Abstract
Grade III fetal bowel hyperechogenicity and/or loop dilatation observed at the second trimester of pregnancy can be due to several disease conditions, including cystic fibrosis (CF). Screening for frequent CF mutations is performed as a first step and, in certain situations, such as when a frequent CF mutation is found in the fetus, the increased risk of CF justifies an in-depth study of the second allele. To determine the contribution of large CFTR gene rearrangements in such cases, detected using a semiquantitative fluorescent multiplex PCR (QFM-PCR) assay, we collated data on 669 referrals related to suspicion of CF in fetuses from 1998 to 2009. Deletions were found in 5/70 cases in which QFM-PCR was applied, dele19, dele22_23, dele2_6b, dele14b_15 and dele6a_6b, of which the last three remain undescribed. In 3/5 cases, hyperechogenicity was associated with dilatation and/or gallbladder anomalies. Of the total cases of CF recognized in the subgroup of first-hand referrals, deletions represent 16.7% of CF alleles. Our study thus strengthens the need to consider large CFTR gene rearrangements in the diagnosis strategy of fetal bowel anomalies, in particular in the presence of multiple anomalies.
Similar content being viewed by others
Introduction
Fetal hyperechogenic bowel is a relatively common event detected at the second-trimester ultrasound scan in 0.04–1.80% fetuses.1, 2 It can be benign and transient, or an indicator of varied fetal pathologies including cystic fibrosis (CF) (MIM 219700), which has been reported in 2.5–10.0% of cases.2, 3, 4, 5, 6, 7
CF is the most common severe autosomal recessive disorder in the Caucasian population with an overall incidence of approximately 1 in 3500 live births.8, 9, 10 Although patients’ life expectancy has greatly improved over past decades, CF remains a severe disease, allowing prenatal diagnosis when both parents are known carriers for CF mutations or when abnormal ultrasound signs are detected during pregnancy. It is now a European recommendation that mutations in the Cystic Fibrosis Transmembrane conductance Regulator (CFTR) gene (OMIM*602421) should be investigated in fetuses with bowel anomalies such as hyperechogenic bowel and loop dilatation.11
More than 1700 CFTR variations have been reported so far (http://www.genet.sickkids.on.ca/cftr/). Most are point mutations, with variable distribution and frequency according to ethnic/geographic origin.12 Large CFTR rearrangements such as deletions, insertions or duplications, which can be identified using semiquantitative assays,13, 14, 15 account for 1–3% of CF mutations and 14–26% of unidentified alleles in CF patients.13, 16, 17 To assess the frequency of large CFTR rearrangements in ultrasound signs suggestive of CF, data of 669 referrals related to suspicion of CF in fetuses were collated. In this study we report 5 cases out of 27 CF solved using a semiquantitative fluorescent multiplex PCR (QFM-PCR) assay.
Subjects and methods
Subjects
A series of 669 cases referred from different French clinics were investigated from 1992 to October 2009 because of fetal bowel anomalies detected by routine ultrasound examination during the second or third trimester of pregnancy. Ultrasound signs were hyperechogenic bowel (sonographic density greater than that of the surrounding bone), intestinal loop dilatation, intra-abdominal calcifications, meconium peritonitis and absence of gallbladder, with each sign being isolated or combined. According to French legislation, informed consent was obtained from the parents during a genetic counseling session. Data from amniotic fluid digestive enzyme (AF-DE) activities18, 19 were collected, when available.
Samples from 450/669 couples were referred directly by clinicians to our laboratory (group 1, first-hand referrals). DNA samples from the other 219 couples were referred for further investigation after screening for frequent mutations in another laboratory (group 2).
CFTR gene analysis
Genomic DNAs were extracted from whole blood samples collected on EDTA or from amniotic fluid using varied protocols. DNA concentration and quality were determined for each sample.
Screening for point mutations
Different study levels were carried out, depending on the clinical context.11 As a first step, screening for frequent mutations was performed using diverse commercial assays or, before 1998, using denaturing gradient gel electrophoresis (DGGE)20, 21 focused on at least 10 exons. According to patients’ ethnic/geographic origins, complementary screening for other mutations was carried out using DGGE or denaturing high-performance liquid chromatography (DHPLC).22 In most cases no mutation was found and the couple was reassured. In cases in which one mutation was identified, complete scanning of the 27 CFTR coding regions and a search for large rearrangements was carried out to look for a second CF mutation.
Screening for and characterization of large CFTR rearrangements
We used the QFM-PCR assay developed and implemented in our laboratory in 2004,13, 23 retrospectively in certain cases referred before 2004. Abnormal patterns were confirmed by at least one other QFM-PCR experiment. Real-time quantitative-PCR (TaqMan) was performed in one case. Characterization of the rearrangement breakpoints was achieved by a combination of long-range PCR and sequencing experiments.
