Abstract
Huntington disease (HD) results from CAG expansion in the huntingtin (HTT) gene. Although HD occurs worldwide, there are large geographic differences in its prevalence. The prevalence in populations derived from Europe is 10–100 times greater than in East Asia. The European general population chromosomes can be grouped into three major haplogroups (group of similar haplotypes): A, B and C. The majority of HD chromosomes in Europe are found on haplogroup A. However, in the East-Asian populations of China and Japan, we find the majority of HD chromosomes are associated with haplogroup C. The highest risk HD haplotypes (A1 and A2), are absent from the general and HD populations of China and Japan, and therefore provide an explanation for why HD prevalence is low in East Asia. Interestingly, both East-Asian and European populations share a similar low level of HD on haplogroup C. Our data are consistent with the hypothesis that different HTT haplotypes have different mutation rates, and geographic differences in HTT haplotypes explain the difference in HD prevalence. Further, the bias for expansion on haplogroup C in the East-Asian population cannot be explained by a higher average CAG size, as haplogroup C has a lower average CAG size in the general East-Asian population compared with other haplogroups. This finding suggests that CAG-tract size is not the only factor important for CAG instability. Instead, the expansion bias may be because of genetic cis-elements within the haplotype that influence CAG instability in HTT, possibly through different mutational mechanisms for the different haplogroups.
Similar content being viewed by others
Introduction
Huntington disease (HD) is a progressive neurodegenerative disorder that is dominantly inherited and results from a mutation that expands the polymorphic trinucleotide (CAG) tract in HTT. The average CAG-tract size in the general population is 16–20 repeats. However, in HD patients the CAG-tract has expanded to 36 repeats or greater.1
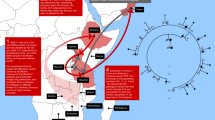
Although HD is found worldwide, there are significant geographic differences in its prevalence.2, 3, 4, 5 The highest prevalence rates are reported for Western populations from Europe, where the minimum prevalence is >5 per 1 00 000 (Figure 1 and Supplementary Table 1). The origins of HD chromosomes in the United States, Canada, South Africa, Australia, the Caribbean, the Indian Subcontinent and Venezuela can be genealogically traced to European origins6 and have similar HD prevalence rates to Europe.
Worldwide estimates of the prevalence of HD. Overall, the prevalence of HD is much higher in European populations than in East Asia. Average minimum prevalence on the basis of several studies are shown (references in Supplementary Table 1). Note that prevalence studies occurring before the discovery of the HD gene in 1993 could underestimate the true prevalence of HD by as much as 14–24%.17, 45 In particular, many of the studies in Africa have small sample sizes and the HD diagnosis has not confirmed by molecular testing. As HD phenocopy disorders are relatively common in Africa,46 these studies could have significantly overestimated the HD prevalence in these regions. Currently, the Maracaibo region of Venezuela has the highest reported worldwide prevalence of HD (700 per 1 00 000).39 Venezuela was colonized by the Spanish in the 16th century, and the origins of HD in Venezuela can be traced back to Europe.38 Also see Harper,4 Conneally47 and Al Jader et al2 for earlier reviews on the worldwide prevalence of HD.
The origins of HD chromosomes in Eastern populations such as China and Japan are less clear. The minimum HD prevalence is estimated to be 0.1–0.5 per 1 00 000 in China7, 8 and Japan.9, 10, 11 Previous studies have speculated that HD chromosomes in the Chinese population have the same origin as the European population.7 HD patients in China and Japan appear to have similar clinical and pathological features, but the disease occurs at a much lower frequency than in Europe.8, 12 The reasons for the 10–100-fold lower prevalence of HD in China and Japan (hereafter referred to as ‘East Asia’) compared with European populations is not well understood.
