Abstract
Large-scale population studies have established that genetic factors contribute to individual differences in smoking behavior. Linkage and genome-wide association studies have shown many chromosomal regions and genes associated with different smoking behaviors. One study was the association of single-nucleotide polymorphisms (SNPs) in the CHRNA5-A3-B4 gene cluster to nicotine addiction. Here, we report a replication of this association in the Italian population represented by three genetically isolated populations. One, the Val Borbera, is a genetic isolate from North-Western Italy; the Cilento population, is located in South-Western Italy; and the Carlantino village is located in South-Eastern Italy. Owing to their position and their isolation, the three populations have a different environment, different history and genetic structure. The variant A of the rs1051730 SNP was significantly associated with smoking quantity in two populations, Val Borbera and Cilento, no association was found in Carlantino population probably because difference in LD pattern in the variant region.
Similar content being viewed by others
Introduction
Cigarette consumption is a major health problem worldwide. It is the cause of about 5 million deaths each year and this number is estimated to double by 2020 (World Health Organization: Economics of Tobacco Control1; World Health Organization: World Health Statistics 20082). Tobacco use is a strong risk factor for leading causes of death in the world: a significant proportion of ischemic heart disease, cerebrovascular disease, respiratory infections, tuberculosis, chronic obstructive pulmonary disease and cancers of the lung, trachea and bronchus are attributable to smoking behavior. Despite its recognized deleterious impact on health, smoking prevalence is very high in many countries with about 22% of the worldwide adult population currently consuming tobacco. Many studies have revealed that genetic factors, together with environmental factors, have a strong impact on smoking-related behavior. A meta-analysis of different twin studies estimated a mean heritability of 0.5 for smoking initiation and of 0.59 for nicotine dependence.3 Identification of genetic factors affecting smoking-related behavior is consequently a major interest of medicine today, to assess individual vulnerability and to develop more effective intervention.
Recently, several genome-wide association (GWA) studies have established the association of one locus on the chromosome 15q25 with nicotine dependence and smoking quantity. This locus implicates a cluster of three genes, CHRNA5, CHRNA3 and CHRNB4, which encode neuronal nicotinic acetylcholine receptor subunits (-5, -3 and -4), acting as risk factors for nicotine dependence or heavy smoking.3, 4, 5, 6, 7, 8, 9, 10 The same variants were also found to be associated with lung cancer and peripheral arterial diseases.5, 11, 12 However, it is still debated whether there is a direct effect on disease etiology or an indirect effect through smoking.
So far, different variants in the 15q25.1 region, strongly correlated by linkage disequilibrium, were reported as associated with smoking behavior. One of them caused an amino acid change in the CHRNA5-encoded protein but its functional role is still unclear. One study also showed evidence for the presence of at least two independent variants in the CHRNA5–CHRNA3–CHRNB4 gene cluster region contributing to nicotine dependence.6 In another study, this nicotine dependence locus has been tagged by the single-nucleotide polymorphism (SNP) rs1051730, located in the CHRNA3 gene, which has been reported to have an effect on the number of cigarettes smoked per day (CPD).5 This variant has already been associated with nicotine dependence13 in a previous genome-wide study that used the low-quantity smokers as controls.
The main focus of our analysis was to evaluate the influence of the rs1051730 variant on smoking behavior, smoking quantity within current and former smokers and related diseases in three Italian isolated populations, belonging to the Italian Network on Genetic Isolates (http://www.netgene.it/ita/ingi_6.asp).
Materials and methods
Study participants and data collection
Individuals enrolled in the study belong to three isolated populations from different parts of Italy; the basic characteristics of the three populations have been previously described.14 All individuals recruited gave their written informed consent and the study protocol was approved by the ethical committees of the institutions involved. All participants provided a detailed medical and lifestyle history collected through structured questionnaires. The information about the smoking behavior and smoking quantity was acquired using a specific questionnaire. With regard to the smoking habit, three categories were available: never, former and current smokers. According to the number of CPD the individuals were clustered in two categories: low-dose smokers (10 or fewer cigarettes per day) or high-dose smokers (>10 cigarettes per day). A dichotomous trait was derived from the CPD categories and it was used for association analysis: the subjects belonging to the low-dose category were classified as controls and those from the high-dose categories as cases.
For the Val Borbera population, information was available for 1007 never smokers, 424 former smokers and 283 current smokers. In this sample, all smoking individuals (current and former smokers) were used for clustering in the two CPD categories: 304 individuals were classified as low-dose smokers and 402 people as high-dose smokers. In the Cilento population, we had 861 never smokers, 211 former smokers and 254 current smokers. However, for this sample the CPD information was available only for current smokers generating two CPD categories with 89 light and 165 heavy smokers. In the Carlantino population, no information was available to distinguish between current and former smokers (=304) then the CPD categories included 191 heavy smokers and 113 light smokers, the total number of never smokers was 806.
