Abstract
Multiple Sulfatase Deficiency (MSD) is caused by mutations in the sulfatase-modifying factor 1 gene encoding the formylglycine-generating enzyme (FGE). FGE post translationally activates all newly synthesized sulfatases by generating the catalytic residue formylglycine. Impaired FGE function leads to reduced sulfatase activities. Patients display combined clinical symptoms of single sulfatase deficiencies. For ten MSD patients, we determined the clinical phenotype, FGE expression, localization and stability, as well as residual FGE and sulfatase activities. A neonatal, very severe clinical phenotype resulted from a combination of two nonsense mutations leading to almost fully abrogated FGE activity, highly unstable FGE protein and nearly undetectable sulfatase activities. A late infantile mild phenotype resulted from FGE G263V leading to unstable protein but high residual FGE activity. Other missense mutations resulted in a late infantile severe phenotype because of unstable protein with low residual FGE activity. Patients with identical mutations displayed comparable clinical phenotypes. These data confirm the hypothesis that the phenotypic outcome in MSD depends on both residual FGE activity as well as protein stability. Predicting the clinical course in case of molecularly characterized mutations seems feasible, which will be helpful for genetic counseling and developing therapeutic strategies aiming at enhancement of FGE.
Similar content being viewed by others
Introduction
Multiple sulfatase deficiency (MSD, MIM no. 272200) is a rare autosomal recessive inborn error of metabolism. Worldwide less than 50 cases have been published so far.1 The clinical picture of MSD combines symptoms of the different sulfatase deficiencies. Patients show neurological deterioration and a neurodegenerative course of disease similar to metachromatic leukodystrophy. Developmental delay, dysmorphism and organomegaly are present as found in various mucopolysaccharidoses. Skeletal abnormalities remind one of Chondrodysplasia punctata type I and skin changes of X-linked ichthyosis.1, 2, 3 According to the age of onset, neonatal, late infantile and juvenile disease subtypes can be distinguished.4 Neonatal MSD is the most severe form with postnatal onset and a broad range of mucopolysaccharidosis-like clinical symptoms. According to the few published cases, all neonatal MSD patients died within the first year of life.5, 6, 7, 8 Late infantile MSD resembles late infantile metachromatic leukodystrophy with progressive loss of mental and motor abilities; these clinical features are combined with other symptoms of single sulfatase deficiencies like dysmorphism, skeletal changes and ichthyosis.2, 3, 9, 10, 11, 12 Late infantile MSD can be subdivided into a severe (LIS) and an attenuated (late infantile mild, LIM) form, the latter showing a reduced number of symptoms and later onset beyond the second year of life.13 Late infantile forms include the majority of MSD cases; a Saudi variant form was described separately.14 Up to now, only one juvenile case of MSD has been published with an onset of symptoms in late childhood.15
In MSD, enzymatic activity of every sulfatase is reduced or even absent. A total of 17 sulfatases are encoded in the human genome, 13 of them have been characterized biochemically and 8 are associated with diseases due to single sulfatase deficiencies.10, 16 At least five sulfatases are necessary for non-redundant desulfation of glycosaminoglycans in the lysosome. Defective sulfatase function in MSD thus leads to intralysosomal storage and cellular damage.10
The primary defect in MSD, causing deficiency of a whole enzyme family, affects a specific posttranslational modification of sulfatases that is strictly required for catalytic activity. All sulfatases contain in their active site the so-called sulfatase signature I, comprising the linear amino acid sequence CxPxR. The cysteine within this motif has to be converted to Cα-formylglycine (FGly).17, 18, 19 This modification occurs late co or early posttranslationally in the endoplasmic reticulum (ER) and is mediated by the FGly-generating enzyme (FGE), an unusual mono-oxygenase using molecular oxygen for oxidation of Cβ of the substrate cysteine through its catalytic cysteines 336 and 341.20, 21 FGE is encoded by the sulfatase-modifying factor 1 gene (SUMF1). Patients with MSD carry mutations in this gene.22, 23 Null mutations result in the complete absence of sulfatase activities, as shown in a SUMF1 gene-trap mouse.24 Expression of FGE carrying MSD patient mutations in SUMF1-deficient mouse fibroblasts results in detectable, although extremely low sulfatase activities, underlining that MSD is a monogenetic disease caused by hypomorphic mutations.25 Until now, more than 30 different SUMF1 mutations were found, most of which are missense mutations.9, 23, 26, 20, 27