New mutations make a significant contribution to the prevalence of HD. Expanded CAG-tracts can be unstable and have a tendency to increase in size across generations, particularly when paternally transmitted.13, 14 New mutations for HD occur when the CAG-tract of a transmitted chromosome expands to 36 repeats or greater. The intermediate allele (IA; 27–35 CAG) class of chromosomes have a larger than normal CAG-tract in HTT. Individuals with an IA will not develop HD themselves, but their offspring may be at risk of inheriting a CAG-tract that has expanded into the HD range (>35 CAG).15 Estimates of the mutation flow of the CAG-tract16 and the frequency of HD cases that do not have a previous family history of HD17 suggest the new mutation rate may be as high as 10% in some populations.
Furthermore, there is an inverse relationship between the CAG-tract size and the age of onset of the disease that accounts for the ‘anticipation’ of the disease (earlier onset) in subsequent generations.18, 19 As the CAG-tract continues to expand from generation-to-generation, the age of onset gets earlier. Eventually, with large CAG-tract sizes (>60 repeats), the onset of the disease occurs before reproductive age (juvenile onset), and the disease chromosome does not have an opportunity to be passed on to the next generation. The current prevalence of HD in any population is therefore the result of the balance between the incidence of new mutations caused by CAG expansion into the disease range and the incidence of CAG expanded chromosomes eliminated by anticipation.20
CAG expansion in European populations does not occur randomly, but is associated with specific HTT haplotypes. In a previous study, three broadly defined haplogroups (A, B and C) were found in the HTT region in the general population, but both HD chromosomes and intermediate alleles for HD were almost exclusively found on haplogroup A.21 Further, CAG-expanded chromosomes were significantly more likely to be associated with two specific haplogroup A variants (A1 and A2) than any other HTT haplotype. However, these variants (A1 and A2) were not exclusive to CAG-expanded chromosomes, suggesting that these variants might have an increased mutation rate relative to other HTT haplotypes in the general population. We have previously proposed that this increased association of CAG expansion with A1 and A2 in the European population is most consistent with the hypothesis that genetic cis-elements found on these haplotypes are predisposed to CAG expansion.21, 22 The purpose of this study was to determine if HD chromosomes in populations of East-Asian descent arise on the same haplotype as seen in European populations.
Methods
In order to determine the haplotypes of HD chromosomes in European (n=199 independent chromosomes from unrelated individuals >35 CAG, including 65 samples reported previously 21) and Chinese and Japanese10, 23 (n=31 independent unrelated chromosomes >35 CAG) populations, genotyping was performed at 21 tagSNP positions in the HTT region using a customised GoldenGate assay on the Illumina BeadArray platform. Phasing was performed using PHASE-224 with family trio information to assign the SNP alleles to chromosomes of known CAG size. CAG-tract size was measured using methods described previously.25 Haplogroups were called using the criteria developed previously21 (Supplementary Table 2). Statistical analysis was performed using Excel or ‘R’ software (www.r-project.org) as appropriate. P-values are reported and indicated with nonsignificant (n.s.) when they exceed alpha=0.05. Ethnicity of subjects was self-reported and confirmed by independent self-reports of family members. The use of both previously collected de-identified archived samples and those collected from consenting UBC HD Clinic patients and families was approved for this study by the UBC/Children's and Women's Health Centre of British Columbia Research Ethics Board (UBC C&W REB H05-70532).
Results and discussion
Confirming our previous results, HD chromosomes in European populations are most likely to occur on a subset of HTT-specific haplotypes within haplogroup A (Supplementary Table 3).21 Variants A1 and A2 have the highest risk for CAG expansion in European populations (Odds Ratio 13.9 and 2.8; P-value 4.8 × 10−36 and 9.9 × 10−7, respectively). CAG expansion may still occur on variant A3 (OR 0.9; P-value 0.82 n.s.) but in contrast, variants A4 and A5 (OR 0.2 and 0.0; P-value 6.6 × 10−4 and 2.9 × 10−4) appear to be protected from the mutation relative to other haplotypes in European populations.