Genotyping
Genotypes for SNP rs1051730 were determined by TaqMan genotyping (Applied Biosystems, cat. no. 4351379, assay C___9510307_20). Genotype calls were done with the SDS 1.08 software (Applied Biosystems, Carlsbad, CA, USA). Call rates were 98.4, 93.8 and 89.4% for the Val Borbera (1713/1741), Cilento (1326/1414) and Carlantino (1110/1241) populations, respectively. SNP genotypes used for LD estimation were obtained from Illumina 370K chip (San Diego, CA, USA) data available for each population study.
Association analysis
Association analyses were performed using a χ2 test implemented in PLINK software (http://pngu.mgh.harvard.edu/~purcell/plink/) and an additive genetic model was tested. The analyses were carried out considering both, each population separately and combining the three populations all together.
Relatedness between individuals was considered in the association testing taking advantage of genealogical information available for each population. A corrected statistics was obtained by the estimation of a correction factor λ through simulations following the procedure proposed by Thorgeirsson et al.5 In detail, the genotype of the rs1051730 variant was randomly assigned to the founders of each genealogy using the allele frequency distribution estimated with the BLUE estimator on the data and assuming the Hardy–Weinberg equilibrium for the marker. Genotypes of all non-founder individuals were then randomly drawn conditional on their parent genotypes and independently from their phenotype. Only genotypes of individuals available in the real data were considered, all others were set to missing. A number of 1 00 000 simulations were performed using this gene-dropping method implemented in the Genedrop program from the MORGAN package (http://www.stat.washington.edu/thompson/Genepi/).
For each simulation, a χ2 test was carried out and the mean of χ2 distribution was considered as λ. Moreover, for the Cilento sample, the λ was also estimated from genetic data (1073 microsatellite markers distributed in the genome) by applying the genomic control approach.16
For the combined sample (Val Borbera–Cilento–Carlantino), the Cochran–Mantel–Haenszel (CMH) and Breslow–Day tests (both implemented in PLINK) were performed. The CMH provides a test based on an ‘average’ odds ratio (OR) adjusted for the possible confounding effects because of population stratification; whereas the Breslow–Day statistic tests for homogeneity of effects between the three groups taking into account the population of origin.
Linkage disequilibrium and HW estimation
The Hardy–Weinberg (HW) equilibrium were assessed using the Haploview v. 4.1 program (http://www.broad.mit.edu/mpg/haploview/). For each population, the LD pattern and the HW equilibrium were evaluated on a subset of poor-correlated individuals. The LD pattern in the variant region was estimated using 18 SNPs located approximately 200 kb around the rs1051703 variant. The method proposed by Zaykin et al17 was applied to compare the LD matrices between each couple of samples (Cilento/ValBorbera, Cilento/Cardile and Valborebera/Cardile). Briefly, this method tests the difference of pair-wise Δ-prime matrix between two groups (ie, Cilento/ValBorbera) and applies a simulation procedure to assess the type I error. For each simulation, the individuals are randomly assigned to the two populations and an empirical statistics is recomputed. In our study, 100 000 simulations were applied.
Results
Allele frequency of the rs1051730 variant was calculated in each population sample. Interestingly, the minor allele frequency (fA) in the Cilento population (fA=0.366) was similar to the frequency in the Hap Map CEU reference population (fA=0.385); whereas, in the Val Borbera (fA=0.412) and Carlantino (fA=0.417) populations the fA was higher and comparable to the frequency of the Hap Map ITS (sample from Tuscany region of Italy) reference population (fA=0.412). The HW equilibrium was respected in all samples (Val Borbera P=0.11, Cilento P=0.1 and Carlantino P=0.42).
A comparison of the rs1051730 variant allele frequency between the current and never smokers and between the former and the current smokers was performed and no significant difference was observed (not shown).
A significant difference was evident between low-dose smokers and high-dose smokers. In Val Borbera and Cilento populations, a strong increase in the A allele (+5.8 and +10.2%, respectively) was observed among the heavy smokers (Table 1). In both cases the enrichment reached the threshold of statistical significance also after correction for relatedness among individuals (Val Borbera P=0.0151 and Cilento P=0.022). However, as in the Cilento sample the value of λ was <1, it did not affect the original statistics. The λ was estimated by a genealogy-based approach in the three populations. For the Cilento population, a genomic-based approach was also used, as described in Materials and methods section. Comparable values of the λ were obtained from the two approaches (0.984 versus 1 from genealogy and genome, respectively).
The A allele of rs1051730 resulted only slightly enriched (+1%) among high-dose smokers in the Carlantino population. Interestingly, if the threshold for low-/high-dose smokers definition was increased to 15 CPD, the difference in the A allele frequency among cases and controls resulted more marked (+3.2%) but still not statistically significant.
