence of a balanced translocation to determine the recurrence risk in the family.
Balanced insertional translocations (ITs) refer to the intercalation of a part of one chromosome into another non-homologous chromosome (interchromosomal ITs, Figure 1a) or into another part of the same chromosome (intrachromosomal insertion, Figure 2a). A balanced, interchromosomal IT is characterized by one chromosome with an interstitial deletion and another chromosome with an interstitial insertion. As there is no reciprocal segment involved in IT formation, the imbalances that result from segregation can be a pure segmental monosomy or trisomy, respectively (Figure 1b).
Insertional rearrangements imply one of the highest reproductive risk, and families usually come to the attention of a geneticist when an affected first child is born.3 For interchromosomal insertions, the unbalanced chromosome in the offspring results from simple meiotic segregation of either one of the bivalents of IT's chromosomes.4 The risk for unbalanced offspring reaches theoretically 50%. Pooled data from a number of carriers of cytogenetically visible IT families indicate the average risk of having an abnormal child for male and female carriers as 32% and 36%, respectively.4 The discrepancy between the theoretical and observed risk is due to embryonic lethality for some imbalances. It can be assumed that the survival rate would increase with decreasing insertion size, and with gains more likely to be viable than losses. Unbalanced chromosomes can also result from recombinations within the inserted segment. Recombinants require the presence of inserted chromosomal region large enough to enable the formation of a quadrivalent between both pairs of IT chromosomes. Crossingover within the insertion loop can lead to up to 26 possible different segregants.5 In contrast to interchromosomal insertions, abnormal offspring in intrachromosomal translocation carriers are associated only with recombinant chromosomes (Figure 2b).
For an IT, at least three breakpoints have to occur, and as a consequence, the prevalence of ITs is lower than for reciprocal translocations. The first estimation of the occurrence of ITs at the cytogenetic level was undertaken by Van Hemel and Eussen.4 Among 40 000 patients referred to the clinic for routine cytogenetic analysis, they identified five unbalanced rearrangements that resulted from a balanced IT in one of the parents (1:8000 patients referred to clinic). Owing to the low number of recognized aberrations, they proposed an alternative estimation of IT occurrence. By comparing the number of ITs with the number of chromosomal abnormalities associated with Down's syndrome in the same group of patients, they estimated the birth prevalence of ITs near to 1:80 000.4 Intrachromosomal insertions are considered to be even less frequent,6 but because of the scarcity of reports, the frequency remains unknown.
With the introduction of microarray analysis as the first tier diagnostic test for patients with developmental anomalies, large numbers of submicroscopic, pathogenic copy number variants (CNVs) have been uncovered. When a CNV is detected in a patient and is not shown by array in the parental genomes, the deletion or gain is considered to have occurred de novo in the patient. However, in contrast to conventional karyotyping, genomic microarrays do not reveal the structural configuration of the abnormal chromosome. Therefore, it remains possible that the imbalance is a consequence of a balanced IT in one of the parents. Despite the clinical importance for assessing the recurrence risk and a substantial number of sporadic reports of ITs7, 8, 9, 10, 11 (for a review see Van Hemel and Eussen4), the frequency of submicroscopic, balanced, ITs that may precede a pathogenic copy number change remained unknown. Recently, two groups, Kang et al12 and Neill et al,13 attempted to estimate the frequency of submicroscopic ITs. Kang et al12 presented results from 18 000 patients referred for molecular diagnostics and found 40 IT events (∼1:500). Similar results were obtained by Neill et al.13 They found 71 cases with an unbalanced insertion among ∼40 000 patients tested by array CGH (∼1:560). However, in both studies, the authors included also complex rearrangements and inherited, unbalanced insertions, which most likely represent benign structural variants.
Because of the low incidence of cytogenetically visible ITs, the potential parental carrier status is rarely investigated. In this study, we set out to determine the frequency of submicroscopic, ITs underlying apparently de novo CNVs in patients with intellectual disability/development delay (ID/DD) and/or multiple congenital anomalies (MCAs).
Methods
All patients were referred to one out of five medical centers (Center for Human Genetics, KU Leuven, Radboud University Nijmegen Medical Centre, Leiden University Medical Center, University Medical Centre Groningen and the Wessex Regional Genetics Laboratory, Salisbury District Hospital) for molecular karyotype analysis.