In contrast, HD chromosomes in East Asia were associated with different haplotypes than seen in Europe. In Chinese and Japanese, CAG expansion was most frequently associated with haplogroup C (OR 5.2; P-value 4.4 × 10−4) and less commonly associated with A5 and haplogroup B (OR 0.4 and 0.5, P-value=0.11 n.s. and P-value=0.37 n.s.; Figure 2). We did not find any Chinese or Japanese chromosomes with CAG expansion on haplogroup variants A1 or A2, which was most common in Europe.
HTT haplogroups of CAG-expanded chromosomes (>35CAG). The HD mutation occurs on different haplogroups in East-Asian and European populations. In Europe, CAG expansion is most likely to be found on haplogroup A, particularly variants A1 and A2. In East Asia, CAG expansion is associated with different haplogroups, including haplogroup C. Note that ‘Other’ haplotypes (white) could not be easily categorized into haplogroup A, B or C. ‘A-Other’ haplogroup variants (grey) could not be easily categorized into a variant of haplogroup A. Number of chromosomes are indicated in brackets. The data are also presented in Supplementary Table 3.
To determine the frequency of the different haplogroups on control chromosomes (<26CAG), genotyping and phasing was performed on DNA from family trios in the general population from Europe (n=428 trios), China and Japan (n=73 trios). Haplogroup frequencies among control chromosomes in the general population were similar within the different European populations and within the East-Asian populations. (Supplementary Table 3). However, the HTT haplogroup frequencies in the general population (<26CAG) are very different between the European and East-Asian populations (Figure 3). The A1 and A2 variants, highly associated with European CAG expanded chromosomes, were found in up to 20% of the general population in Europe. In contrast, these two highest risk haplogroup variants are completely absent from the general population in East Asia in our sample. One explanation for the lower prevalence of HD in East Asia could therefore be the absence of predisposing A1 and A2 haplotypes in their general population.
HTT haplogroups of the general population (<27CAG). There is a diversity of haplogroups found in the general population of Europe, despite the fact that CAG expansion is most likely to occur on haplogroup A in this population (compare with Figure 2). Note that haplogroup A, and the variants with the highest risk of CAG expansion in the European population (A1 and A2, in red) are absent from the general population of China and Japan. Number of chromosomes are indicated in brackets. The data are also presented in Supplementary Table 3.
Overall, the average CAG-tract size in the East-Asian general population was 16.9 repeats (Table 1). This is lower than the average CAG-tract size of 17.8 repeats in Europeans and consistent with previous reports.26 In East Asia, the frequency of haplogroup C on HD chromosomes is significantly increased relative to the frequency of haplogroup C in the general population (Supplementary Table 4, χ2 P-value 4.4 × 10−4). This suggests that new mutations for HD do not occur randomly on any chromosome, but that there is a mutation bias that increases the probability of CAG-tract expansion on haplogroup C in this population.
Interestingly, HD occurs on haplogroup C at a similar frequency in both Europe and East-Asian populations, and may be the result of a baseline level of CAG instability of this specific haplogroup. HD prevalence in Europe on average is estimated at 7.5 in 100 000 and expansion occurs on haplogroup C only 2% of the time in this population. Therefore, the specific prevalence of HD on haplogroup C in Europe would be estimated at 0.15 in 100 000. Although the overall prevalence of HD in East Asia is 10–100 times lower than in Europe, this is similar to the estimated specific prevalence of HD on haplogroup C in the East of 0.19 per 100 000 (Table 2). Although the frequency of HD on haplogroup C in Europe is only a minor proportion of the overall prevalence, the specific prevalence of HD on haplogroup C is very similar to that in East Asia. Haplogroup C remains the most susceptible haplogroup in East Asia because the haplogroup variants with higher risk (A1 and A2) are absent from this population.