Results
In group 1 of first-hand referrals, 408/450 couples (91%) were tested for frequent mutations according to their geographic origin. Comprehensive CFTR gene studies were carried out in 42 cases, including a search for large rearrangements in 17 of them (Table 1). CF was confirmed in 9 fetuses (2.0% of 450 fetuses), including 2 with a deletion (cases 1 and 4).
In group 2 of second-hand referrals composed of 219 couples, 101 cases were referred for complementary screening depending on geographic origin, 92 because one frequent mutation was identified, 10 because of suggestive ultrasound signs and 16 because of technical problems. Search for large rearrangements by QFM-PCR was undertaken in 53 cases (Table 1). CF was confirmed in 18/219 cases (8.2%), including 3 with a large deletion (cases 2, 3 and 5).
Overall, of the 70 cases screened for gross rearrangements, a partial gene deletion was characterized in 5 cases (7.1%; Tables 2 and 3). Large CFTR deletions represent in this study 11.1% (6/54) of alleles in the 27 CF fetuses: 16.7% (3/18) in the 9 CF fetuses of group 1 and 8.3% (3/36) in the 18 CF fetuses of group 2.
Case 1
First cousins of Turkish origin were referred for fetal hyperechogenic bowel at 20 weeks of gestation.24 Screening for frequent mutations by the CF-OLA assay (PE Biosystems) was negative in both parents, whereas three peaks corresponding to exon 19 were missing in the fetus. The deletion was confirmed using real-time quantitative-PCR (TaqMan), in heterozygosity in the parents and homozygosity in the fetus. Given the late stage of gestation, the pregnancy was continued to term. The 10-year-old girl has classical CF with a positive sweat test (90 mEq/l).
Case 2
The couple was referred at 20 weeks because of fetal bowel hyperechogenicity and dilatation associated with nonvisualization of gallbladder. Screening for frequent mutations in another laboratory identified W1282X in the father and the fetus. Complementary analysis led to the identification of a new heterozygous 55 kb deletion in the mother and the fetus, encompassing exons 2–6b. An 11-bp sequence homology overlapped the inserted sequence and the 3′ breakpoint. The genotype was compatible with classical CF and, given the persistence of ultrasound signs, the parents chose to terminate the pregnancy at 26 weeks.
Case 3
The couple was referred at 30 weeks for fetal ascites, hyperechogenic bowel wall with hypoechogenic loop content, which is indicative of incipient loop dilatation, and gallbladder sludge. The father and the fetus were found F508del heterozygous in another laboratory. Abnormally high AF-DE activities were compatible with ascites, probably caused by meconial peritonitis. We identified a heterozygous previously described deletion of exons 22 and 23 in the mother and the fetus.14 Termination of pregnancy was performed and the diagnosis of CF confirmed by fetopathological examination.
Case 4
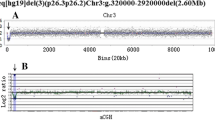
The couple was referred at 25 weeks because of nonvisualization of fetal gallbladder associated with moderate digestive dilatation and hyperechogenicity. As screening for frequent mutations according to their geographic origin was negative in both parents, the study was stopped and the pregnancy carried out to term. However, neonatal screening for CF by determination of immunoreactive trypsinemia was positive in the baby, as was the sweat test (97 mEq/l). Complete analysis of the CFTR gene led to identification of the baby as compound heterozygous for the rare 2347delG mutation, inherited from the mother, and a paternal new large deletion removing exons 14b and 15 (Figure 1).
Identification and characterization of 2751+1355_3040+243del7613 bp (dele14b_15, case 4). (a) Electropherogram from a QFM-PCR experiment in fetus 4 (blue), carrying a deletion of exons 14b and 15 at the heterozygous state (indicated by arrows), superimposed with that of a normal control (red). The deletion is evidenced by a twofold decrease in fluorescence intensity in the fetus when compared with the normal control. The x axis represents the computed length of the PCR products in base pairs as determined using an internal lane standard (green). The y axis shows fluorescent intensities in arbitrary units. Normalization of the profiles was performed using F9 (chromosome X) or DSCR1 (chromosome 21, not seen in this window) reference genes.13 Presence of double peaks at exon 9 corresponds to heterozygosity at the intron 8 poly(TG)-polyT locus. (b) The breakpoint junctions of the deletion, as determined by direct sequencing. (c) Sequence comparison and alignment between the recombined region (dele14b_15) and the wild-type sequences spanning the 5′- (IVS14a) and 3′- (IVS15) breakpoints. Identical sequences are in black and the direct repeat (GTGGG-CTG) is indicated underlined.