A total of 14 293 DNA samples were analyzed by different genome-wide array platforms; 2105 patients were analyzed with a 1-Mb resolution BAC array (Leuven), 520 patients with a tiling-resolution 32k BAC array (Nijmegen),14 1816 patients with OGT 105K oligo array (744 Leuven and 1072 Groningen), 559 patients with OGT 180K array (Leuven; CytoSure Syndrome Plus v2), 2645 patients with the ISCA design 8 × 60k oligo array (Wessex) provided by OGT (Oxford Gene Technology, Oxford, UK), 298 patients using 180K oligo from Agilent (Groningen; custom design ID: 019015; Agilent Technologies Inc., Santa Clara, CA, USA) and 6350 patients with a SNP array using the Affymetrix NspI 250k SNP array platform (4689 Nijmegen and 1661 Leiden; Affymetrix Inc., Santa Clara, CA, USA).
As the unbalanced transmission of the chromosomes involved in a balanced IT can result in either a deletion or duplication of the translocated region, both apparently de novo interstitial losses and gains, defined after parental array analysis, greater in size than 200 kb, were followed up. For deletions, FISH analysis was performed with at least one probe located within the CNV on metaphase spreads from both parents to determine whether one of the parents is carrier of a balanced IT. For gains, FISH analysis was performed on metaphase spreads of the index patient to determine whether the gain was due to the duplication in tandem, elsewhere on the same chromosome or on another chromosome. In the latter two cases, both parents were tested for the presence of an IT.
Results
Using genome-wide array, we analyzed 14 293 index patients referred for clinical diagnostics because of ID/DD and/or MCAs. In ∼60% of the patients with an interstitial, potentially pathogenic CNV, parental DNA was available for inheritance testing. A total of 771 interstitial CNVs detected in our patients were concluded to have occurred de novo based on array analysis in the parents. Among these, 144 represented recurrent, repeat-mediated CNVs (Table 1). In ∼5% of the apparently de novo unbalances, the size was lower than 200 kb; therefore, the CNV was too small to be validated by FISH. Parental peripheral blood lymphocytes were available in 62% of all these apparently de novo cases. Follow-up FISH analyses were performed in 477 cases. Among these 477 cases analyzed, 10 imbalances were identified as a consequence of the presence of an IT in one of the parents (Table 1, Figure 3). The imbalances ranged in size from 0.76 to 12.3 Mb. In two families, an intrachromosomal IT was identified, whereas in the eight other patients an interchromosomal IT was identified in one of the parents. In three cases, the mother and in seven cases the father was carrier of a balanced IT. None of the 10 cases was classified as a repeat-mediated CNV. Thus, 2.1% (∼1:50) of FISH analyzed and 0.07% (∼1:1400) of all referred to clinic patients carried apparently de novo CNVs as a consequence of the unbalanced transmission of a rearranged chromosome.
Balanced insertional translocations in nine balanced carriers. (a) FISH result with BAC clone RP11-809M12 showing translocation of 10q21.1. (b) FISH with RP11-98F19 probe showing translocation of 2q37.1. (c) FISH with RP11-679B20 showing intrachromosomal IT of region 19q13.12. (d) FISH with clone RP11-27N21 (red) showing translocated 8q21.13 and RP11-279K24 (green) indicating chromosome 5p15.2. (e) FISH with probe RP11-95M15 showing translocated region 6q23.3 (red), probe P308 (green) indicating centromere of chromosome 6. (f) FISH with RP11-13F20 (red) showing translocated region 4q22.3 and P63K5 (green) indicating chromosome 4qter. (g) FISH result with whole chromosome painting probes for chromosome 6 (green) and 11 (red). (h) FISH with FAT5 probe (red) showing translocated region 11p14.3 and centromere probe (green) indicating chromosome 11. (i) FISH with clone RP11-131P10 (red) showing translocated 13q13.3 region and probe indicating centromeres of chromosomes 13/21 (green).