The overall prevalence of HD in any population is the balance between the incidence of new mutations for HD and loss of HD chromosomes because of negative selection of very large CAG-tracts that result in juvenile onset HD. This equilibrium is therefore critically dependant on forces that alter the CAG-tract size of normal (<27 CAG) and intermediate alleles (27–35 CAG) of HTT, which are not subject to significant negative selection. Why is the HD mutation more likely to occur on specific HTT haplotypes? We propose two potentially complementary explanations:21, 22, 27
-
1)
The bias is due to a larger average CAG-tract size of that haplogroup in the general population. The larger average CAG size could be due to a founder mutation or genetic drift. CAG instability is positively and exponentially related the size of the CAG-tract,16, 28 and a larger average CAG-tract size would make that haplogroup more likely to expand and become a new mutation for HD.
-
2)
Certain haplogroups contain genetic cis-elements in the HTT region that increase the probability of CAG expansion relative to other haplogroups. These cis-elements may be particularly important when the CAG-tract size is in the normal or intermediate range and the influence of CAG-tract size itself is reduced.
To discriminate between these two possibilities, we compared the CAG-tract size distribution between C and non-C haplotypes in East Asia (Figure 4). Interestingly, the mean CAG-tract size of haplogroup-C chromosomes (16.0 repeats) was not higher than non-C chromosomes (17.2 repeats) in the general population of East Asia. This is not consistent with the first explanation above because the CAG expanded chromosomes on haplogroup C do not appear to have come from a pool of intermediate alleles on the same haplotype in East Asia.
CAG size distributions of haplogroup C (a) East Asia and (b) Europe. In East Asia, the majority of HD mutations occur on haplogroup C. In the general population of East Asia, the average CAG-tract size of haplogroup C is less than other haplogroups, suggesting that the bias of expansion is not simply because of a large average CAG-tract size (Note that the CAG-tract size distribution is discontinuous because of the fact that there is an ascertainment bias towards discovering chromosomes with >36CAG due to HD symptoms).
Instead, the data argue that CAG-tract size cannot be the only factor important for CAG instability and supports the second explanation above. This conclusion is further supported by a clear precedent for genetic cis-elements influencing CAG instability for other genes.29 Although there are many CAG-tract-containing genes in the genome, each has a different propensity to CAG instability that is not explained by differences in mean CAG-tract size30 but could be explained by cis-elements that influence CAG instability. For example, CTCF binding sites located cis to the ATXN7 gene can regulate CAG hyperinstability at that locus.31, 32 We hypothesize that genetic cis-elements present in different HTT haplogroups may be regulating CAG instability at the HTT locus. Further experiments are clearly needed to demonstrate this.
Although the sample size is relatively small, particularly in the East-Asian samples, our data are consistent with the hypothesis that different HTT haplotypes have different mutation rates, and geographic differences in HTT haplotypes explain the difference in prevalence of HD (Figure 5a). The A1 and A2 haplotypes that comprise the majority of European HD chromosomes are not present in the East-Asian populations, and may have arisen in Europe following the separation of these two populations (Figure 5b) about 25 000 years ago.33, 34 There also appears to be a consistent trend of higher prevalence estimates in Western Europe compared with Eastern Europe, suggesting the A1 and A2 haplotypes may have originated from the Western coast of Europe. This prevalence trend is emphasised in Finland35 (0.5 per 100 000), where geographic isolation and reduced immigration is believed to result in a lower prevalence of HD than its coastal neighbours Sweden36 (5 per 100 000) and Norway37 (7 per 100 000). The genealogical path of HD into many European-derived populations in North America and Australia is known,6 although there is a notable lack of basic prevalence information from South America, aside from Venezuela.38, 39 Prevalence estimates from the Indian subcontinent are also limited to one study of immigrants to the UK,40 but these data suggest the prevalence of HD in India may be higher than in East Asia. This is supported by further molecular analysis of HTT in the Indian population41 but warrants further examination. More data are also needed from Africa, where a number of studies have found that the prevalence of HD is much higher in Whites than Blacks,42, 43 presumably because of the introduction of HD in South Africa and Mauritius from Europe.6, 44