A comparison of the LD pattern in the variant region across the three population samples was performed to investigate about the lack of association in Carlantino. The LD between 18 SNPs, located approximately 100-kb upstream and downstream the rs1051730 variant, was analyzed. Interesting, LD pattern was significantly different between Carlantino and the other two populations: Val Borbera (P<10−6) and Cilento (P=9.10−4). As expected, no significant difference in the LD pattern was found between Val Borbera and Cilento (P=0.514) (see Supplementary Figure 1). This result suggests that LD difference in Carlantino could be a possible explanation of the association result observed in this population.
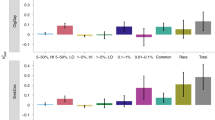
In order to increase the power of the analysis the case–control association was also carried out on the combined sample (Val Borbera–Cilento–Carlantino). The CMH was applied to avoid the confounding effects because of the population origin and an OR of 1.26 (95% confidence interval (CI) 1.07–1.49, P=0.0057) was obtained confirming the association of the variant A with the heavy smoking. This result was strengthened by the Breslow–Day test that supported the hypothesis of the homogeneity of OR giving a P-value of 0.285, data shown in the Figure 1.
OR and CI from all studies. A forest plot for the three studies with OR and 95% CI is shown. A log scale is used on the x axis. The width of the boxes is proportional to the precision of the study and the ends of the horizontal lines represent lower and upper 95% confidence limits. The left vertical line marks an OR of 1.0, corresponding to any risk. For the Val Borbera and Cilento populations, the reported OR are greater than unity indicating an increase in the risk of becoming a heavy smoker. The central diamond corresponds to the OR for the combined sample and it is obtained applying the CMH test. The homogeneity of effects between the three groups is tested by Breslow–Day statistic test.
The effect of the rs1051730 variant in the etiology of some diseases for which smoking is a strong risk factor was examined. A total of 138 cases of malignant tumors (about one-third being breast cancers) were assessed in the three populations and were compared with 3185 not affected individuals. No association was found taking into account sex, age and smoking as covariates in logistic regression analysis. Similarly, no association was found comparing 160 individuals presenting cardiovascular disorders (stroke and myocardial infarction) with 3979 control individuals (data not shown).
Discussion
Genetic factors have been suggested to contribute to individual variations in smoking behavior and several have been identified in recent years through GWA studies and linkage analysis.5, 6, 7, 8, 9, 18 Many have been replicated in different populations and genetic factors for smoking cessation, number of CPD, smoking initiation and nicotine addiction have been confirmed.5, 6, 7, 8, 9, 19, 20 Here, we report a replication of the rs1051730 variant in the CHRNA5-A3-B4 gene cluster, associated with smoking quantity, in isolated populations from Italy. As shown by recent studies of the European population,21 the Italian population is rather heterogeneous from the genetic point of view, possibly because of its geographical position and its history, characterized by multiple population admixtures. The study of isolated populations from different parts of the country seems therefore a useful tool to define the complexity of the population structure and to highlight genetic differences that may lead to discovery of less common genetic variations.22
The A allele of the rs1051730 variant, in all three isolated populations, showed a frequency similar to those described in European and Middle Eastern population,11 even if in Cilento sample this variant was less frequent. The association was replicated in two isolates, Val Borbera (in the North) and Cilento (in the South) thus, confirming the association of this variant to smoking quantity also in our country. Interestingly, Carlantino did not show any significant association but only a trend that was stronger if the cut-off of smokers was set at 15 CPD. The lack of association observed in Carlantino could be attributed to a reduced power in this sample. However, the different LD pattern detected in Carlantino in the variant region, suggests that the rs1051730 variant is not the functional variant but is in LD with the causal variant in Cilento and Val Borbera populations. A different proxy should be present in Carlantino. Further study of the region, including a sequencing of the region in larger sets of individuals, will be required to elucidate this hypothesis.
References
World Health Organization: Economics of Tobacco Control, (www.who.int/gb/fctc/PDF/wg1/e1t2.pdf).
World Health Organization: World Health Statistics 2008, (http://www.who.int/whosis/whostat/2008/en/index.html).
Li MD, Cheng R, Ma JZ, Swan GE : A meta-analysis of estimated genetic and environmental effect on smoking behaviour in male and female adult twins. Addiction 2003; 98: 23–31.
Li MD : Identifying susceptibility loci for nicotine dependence: 2008 update based on recent genome-wide linkage analyses. Hum Genet 2008; 123: 119–131.
Thorgeirsson TE, Geller F, Sulem P et al: A variant associated with nicotine dependence, lung cancer and peripheral arterial disease. Nature 2008; 452: 638–642.