Case 5
The couple was referred at 32 weeks because of fetal ascites and polyhydramnios. The father and the fetus were found F508del heterozygous in another laboratory. A novel deletion of exons 6a and 6b was identified by QFM-PCR in the mother and the fetus. Further characterization showed that it retains the first 12 codons of exon 6a, with a 7 bp homology at the breakpoints in exon 6a and intron 6b. Given the late stage of pregnancy, it was continued to term and the mother gave birth to a CF baby.
Discussion
This is the first study reporting on the occurrence and frequency of large CFTR rearrangements in cases of bowel anomalies, hereby showing their notable contribution to the diagnosis of CF. The frequency of gross deletions among CF alleles found in affected fetuses (16.7% of first-hand referrals), which seems much higher than the 1.3% found in postnatally diagnosed CF patients in the same geographic population,13 makes our QFM-PCR assay the method of choice for second-line screening after testing for frequent mutations, especially in emergency cases. With the development of commercial kits such as the Multiplex ligation-dependent probes amplification kit (MLPA; MRC Holland, Amsterdam, The Netherlands) and the implementation of laboratory networks for the molecular diagnosis of CF,11 the frequency of CFTR gross rearrangements could even be re-evaluated upwards in the next few years.
We here refer to five large deletions, all of which could be detected using the MLPA assay, including three as yet undescribed: dele2_6b, dele6a_6b and dele14b_15.
As for other rearrangements described,13, 16 these deletions seem to result from non-homologous recombination, with short sequence homologies at the breakpoints. Such deletions can result from slipped mispairing during DNA replication.14 The ‘super hotspot’ CCAAR, described in 10/40 breakpoints of CFTR large rearrangements.25 was however not found in any of the five deletions reported in our study. Although the consequences of the deletions have not been studied at the functional level, it is presumed that they truncate the CFTR protein and can thus be considered to have a severe effect. Identification of deletions and consecutive confirmation of the diagnosis of CF had implications for genetic counseling in the families, as it enabled prenatal diagnosis to be offered in further pregnancies as well as cascade testing in relatives.
The clinical presentations of these five cases were representative of the signs suggestive of CF in fetuses.7 It is noteworthy that 3/5 CF fetuses carrying a deletion had associated digestive signs, all 3 with gallbladder anomalies. Few studies have been published about the association between CF and fetal gallbladder anomalies,7, 26, 27 thus calling for larger studies to assess better the frequency of CF in fetuses with such anomalies.
The present cases also further illustrate the variety of patterns of AF-DE activities. As recently evidenced in a wide retrospective study, AF-DE could still be of value for prenatal diagnosis of CF, although after 20 weeks, a physiological low value may be impossible to distinguish from a pathological one.28
In conclusion, our study strengthens the need to consider large CFTR gene rearrangements in the diagnosis strategy of fetal bowel anomalies, in particular in the presence of multiple anomalies.
References
Penna L, Bower S : Hyperechogenic bowel in the second trimester fetus: a review. Prenat Diagn 2000; 20: 909–913.
Scotet V, De Braekeleer M, Audrezet MP et al: Prenatal detection of cystic fibrosis by ultrasonography: a retrospective study of more than 346 000 pregnancies. J Med Genet 2002; 39: 443–448.
Ghose I, Mason GC, Martinez D et al: Hyperechogenic fetal bowel: a prospective analysis of sixty consecutive cases. BJOG 2000; 107: 426–429.
Corteville JE, Gray DL, Langer JC : Bowel abnormalities in the fetus--correlation of prenatal ultrasonographic findings with outcome. Am J Obstet Gynecol 1996; 175: 724–729.
Slotnick RN, Abuhamad AZ : Prognostic implications of fetal echogenic bowel. Lancet 1996; 347: 85–87.
Muller F, Dommergues M, Simon-Bouy B et al: Cystic fibrosis screening: a fetus with hyperechogenic bowel may be the index case. J Med Genet 1998; 35: 657–660.
Muller F, Simon-Bouy B, Girodon E, Monnier N, Malinge MC, Serre JL : Predicting the risk of cystic fibrosis with abnormal ultrasound signs of fetal bowel: results of a French molecular collaborative study based on 641 prospective cases. Am J Med Genet 2002; 110: 109–115.
Farrell PM : The prevalence of cystic fibrosis in the European Union. J Cyst Fibros 2008; 7: 450–453.
Southern KW, Munck A, Pollitt R et al: A survey of newborn screening for cystic fibrosis in Europe. J Cyst Fibros 2007; 6: 57–65.
Welsh MJ, Smith AE : Cystic fibrosis. Sci Am 1995; 273: 52–59.
Dequeker E, Stuhrmann M, Morris MA et al: Best practice guidelines for molecular genetic diagnosis of cystic fibrosis and CFTR-related disorders--updated European recommendations. Eur J Hum Genet 2009; 17: 51–65.