In a first approach, we investigated the consequences of four different MSD causing SUMF1 mutations on FGE13 expression, localization, stability and activity in either an overexpressing cell-culture model or in patient fibroblasts. We found that all four variant FGE proteins could be expressed in HT-1080 cells, correctly localizing to the ER. Expressing FGE A177P and W179S led to stable proteins, whereas FGE A279V and R349W were unstable. The specific enzyme activity of the purified FGE proteins varied between 1 and 23% as compared with wild-type (wt) FGE. All three patients with both unstable FGE protein and low residual activity expressed FGE R349W and presented a LIS clinical phenotype. All other patients exhibited a LIM phenotype and expressed either unstable FGE with high residual activity or stable FGE with low residual activity. Intracellular sulfatase activities were massively reduced in fibroblasts from patients with the LIS phenotype. However, when measured in cultured fibroblasts of more mildly affected patients, these sulfatase activities were surprisingly variable. Therefore, the molecular analysis of FGE activity and stability seems to provide a more convincing correlation with the clinical presentation of patients of a given genotype.
In this study, we further tested this hypothesis. We extended our previous clinical and biochemical characterization of MSD patients to confirm a potential genotype/phenotype correlation. We included another ten patients showing different clinical presentations and various MSD causing SUMF1 missense and also nonsense mutations. For the first time, a neonatal very severe case with nonsense mutations on both alleles is studied at the molecular level, thus, challenging the concept that only hypomorphic mutations cause MSD.
Materials and methods
Patients and patient's cells
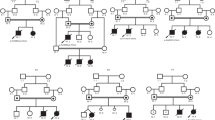
Clinical data were obtained from patients’ relatives by personal communication, patient's data sheets and from already published case reports.28 For ethnic origin and family history of the nine homozygous patients see Supplementary Table S1. Patient skin fibroblasts were grown in Dulbecco's modified Eagle's medium supplemented with 10% fetal calf serum and 1% penicillin/streptomycin (Invitrogen, Karlsruhe, Germany) under 5% CO2 at 37°C.
Western blot analysis
Equivalent aliquots of medium (40 μl) or cell lysate (50 or 100 μg cell protein) were analyzed by western blotting using FGE antiserum or a Hexa-His antibody (Sigma, Munich, Germany) as described earlier.13, 20, 29, 21 ECL signals were quantified using the AIDA software package (Raytest, Straubenhardt, Germany). Endogenous FGE in MSD fibroblasts was detected using ECL femto substrate and enhancer diluted 1:10 in ECL solution (Pierce, Bonn, Germany).
Enzyme assays
Sulfatase activities were measured from whole-cell lysates as described.30, 31 FGE activity assays were performed with FGE enriched from cells and media using the synthetic ASA-derived peptide P23 as substrate.23 See also supplementary information.
Metabolic labeling and immunoprecipitation
Cells stably expressing FGE proteins were grown on 3 cm plates to 80–90% confluence, starved for 1 h in medium depleted of methionine and cysteine and pulsed with 3.7 MBq/ml [35S]methionine/cysteine also for 1 h. Medium was changed to 2 ml medium with 10% fetal calf serum, supplemented with 200 μg/ml methionine and 100 μg/ml cysteine, for 20 min. Cells and medium were harvested either directly or after another change of medium and growth for 1.5, 3 or 6 h (chase). Immunoprecipitation of FGE was performed with FGE antiserum according to a protocol described earlier.32 Proteins were solubilized and subjected to SDS-PAGE. Dried gels were analyzed by phosphorimaging (BAS 1000, Fuji, Düsseldorf, Germany) and densitometric quantification using the MacBAS software (Raytest, Straubenhardt, Germany).