Bierut LJ, Stitzel JA, Wang JC et al: Variants in nicotine receptors and risk for nicotine dependence. Am J Psychiatry 2008; 165: 1163–1171.
Bierut LJ, Madden PA, Breslau N et al: Novel genes identified in a high-density genome wide association study for nicotine dependence. Hum Mol Genet 2007; 16: 24–35.
Berrettini W, Yuan X, Tozzi F et al: Alpha-5/alpha-3 nicotinic receptor subunit alleles increase risk for heavy smoking. Mol Psychiatry 2008; 13: 368–373.
Weiss RB, Baker TB, Cannon DS et al: A candidate gene approach identifies the CHRNA5-A3-B4 region as a risk factor for age-dependent nicotine addiction. PLoSGenet 2008; 4: e1000125.
Caporaso N, Gu F, Chatterjee N et al: Genome-wide and candidate gene association study of cigarette smoking behaviors. PLoS One 2009; 4: e4653.
Hung RJ, McKay JD, Gaborieau V et al: A susceptibility locus for lung cancer maps to nicotinic acetylcholine receptor subunit genes on 15q25. Nature 2008; 452: 633–637.
Amos CI, Wu X, Broderick P et al: Genome-wide association scan of tag SNPs identifies a susceptibility locus for lung cancer at 15q25.1. Nat Genet 2008; 40: 616–622.
Saccone SF, Hinrichs AL, Saccone NL et al: Cholinergic nicotinic receptor genes implicated in a nicotine dependence association study targeting 348 candidate genes with 3713 SNPs. Hum Mol Genet 2007; 16: 36–49.
Sala C, Ciullo M, Lanzara C et al: Variation of hemoglobin levels in normal Italian populations from genetic isolates. Haematologica 2008; 93: 1372–1375.
Grant SF, Thorleifsson G, Reynisdottir I et al: Variant of transcription factor 7-like 2 (TCF7L2) gene confers risk of type 2 diabetes. Nat Genet 2006; 38: 320–323.
Bacanu S, Devlin B, Roeder K : Association Studies for quantitative traits in structured populations. Genet Epidemiol 2002; 22: 78–93.
Zaykin DV, Meng Z, Ehm MG : Contrasting linkage-disequilibrium patterns between cases and controls as a novel association-mapping method. Am J Hum Genet 2006; 78: 737–746.
Li MD : Identifying susceptibility loci for nicotine dependence: 2008 update based on recent genome-wide linkage analyses. Hum Genet 2008; 123: 119–131.
Uhl GR, Liu QR, Drgon T et al: Molecular genetics of successful smoking cessation. Arch Gen Psychiatry 2008; 65: 683–693.
Vink JM, Smit AB, de Geus EJC et al: Genome-wide association study of smoking initiation and current smoking. Am J Hum Genet 2009; 84: 367–379.
Novembre J, Johnson T, Bryc K et al: Genes mirror geography within Europe. Nature 2008; 456: 98–101.
Kristiansson K, Naukkarinen J, Peltonen L : Isolated populations and complex disease gene identification. Genome Biol 2008; 9: 109.
Acknowledgements
We are indebted to the inhabitants of the three isolates, Val Borbera, Carlantino and Cilento (Gioi/Cardile and Campora) who, participating to the projects, have made this study possible. We also thank the local administrations, the MD and the numerous people from each village who helped with data collection. The work was supported by Compagnia di San Paolo, Torino, Cariplo Fundation, Milano, Italian Ministry of Health (Grants Ricerca Corrente and Grants Ricerca Finalizzata, to PG and DT), Telethon Fundation, the Italian Ministry of Universities (FIRB -RBIN064YAT) to MC. RS and TN were supported by a fellowship of Regione Campania, Italy.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on European Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Sorice, R., Bione, S., Sansanelli, S. et al. Association of a variant in the CHRNA5-A3-B4 gene cluster region to heavy smoking in the Italian population. Eur J Hum Genet 19, 593–596 (2011). https://doi.org/10.1038/ejhg.2010.240
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2010.240
Keywords
This article is cited by
-
Nicotinic acetylcholine gene cluster CHRNA5-A3-B4 variants influence smoking status in a Bangladeshi population
Pharmacological Reports (2021)
-
Human Genetics of Addiction: New Insights and Future Directions
Current Psychiatry Reports (2018)
-
Converging findings from linkage and association analyses on susceptibility genes for smoking and other addictions
Molecular Psychiatry (2016)
-
Gene network analysis shows immune-signaling and ERK1/2 as novel genetic markers for multiple addiction phenotypes: alcohol, smoking and opioid addiction
BMC Systems Biology (2015)
-
Association between CHRNA3 rs1051730 genotype and lung cancer risk in Chinese Han population: A case-control study
Journal of Huazhong University of Science and Technology [Medical Sciences] (2013)