Bobadilla JL, Macek Jr M, Fine JP, Farrell PM : Cystic fibrosis: a worldwide analysis of CFTR mutations--correlation with incidence data and application to screening. Hum Mutat 2002; 19: 575–606.
Niel F, Martin J, Dastot-Le Moal F et al: Rapid detection of CFTR gene rearrangements impacts on genetic counselling in cystic fibrosis. J Med Genet 2004; 41: e118.
Audrezet MP, Chen JM, Raguenes O et al: Genomic rearrangements in the CFTR gene: extensive allelic heterogeneity and diverse mutational mechanisms. Hum Mutat 2004; 23: 343–357.
Hantash FM, Milunsky A, Wang Z et al: A large deletion in the CFTR gene in CBAVD. Genet Med 2006; 8: 93–95.
Ferec C, Casals T, Chuzhanova N et al: Gross genomic rearrangements involving deletions in the CFTR gene: characterization of six new events from a large cohort of hitherto unidentified cystic fibrosis chromosomes and meta-analysis of the underlying mechanisms. Eur J Hum Genet 2006; 14: 567–576.
Tomaiuolo R, Sangiuolo F, Bombieri C et al: Epidemiology and a novel procedure for large scale analysis of CFTR rearrangements in classic and atypical CF patients: a multicentric Italian study. J Cyst Fibros 2008; 7: 347–351.
Brock DJ, Hayward C, Gosden C : Amniotic fluid GGTP in prenatal diagnosis of cystic fibrosis: a word of warning. Lancet 1983; 1: 1099.
Mulivor RA, Cook D, Muller F et al: Analysis of fetal intestinal enzymes in amniotic fluid for the prenatal diagnosis of cystic fibrosis. Am J Hum Genet 1987; 40: 131–146.
Fanen P, Ghanem N, Vidaud M et al: Molecular characterization of cystic fibrosis: 16 novel mutations identified by analysis of the whole cystic fibrosis conductance transmembrane regulator (CFTR) coding regions and splice site junctions. Genomics 1992; 13: 770–776.
Costes B, Girodon E, Ghanem N et al: Frequent occurrence of the CFTR intron 8 (TG)n 5T allele in men with congenital bilateral absence of the vas deferens. Eur J Hum Genet 1995; 3: 285–293.
Le Marechal C, Audrezet MP, Quere I, Raguenes O, Langonne S, Ferec C : Complete and rapid scanning of the cystic fibrosis transmembrane conductance regulator (CFTR) gene by denaturing high-performance liquid chromatography (D-HPLC): major implications for genetic counselling. Hum Genet 2001; 108: 290–298.
Ratbi I, Legendre M, Niel F et al: Detection of cystic fibrosis transmembrane conductance regulator (CFTR) gene rearrangements enriches the mutation spectrum in congenital bilateral absence of the vas deferens and impacts on genetic counselling. Hum Reprod 2007; 22: 1285–1291.
Costes B, Girodon E, Vidaud D et al: Prenatal detection by real-time quantitative PCR and characterization of a new CFTR deletion, 3600+15kbdel5.3 kb (or CFTRdele19). Clin Chem 2000; 46: 1417–1420.
Quemener S, Chen JM, Chuzhanova N et al: Complete ascertainment of intragenic copy number mutations (CNMs) in the CFTR gene and its implications for CNM formation at other autosomal loci. Hum Mutat 31: 421–428.
Duchatel F, Muller F, Oury JF, Mennesson B, Boue J, Boue A : Prenatal diagnosis of cystic fibrosis: ultrasonography of the gallbladder at 17–19 weeks of gestation. Fetal Diagn Ther 1993; 8: 28–36.
Ochshorn Y, Rosner G, Barel D, Bronshtein M, Muller F, Yaron Y : Clinical evaluation of isolated nonvisualized fetal gallbladder. Prenat Diagn 2007; 27: 699–703.
Oca F, Dreux S, Gerard B et al: Amniotic fluid digestive enzyme analysis is useful for identifying CFTR gene mutations of unclear significance. Clin Chem 2009; 55: 2214–2217.
Acknowledgements
We thank Drs C Philippe, D Feldmann, JP Bonnefont, J Kaplan, A Martin and ML Dalphin for referring the DNA samples. B Boissier, R Medina, C Gameiro and F Rossi are thanked for valuable technical assistance. D Kerridge is acknowledged for re-reading the paper.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
de Becdelièvre, A., Costa, C., LeFloch, A. et al. Notable contribution of large CFTR gene rearrangements to the diagnosis of cystic fibrosis in fetuses with bowel anomalies. Eur J Hum Genet 18, 1166–1169 (2010). https://doi.org/10.1038/ejhg.2010.80
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2010.80