Discussion
Using a combination of genome-wide array (CGH) analysis and FISH, we show that ITs underlie ∼2% of all apparent de novo CNVs. Those families are at 50% risk to have children with chromosomal imbalances. This is illustrated by the proband with a deletion on chromosome 2q, who comes from a family of six individuals with severe unexplained learning disability and dysmorphisms in two generations, who had been repeatedly investigated over a 15-year period, with no abnormality found (Figure 4). This high incidence together with the high-risk warrants parental FISH follow-up as part of the clinical routine testing in all families where apparently de novo, interstitial aberrations are detected. In addition to differences in risk counseling for carriers of a balanced IT, both prenatal and preimplantation genetic diagnosis (PGD) could be offered to reduce the risk of miscarriages and chromosomally abnormal offspring. Both PGD for intrachromosomal ITs15 as well as PGD for interchromosomal IT5, 16 has been successfully applied, and reported.
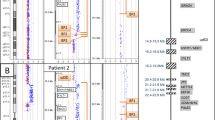
Pedigree of the proband with a der(2)ins(1;2)(p13;q36.3q37.1) pat, who comes from a family of six individuals with severe unexplained learning disability and dysmorphisms in two generations, who had been repeatedly investigated over a 15-year period, with no abnormality found. A cryptic chromosomal rearrangement had always been highly suspected but not proven until genome-wide 32k BAC array CGH analysis was available. Affected family members showed similar facies of bitemporal narrowing, prominent jaw and abnormal palmar creases. In addition to the proband, array analysis revealed an interstitial, 2.1 Mb loss in 2q36.3q37.1 in each of the three affected family members. The unaffected parent of each person carried a cryptic balanced 1:2 insertional translocation (IT) of 2.1 Mb of 2q36.3q37.1 into 1p13. Carrier testing has been carried out in nine healthy family members, and in one an asymptomatic duplication of 2q36.3q37.1 was noted. One prenatal FISH analysis was performed and also revealed a duplication of 2q36.3q37.1, but because the aunt of this unborn child was an asymptomatic carrier of the same imbalance, the parents were reassured and continued the pregnancy. Apart from stressing the importance of being able to specifically test family members who are at risk of being a carrier of such a balanced IT, this family also exemplifies how large the difference can be between the clinical consequences of a loss versus a gain of the same chromosomal region.
Overall, ITs were shown to underlie imbalances in 1:1400 of all patients referred for chromosome analysis. This result demonstrates that the frequency of unbalanced, submicroscopic, ITs among all patients referred for chromosomal evaluation is at least sixfold higher as compared with cytogenetically visible ITs.4 Considering, that in ∼40% of all cases, and in 38% of apparently de novo CNVs the parental material was not available, and in ∼5% of apparently de novo CNVs the size of the aberration does not allow follow-up FISH studies, the incidence of ITs can be extrapolated to be ∼1:900 of all patients referred to clinic or eightfold higher as compared with cytogenetically visible ITs.
Recently, two studies estimated the frequency of ITs at the submicroscopic level at ∼1:500 for patients referred to clinic.12 They detected 40 and 71 unbalanced aberrations caused by IT, out of 18 000 and 40 000 tested patients, respectively. In both reports, the overall occurrence of unbalanced IT is higher than in this study. Contrary to our results, these two studies included both de novo and inherited unbalanced ITs, whereas in this study only the de novo ITs were included. An inherited unbalanced translocation will not be detected as a de novo event and, hence, the clinical interpretation is different and needs to be individually investigated.
Carriers of a balanced IT may demonstrate a clinically abnormal phenotype because the IT events may affect gene function through disruption of the genes localized in the translocated region, genes located on the insertion site or due to position effect in abnormal gene expression in the flanking regions.12, 13 In this study, all parent carriers of an IT were phenotypically normal, and hence it can be assumed that the translocation in itself is harmless. When the translocated region contains dosage-sensitive genes, the unbalanced transmission will result in an abnormal phenotype.
In conclusion, ITs occur quite frequently in the population and underlie pathogenic CNVs in a significant fraction of patients with developmental abnormalities and/or intellectual disabilities.
References
Hook EB, Hamerton JL : The frequency of chromosome abnormalities detected in consecutive newborn studies - differences between studies - results by sex and by severity of phenotypic involvement; in Hook EB, Porter LH (eds): Population Cytogenetics. New York: Academic Press, 1977.
Jacobs PA, Browne C, Gregson N, Joyce C, White H : Estimates of the frequency of chromosome abnormalities detectable in unselected newborns using moderate levels of banding. J Med Genet 1992; 29: 103–108.