Molecular analysis of SUMF1 mutations, FGE expression, FGE purification, immunofluorescence microscopy
Results
Clinical presentation and mutational analysis of ten patients with MSD
On the basis of Eto's classification,4 we distinguished one neonatal (NVS) and nine late infantile patients. The latter were further subdivided into seven severe (LIS) and two milder (LIM) cases.10, 13 For classification, we considered ten frequent clinical symptoms and also the time point of manifestation of every single symptom (Table 1). In NVS cases every symptom appears in the first months of life, as holds true for patient 1; (skeletal changes had not been defined). The patient was compound heterozygous for two mutations: one in-frame deletion resulting in a loss of 25 amino acids (p.A149_A173del; Supplementary Figure S1) and one mutation leading to a premature stop, thereby disrupting the active site of FGE (p.R327X; Supplementary Figure S1). All other patients, homozygous for four different SUMF1 missense mutations, did not show all of the ten symptoms and their onset varied between LIS and LIM patients, as the latter displayed less symptoms or a later onset of symptoms. Accordingly, the two patients with FGE G263V were assigned to the LIM group, whereas all other patients, expressing FGE S155P (three patients), G247R (three) or R345C (one) (Supplementary Figure S1), belong to the LIS group (Table 1).
Expression of recombinant FGE mutant proteins
All FGE proteins, resulting from the individual SUMF1 mutations, were stably expressed in HT-1080 fibrosarcoma cells. Expressed protein was detected in cell lysates by western blotting using either an anti-hexahistidine or a polyclonal FGE antibody (Figure 1). Expression levels were comparable among different cell lines; only FGE S155P was expressed at a severely reduced level. Even the two nonsense mutations, p.R327X and p.A149_A173del, showed intracellular protein expression, resulting in smaller versions of FGE (Figure 1, upper panel). wt FGE is secreted and processed in the Golgi leading to an N-terminally truncated form in the medium.29, 33 Only three of the FGE variants under study also showed secretion into the cell-culture medium (p.G247R, p.G263V and p.R345C). All other variants, despite their comparable intracellular expression, were not secreted (Figure 1, lower panel). Functionally impaired FGE protein, which is not secreted, is retained intracellularly (and degraded, see below). Among the retained proteins are those resulting from the two nonsense mutations probably leading to a drastically impaired FGE structure (see Supplementary Figure S1). For all secreted FGE variants it is concluded that they are able to pass quality control mechanisms of the ER in spite of protein instability predictions.20
Expression and secretion of mutant FGE in HT-1080 cells. Cells stably overexpressing His-tagged forms of wt FGE and FGE variants were grown to confluence and incubated with fresh medium for 24 h. Equal amounts of cell lysate protein and corresponding medium aliquots were analyzed by western blotting using a polyclonal FGE antibody. The bands visible in the cell lysates (upper panel) represent full-length FGE at 42 kDa. The lower bands labeled with asterisks result from N-terminal processing of wt FGE in the secretory route and are not relevant for intracellular FGE function. FGE A149_A173del and R327X were reduced in size because of the deletion/truncation resulting from the respective mutation. In the medium (lower panel) secreted FGE, detectable almost exclusively as an N-terminally trimmed 38 kDa form, was only visible for wt FGE, as well as for FGE G247R, G263V and R345C. Expression levels varied among the stable cell lines. The data shown were representative for three independent experiments. For endogenous expression of FGE variants in patient fibroblasts see Supplementary Figure S2.