Model of CAG expansion in HTT. (a) At least two mutation rates and potentially two mutational mechanisms, may contribute to CAG expansion in HTT. The low prevalence of HD on haplogroup C is consistent in both East-Asian and European populations and is explained by a low level of CAG expansion that occurs on haplogroup C (blue). The mutation rate of the second mechanism (red) is much higher and therefore results in a higher overall prevalence of HD in European populations. (b) The genetic changes that produced haplotypes A1/A2 appear to have occurred after the separation of Asian and European chromosomes (red arrows), as these haplotypes are completely absent from the Chinese and Japanese populations. The genetic changes that produced A1/A2 may have altered cis-elements in and around the HTT gene, resulting in a higher rate of CAG-expansion. Chromosomes from Europe serve as the origin of HD in several other populations including North and South America, South Africa and Australia through human migration.6
Despite the similarities in clinical phenotype,8, 26 the origin of the majority of HD chromosomes in East Asia and Europe are different. The prevalence of HD is 10–100-fold greater in Europe than in East Asia, and the haplogroups that are closely associated with CAG expansion in European populations (haplogroups A1 and A2), are found in up to 47% of a sample of the general population in Europe.21 In contrast, these haplogroups are absent from populations of Chinese or Japanese descent, and provide an explanation for why the HD prevalence is low in East-Asian populations. However, there may be a common origin of a subset of HD cases found on haplogroup C, as the prevalence of HD was similar, although at a low level in both populations. Overall, we find that CAG expansion at the HTT locus occurs preferentially on specific haplogroups. The highest risk haplogroups are found in the European population, which presumably emerged after the separation of European and Asian populations.
References
Walker FO : Huntington's disease. Lancet 2007; 369: 218–228.
Al-Jader LN, Harper PS, Krawczak M, Palmer SR : The frequency of inherited disorders database: prevalence of Huntington disease. Community Genet 2001; 4: 148–157.
Andrew SE, Hayden MR : Origins and evolution of Huntington disease chromosomes. Neurodegeneration 1995; 4: 239–244.
Harper PS : The epidemiology of Huntington's disease. Hum Genet 1992; 89: 365–376.
Squitieri F, Andrew SE, Goldberg YP et al: DNA haplotype analysis of Huntington disease reveals clues to the origins and mechanisms of CAG expansion and reasons for geographic variations of prevalence. Hum Mol Genet 1994; 3: 2103–2114.
Hayden MR : Huntington's Chorea. New York: Springer, 1981.
Leung CM, Chan YW, Chang CM, Yu YL, Chen CN : Huntington's disease in Chinese: a hypothesis of its origin. J Neurol Neurosurg Psychiatry 1992; 55: 681–684.
Chang CM, Yu YL, Fong KY et al: Huntington's disease in Hong Kong Chinese: epidemiology and clinical picture. Clin Exp Neurol 1994; 31: 43–51.
Nakashima K, Watanabe Y, Kusumi M et al: Epidemiological and genetic studies of Huntington's disease in the San-in area of Japan. Neuroepidemiology 1996; 15: 126–131.
Masuda N, Goto J, Murayama N, Watanabe M, Kondo I, Kanazawa I : Analysis of triplet repeats in the huntingtin gene in Japanese families affected with Huntington's disease. J Med Genet 1995; 32: 701–705.
Adachi Y, Nakashima K : Population genetic study of Huntington's disease—prevalence and founder's effect in the San-in area, western Japan. Nippon Rinsho 1999; 57: 900–904.
Narabayashi H : Huntington's Chorea in Japan: review of the literature. Adv Neurol 1973; 1: 253–259.
De Rooij KE, Koning Gans PA, Skraastad MI et al: Dynamic mutation in Dutch Huntington's disease patients: increased paternal repeat instability extending to within the normal size range. J Med Genet 1993; 30: 996–1002.