Gardner RJM, Sutherland GR : Chromosome Abnormalities and Genetic Counseling, 3rd edn. New York: Oxford University Press, 2004.
Van Hemel JO, Eussen HJ : Interchromosomal insertions. Identification of five cases and a review. Hum Genet 2000; 107: 415–432.
Melotte C, Debrock S, D’Hooghe T, Fryns JP, Vermeesch JR : Preimplantation genetic diagnosis for an insertional translocation carrier. Hum Reprod 2004; 19: 2777–2783.
Madan K, Menko FH : Intrachromosomal insertions: a case report and a review. Hum Genet 1992; 89: 1–9.
Arens YH, Engelen JJ, Govaerts LC et al: Familial insertion (3;5)(q25.3;q22.1q31.3) with deletion or duplication of chromosome region 5q22.1–5q31.3 in ten unbalanced carriers. Am J Med Genet A 2004; 130A: 128–133.
de Ravel T, Aerssens P, Vermeesch JR, Fryns JP : Trisomy of chromosome 16p13.3 due to an unbalanced insertional translocation into chromosome 22p13. Eur J Med Genet 2005; 48: 355–359.
Spreiz A, Muller D, Zotter S et al: Phenotypic variability of a deletion and duplication 6q16.1–>q21 due to a paternal balanced ins(7;6)(p15;q16.1q21). Am J Med Genet A 2010; 152A: 2762–2767.
Van Zutven LJ, van BY, Van Nieuwland CC et al: Interstitial 11q deletion derived from a maternal ins(4;11)(p14;q24.2q25): a patient report and review. Am J Med Genet A 2009; 149A: 1468–1475.
Vermeesch JR, Thoelen R, Fryns JP : A familial complex chromosome translocation resulting in duplication of 6p25. Ann Genet 2004; 47: 275–280.
Kang SH, Shaw C, Ou Z et al: Insertional translocation detected using FISH confirmation of array-comparative genomic hybridization (aCGH) results. Am J Med Genet A 2010; 152A: 1111–1126.
Neill NJ, Ballif BC, Lamb AN et al: Recurrence, submicroscopic complexity, and potential clinical relevance of copy gains detected by array CGH that are shown to be unbalanced insertions by FISH. Genome Res 2011; 21: 535–544.
de Vries BB, Pfundt R, Leisink M et al: Diagnostic genome profiling in mental retardation. Am J Hum Genet 2005; 77: 606–616.
Simopoulou M, Harper JC, Fragouli E et al: Preimplantation genetic diagnosis of chromosome abnormalities: implications from the outcome for couples with chromosomal rearrangements. Prenat Diagn 2003; 23: 652–662.
Vanneste E, Melotte C, Voet T et al: PGD for a complex chromosomal rearrangement by array comparative genomic hybridization. Hum Reprod 2011; 26: 941–949.
Acknowledgements
We are grateful to the patients and their parents, and to the clinicians of the different Clinical Genetics Departments for referring the patients. Furthermore, we thank the technicians and laboratory specialists of the different laboratories for providing the numerous array and FISH experimental data. This work has been made possible by the IWT (SBO 60848), FWO grant G.0320.07 and the KUL PFV/10/016 SymBioSys to JRV. BAN is supported by KOLUMB fellowship from the Foundation for Polish Science.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Nowakowska, B., de Leeuw, N., Ruivenkamp, C. et al. Parental insertional balanced translocations are an important cause of apparently de novo CNVs in patients with developmental anomalies. Eur J Hum Genet 20, 166–170 (2012). https://doi.org/10.1038/ejhg.2011.157
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2011.157
Keywords
This article is cited by
-
Analysis of complex chromosomal rearrangement involving chromosome 6 via the integration of optical genomic mapping and molecular cytogenetic methodologies
Journal of Human Genetics (2024)
-
Combining chromosomal microarray and clinical exome sequencing for genetic diagnosis of intellectual disability
Scientific Reports (2023)
-
Chromosomes in the genomic age. Preserving cytogenomic competence of diagnostic genome laboratories
European Journal of Human Genetics (2021)
-
13q deletion syndrome resulting from balanced chromosomal rearrangement in father: the significance of parental karyotyping
Molecular Cytogenetics (2020)
-
First interchromosomal insertion in a patient with cerebral and spinal cavernous malformations
Scientific Reports (2020)