Subcellular localization of mutant FGE proteins
Immunofluorescence analysis of HT-1080 cell clones revealed a correct localization of each FGE variant in the ER, as revealed by colocalization with the ER marker PDI (Figure 2). In addition, the secreted FGE proteins (p.G247R, p.G263V and p.R345C) were also visible in the Golgi (Figure 2r, s and u; red staining) showing colocalization with a Golgi marker (not shown). No costaining was visible with a lysosomal marker (data not shown). The ER localization could be confirmed for the endogenous FGE proteins in MSD patient fibroblasts even though protein expression was extremely reduced and material cross-reacting with the anti-FGE antibody was hardly detectable (Figure 3). Localization of patient FGE protein in the ER is one necessary precondition for residual FGly generation in newly synthesized sulfatases.
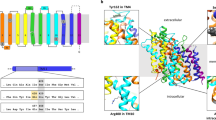
Subcellular localization of FGE variants expressed in HT-1080 cells. Cultured cells, after permeabilization, were stained with a polyclonal FGE antibody (red, a–g) and monoclonal antibodies against organelle markers. Only the ER marker PDI is shown (green, h–n). The merge (yellow, o–u) reveals colocalization of each protein expressed.
Subcellular localization of endogenous FGE in MSD fibroblasts. Endogenous FGE was stained in permeabilized MSD patient fibroblasts and wt control fibroblast cells using a polyclonal FGE antibody (red) and a monoclonal antibody detecting the ER marker PDI (green). The merge (yellow) indicates colocalization inside the ER.
Residual catalytic activity of FGE protein variants
The enzymatic activity of FGE variants was measured using a synthetic peptide derived from arylsulfatase A as substrate (at saturating concentration, 15-times above KM of wt FGE).23 FGE was partially purified from cell lysates or conditioned medium. The activities of variant proteins, relative to wt, ranged from 0 to 15.9% (Figure 4b). FGE R327X, with a disrupted active site, displayed no activity; p.A149_A173del with a 25 amino-acid deletion at the surface of the FGE core domain (Supplementary Figure S1), showed detectable, although very low residual activity of 0.3±0.04%. FGE S155P and R345C showed slightly, but significantly higher residual activities of 1.6±0.4% and 2.0±0.2%, respectively. The highest residual activity was measured for p.G263V (15.9±1.2%). Thus, apart from the catalytically fully impaired FGE R327X, all investigated FGE proteins were able to catalyze at least some FGly generation in vitro.
Stability, retention and residual activity of FGE variants. (a) Stable cell lines were labeled with 35S methionine/cysteine for 1 h and chased for 6 h in label-free medium. FGE was immunoprecipitated from cells and medium, and quantified by densitometry after SDS-PAGE and phosphorimaging. The results shown are representative for three independent experiments. All variant FGE proteins were unstable. Relative secretion (calculated as medium FGE/total FGE after 6 h to correct for degradation within the 6 h) varied, as indicated; n.d., cannot be determined because of massive FGE degradation after 6 h. (b) His-tagged FGE was purified from secretions or cells, respectively. FGE amounts were adjusted according to western blot quantification. Its activity was measured as FGly modification of the synthetic arylsulfatase A substrate peptide P23 using mass spectrometry (see Materials and methods). Specific activities were calculated and expressed as percentage of wt FGE activity. Mean values and SD were calculated from three independent analyses. (c) For a better resolution of FGE secretion and degradation, the pulse chase experiment was repeated, as above (see a), this time with a chase period of only 1.5 h; (relative secretion: medium FGE/total FGE after 1.5 h).