Goldberg YP, McMurray CT, Zeisler J et al: Increased instability of intermediate alleles in families with sporadic Huntington disease compared to similar sized intermediate alleles in the general population. Hum Mol Genet 1995; 4: 1911–1918.
Semaka A, Creighton S, Warby S, Hayden MR : Predictive testing for Huntington disease: interpretation and significance of intermediate alleles. Clin Genet 2006; 70: 283–294.
Falush D, Almqvist EW, Brinkmann RR, Iwasa Y, Hayden MR : Measurement of mutational flow implies both a high new-mutation rate for Huntington disease and substantial underascertainment of late-onset cases. Am J Hum Genet 2001; 68: 373–385.
Almqvist EW, Elterman DS, MacLeod PM, Hayden MR : High incidence rate and absent family histories in one quarter of patients newly diagnosed with Huntington disease in British Columbia. Clin Genet 2001; 60: 198–205.
Langbehn DR, Hayden MR, Paulsen JS : CAG-repeat length and the age of onset in Huntington disease (HD): a review and validation study of statistical approaches. Am J Med Genet B Neuropsychiatr Genet 2010; 153B: 397–408.
Langbehn DR, Brinkman RR, Falush D, Paulsen JS, Hayden MR : A new model for prediction of the age of onset and penetrance for Huntington's disease based on CAG length. Clin Genet 2004; 65: 267–277.
Rubinsztein DC, Amos W, Leggo J et al: Mutational bias provides a model for the evolution of Huntington's disease and predicts a general increase in disease prevalence. Nat Genet 1994; 7: 525–530.
Warby SC, Montpetit A, Hayden AR et al: CAG expansion in the Huntington disease gene is associated with a specific and targetable predisposing haplogroup. Am J Hum Genet 2009; 84: 351–366.
Warby SC, Visscher H, Butland S, Pearson CE, Hayden MR : Response to Falush: a role for cis-element polymorphisms in HD. Am J Hum Genet 2009; 85: 942–945.
Almqvist E, Spence N, Nichol K et al: Ancestral differences in the distribution of the delta 2642 glutamic acid polymorphism is associated with varying CAG repeat lengths on normal chromosomes: insights into the genetic evolution of Huntington disease. Hum Mol Genet 1995; 4: 207–214.
Stephens M, Smith NJ, Donnelly P : A new statistical method for haplotype reconstruction from population data. Am J Hum Genet 2001; 68: 978–989.
Potter NT, Spector EB, Prior TW : Technical standards and guidelines for Huntington disease testing. Genet Med 2004; 6: 61–65.
Soong BW, Wang JT : A study on Huntington's disease associated trinucleotide repeat within the Chinese population. Proc Natl Sci Counc Repub China B 1995; 19: 137–142.
Falush D : Haplotype background, repeat length evolution, and Huntington's disease. Am J Hum Genet 2009; 85: 939–942.
Kremer B, Almqvist E, Theilmann J et al: Sex-dependent mechanisms for expansions and contractions of the CAG repeat on affected Huntington disease chromosomes. Am J Hum Genet 1995; 57: 343–350.
Pearson CE, Edamura KN, Cleary JD : Repeat instability: mechanisms of dynamic mutations. Nat Rev Genet 2005; 6: 729–742.
Butland SL, Devon RS, Huang Y et al: CAG-encoded polyglutamine length polymorphism in the human genome. BMC Genomics 2007; 8: 126.
Libby RT, Monckton DG, Fu YH et al: Genomic context drives SCA7 CAG repeat instability, while expressed SCA7 cDNAs are intergenerationally and somatically stable in transgenic mice. Hum Mol Genet 2003; 12: 41–50.
Libby RT, Hagerman KA, Pineda VV et al: CTCF cis-regulates trinucleotide repeat instability in an epigenetic manner: a novel basis for mutational hot spot determination. PLoS Genet 2008; 4: e1000257.
Gutenkunst RN, Hernandez RD, Williamson SH, Bustamante CD : Inferring the joint demographic history of multiple populations from multidimensional SNP frequency data. PLoS Genetics 2009; 5: e1000695–e1000695.
Garrigan D, Kingan SB, Pilkington MM et al: Inferring human population sizes, divergence times and rates of gene flow from mitochondrial, X and Y chromosome resequencing data. Genetics 2007; 177: 2195–2207.
Palo J, Somer H, Ikonen E, Karila L, Peltonen L : Low prevalence of Huntington's disease in Finland. Lancet 1987; 2: 805–806.
Mattsson B : Huntington's chorea in Sweden. Acta Psychiatrica Scandinavica Supplementum 1974; 255: 211–220.
Saugstad L, Odegård O : Huntington's chorea in Norway. Psychol Med 1986; 16: 39–48.
Paradisi I, Hernández A, Arias S : Huntington disease mutation in Venezuela: age of onset, haplotype analyses and geographic aggregation. J Hum Genet 2008; 53: 127–135.
Avila-Giron R : Medical and social aspects of Huntington's chorea in the state of Zulia, Venezuela. Adv Neurol 1973; 1: 261–266.
Shiwach RS, Lindenbaum RH : Prevalence of Huntington's disease among UK immigrants from the Indian subcontinent. Br J Psychiatry: J Mental Sci 1990; 157: 598–599.
Saleem Q, Roy S, Murgood U et al: Molecular analysis of Huntington's disease and linked polymorphisms in the Indian population. Acta Neurol Scand 2003; 108: 281–286.
Hayden MR, Beighton P : Huntington's chorea in the Cape coloured community of South Africa. S Afr Med J 1977; 52: 886–888.
Hayden MR, MacGregor JM, Beighton PH : The prevalence of Huntington's chorea in South Africa. S Afr Med J 1980; 58: 193–196.
Scholefield J, Greenberg J : A common SNP haplotype provides molecular proof of a founder effect of Huntington disease linking two South African populations. Eur J Hum Genet 2007; 15: 590–595.
Siesling S, Vetger-van de Vlis M, Losekoot M et al: Family history and DNA analysis in patients with suspected Huntington's disease. J Neurol Neurosurg Psychiatry 2000; 69: 54–59.
Margolis RL, Rudnicki DD, Holmes SE : Huntington's disease like-2: review and update. Acta Neurol Taiwan 2005; 14: 1–8.
Conneally PM : Huntington disease: genetics and epidemiology. Am J Hum Genet 1984; 36: 506–526.
Acknowledgements
The authors thank the HD families for the generous use of their DNA for these studies. Funding has been provided by the Canadian Genetic Diseases Network (CGDN), Huntington Society of Canada (HSC), Michael Smith Foundation for Health Research (MSFHR), Canadian Institutes of Health Research (CIHR), Child and Family Research Institute (CFRI) and the Huntington Disease Society of America (HDSA)/High Q Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on European Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Warby, S., Visscher, H., Collins, J. et al. HTT haplotypes contribute to differences in Huntington disease prevalence between Europe and East Asia. Eur J Hum Genet 19, 561–566 (2011). https://doi.org/10.1038/ejhg.2010.229
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2010.229
Keywords
This article is cited by
-
A clinical study and future prospects for bioactive compounds and semi-synthetic molecules in the therapies for Huntington's disease
Molecular Neurobiology (2024)
-
Loss of CAA interruption and intergenerational CAG instability in Chinese patients with Huntington’s disease
Journal of Molecular Medicine (2023)
-
Antisense Oligonucleotide Therapeutics for Neurodegenerative Disorders
Current Geriatrics Reports (2021)
-
Geographic differences in the incidence of Huntington’s disease in Sardinia, Italy
Neurological Sciences (2021)
-
The known burden of Huntington disease in the North of Scotland: prevalence of manifest and identified pre-symptomatic gene expansion carriers in the molecular era
Journal of Neurology (2021)