Stability and intracellular retention of FGE protein variants
On the basis of the crystal structure of wt FGE, all chosen SUMF1 mutations were expected to impair protein stability.20 To prove this prediction, we metabolically labeled FGE-expressing HT-1080 cell clones with [35S]methionine/cysteine. Labeled FGE protein was determined before and after 6 h of chase time, as detected by immunoprecipitation of intracellular and secreted FGE, followed by SDS-PAGE and phosphorimaging (Figure 4a). As expected,13 wt FGE was fully stable during the 6 h chase with ∼50% of initial protein remaining inside the cells and ∼50% being secreted (Figure 4a). FGE variants A149_A173del and R327X both were unstable with only about 1% of cross-reacting material detectable after 6 h; also FGE S155P was unstable (about 2%). These three variants likewise could not be detected in the secretions (Figure 4a). The other FGE variants (G247R, G263V and R345C) showed clearly reduced stability as well. In fact, intracellular protein was hardly detectable after the chase also here (0–2% of the initial protein amount); however, considerable amounts of protein were secreted indicating the passing of intracellular protein quality control.
To further analyze degradation and secretion kinetics, we repeated the experiment with chase times of 1.5 and 3 h (Figure 4c and data not shown). FGE R327X showed the lowest (12% residual protein after 1.5 h) and R345C the highest (44%) stability (at 91% stability observed for wt FGE) (Figure 4c). For FGE G247R and G263V, rather low levels of intracellular protein were detected (∼20%), which in part is because of intense secretion. These results confirm the data of Figure 4a. Taken together, all investigated mutations severely affect intracellular FGE protein levels. Apart from degradation, in some cases also increased secretion, that is, impaired protein retention, contributes to reduced intracellular FGE levels.
Expression level of endogenous FGE protein in MSD patient fibroblasts
We analyzed the amount of endogenous FGE protein in whole-cell lysates from patient fibroblasts by immunofluorescence detection (Figure 3) and also by western blotting (Supplementary Figure S2). Fibroblasts from healthy persons served as a positive control. FGE amounts were extremely reduced for all studied patients. No or only faint bands of cross-reacting material were detectable in western blots (Supplementary Figure S2). The extremely reduced levels of defective protein in the patient cells agree with the results from the pulse-chase experiments showing that all analyzed MSD-causing mutations lead to unstable proteins that are degraded inside the cells and, in some cases, also secreted (see above).
Residual sulfatase activities in MSD patient fibroblasts
To determine the functional consequences of MSD mutations on endogenous sulfatase activities, we measured the residual activities of three sulfatases, namely arylsulfatase A (ASA), arylsulfatase C (ASC) and galactose 6-sulfatase (Gal6S) in cell lysates from patient fibroblasts (Table 2). For FGE G263V, we used a fetal fibroblast cell line, as no other patient cells were available. The lowest sulfatase activities were found in the fibroblasts from the NVS patient displaying no detectable activity for ASA, less than 1% for ASC and 1.4% for Gal6S (relative to wt activities, Table 2). The highest activities were measured in the fetal fibroblasts expressing p.G263V found also in the LIM patients 9 and 10 (Table 1). Residual activities of these cells were 17% for ASA, 61% for ASC and 39% for Gal6S (Table 2).
All other cell lines originated from patients with the LIS form of MSD and displayed reduced sulfatase activities, with 2–12% residual activity measured for ASA and 1–22% for Gal6S. ASC activities were also clearly reduced, but even more variable for the different mutations. Whereas only <1–5% residual ASC activity was measured for fibroblasts expressing FGE R345C and G247R, ASC activity in FGE S155P fibroblasts reached values between 14 and 31% of wt activity. It should be noted that patient 2, from whom the fibroblasts with highest ASC activity were derived, developed ichthyosis like all other patients of this study (Table 1).
Discussion
In a first study,13 we were able to show a correlation of both residual activity and stability of FGE variants with the clinical presentation of selected MSD patients. As this study included only four different SUMF1 mutations (homozygous in the patients), it remained open to discussion if the observed genotype/phenotype correlation holds true also for other mutations and for MSD in general. Here, we extended our analysis including another set of clinically thoroughly characterized and genetically defined patients with different forms of disease.
Robust phenotype classification based on the clinical presentation of MSD patients
We investigated ten MSD patients, either historical or actual cases, and classified the patients according to their clinical presentation into groups of NVS, LIM or LIS cases. Our neonatal patient exhibited the whole range of symptoms possible, which agrees with all published neonatal cases.5, 6, 7, 8 Most symptoms were present at birth and the patient showed a rapidly progressive course of disease. The group of late infantile patients9, 11, 12 included in this study could be clearly distinguished from the neonatal patient, as symptoms varied in their age of onset with some features presenting only in later life or others lacking completely. Two of the late infantile patients could be subgrouped as LIM versions of MSD (Table 1), and differentiated from the very rare juvenile form,15 which is not included in our patient collection. Importantly, we can infer from this classification that independent patients with an identical mutation (in homozygous form) display a comparable clinical phenotype.
Molecular phenotype of MSD: differential functional impairments of recombinant FGE
For extending our pathobiochemical understanding of MSD, we expressed all patient SUMF1 mutants in the established HT-1080 cell line.13, 15, 23, 20, 29 Our approach was substantiated by mapping the amino acid substitutions on the crystal structure of wt FGE (see Supplementary Figure S1), which allowed a solid prediction of structural consequences. Without exception, all missense mutations affected strictly conserved residues and were expected to impair FGE protein stability because of unfavorable side chain interactions resulting in clashes with other residues (p.S155P, p.G247R, p.G263V) and/or losses of specific hydrogen bonds (p.R345C).20 However, the consequences on FGE activity remained difficult to predict, because the substitutions did not directly affect the substrate-binding groove, the oxygen-binding pocket or even the catalytic site close to the redox active cysteines.34
All FGE variants could be expressed, including even the variants resulting from nonsense/deletion mutations. Thus, translation of these mRNAs is not attenuated. All FGE proteins localized to the ER (Table 3), thereby fulfilling one important precondition for function, as generation of FGly is a posttranslational modification of nascent, yet unfolded, sulfatase proteins inside the ER.17, 35 Nevertheless, we cannot exclude that the ER localization of defective FGE proteins just mimics a correct localization and in fact is mainly caused by the quality control mechanism inside the ER.36, 37 Importantly, the four studied FGE variants with single substitutions were more (p.G247R, p.G263V and p.R345C) or less (p.S155P) efficient in passing this quality control mechanism, as these proteins were also secreted – just like wt FGE (Figures 1, 2 and 4), which is secreted even at normal expression levels.29, 38, 39
In fact, all FGE variants except p.R327X and p.A149_A173del (see below) were able to catalyze a significant substrate turnover in vitro, as determined after isolation from the medium or cell lysates. The highest residual activity (16% of wt) was exhibited by FGE G263V and the lowest by FGE S155P (1.6%). These data, obtained by measuring directly the FGly formation in a sulfatase substrate peptide,23, 29, 40 are in accordance with observations made by Annunziata et al,25 who expressed different MSD causing SUMF1 mutations together with reporter sulfatases in FGE-deficient cells and observed detectable, although reduced sulfatase activities in all cases. Thus, all MSD causing mutations analyzed so far are hypomorphic – at least one per patient. In conclusion, a combination of two null mutations should be lethal. This concept, however, was challenged by our NVS patient, who carried two alleles with nonsense mutations, one of which (p.R327X) definitely disrupting the active site (Supplementary Figure S1), as confirmed in our experiments by the total absence of activity. The other mutation, p.A149_A173del, leads to an internal in-frame deletion of 25 amino acids. Interestingly, this defective FGE at high concentrations of purified protein indeed led to detectable substrate turnover corresponding to a residual activity of 0.3%. Thus, the protein obviously adopts a conformation allowing substrate turnover, although with strong impairments. Notably, the deleted 25 residues (pink colored in Supplementary Figure S1) are located on the surface of the FGE core domain. Residues 161−176 form a loop that is easily cleaved off by proteases.20 The deletion further removes β-strand 5 (residues 153–158, pink), which normally pairs with the adjacent β-strand 6 (residues 180–184, blue) to form an antiparallel two-stranded sheet on the surface of the molecule. The residues forming β-strand 6 are also present in FGE A149_A173del. Upon losing the anti-parallel interaction partner β-strand 5, β-strand 6 is predicted to change at least its orientation, which obviously allows bridging the distance to the deletion breakpoint at position 148 without complete disruption of the catalytic core domain architecture.
Taken together, we found extensive instability for all FGE variants, thereby confirming the predictions from mapping the amino acid substitutions onto the FGE crystal structure. The degradation kinetics, however, varied among the different proteins (Figure 4). Furthermore, we found different secretion properties of FGE variants. About 50% of wt FGE protein is secreted after 6 h. Three impaired FGE proteins, among them the two expressed nonsense mutations, were completely or almost completely (p.S155P) retained inside the cells and degraded. The other three (p.G247R, p.G263V and p.R345C) were partially secreted, two of them (p.G247R and, in particular, p.G263V) even much more rapidly than wt FGE (Figure 4c). As expression was comparable or even reduced, the enhanced secretion is not caused by simple saturation of retention mechanisms inside the cells. Rather the interaction with the retention machinery might be affected, which is compatible with the surface exposure of both glycine residues G247 and G263 (Supplementary Figure S1). For wt FGE, an interaction with ERp44, protein disulfide isomerase and ERGIC-53 is already known; these components act in different ER and Golgi subcompartments.21, 38 The exact way of retention of defective FGE variants as well as their delivery for degradation is still unknown. As secretion of wt FGE was described also at physiological expression levels,39 we conclude that impaired retention contributes to disease severity in patients with p.G247R and p.G263V mutations. Specific interactions with impact on the degradation and secretion kinetics may be affected. Whatever mechanism applies, impaired retention in addition to impaired stability, observed for all FGE variants tested, both lead to reduced intracellular levels of FGE.
Cellular phenotype in MSD fibroblasts: impairments of endogenous FGE and sulfatases
The consequences of impaired FGE structure and function, as observed in overexpressing cells, could be confirmed in patient fibroblasts. FGE was hardly detectable here, probably because of degradation. Also no intracellular mislocalization was visible. The activities of endogenous sulfatases were differentially reduced, roughly mirroring residual FGE functionality. Lowest activities were found in fibroblasts combining the two FGE nonsense/deletion mutations, that is, in the NVS case. Highest sulfatase activities were found in LIM fibroblasts homozygous for the p.G263V mutation leading to FGE that is unstable but shows considerable residual activity (as high as 16%). This observation confirms our previous results of high sulfatase activities in fibroblasts expressing unstable mutant FGE with high residual activity.13
In LIS fibroblasts residual sulfatase activities can vary considerably. Residual ASC activities in the three p.S155P patients reached 14–32%, whereas only 0.3–3% were measured for the other three patients of this phenotypic group. Interestingly, the variability was clearly less pronounced for the other two sulfatases tested. This might indicate that individual sulfatases exhibit different affinities to the various compromised FGE proteins. The resulting differential residual activities of specific sulfatases, however, can not be correlated with the onset of specific clinical symptoms during manifestation of the MSD syndrome. Nevertheless, the severity of the disease is reflected by the total residual sulfatase activity (Table 3).
Perspective: genotype/phenotype correlation and clinical predictions in MSD patients
Comparing the biochemical impairments of FGE caused by individual SUMF1 mutations with the clinical phenotype of the corresponding MSD patients, we find clear correlations (see Table 3). Patients with drastic impairments of both FGE stability and residual enzyme activity displayed the most severe clinical phenotype. In our present study, this was the case for the NVS MSD patient. The mildest phenotype presented in the patient with FGE G263V, which is associated with the highest residual FGE activity among the studied variants; further FGE G263V, though present at only rather low intracellular levels, can be assumed to truly perform this residual activity, as it efficiently passed intracellular quality control mechanisms.
Taking all available molecular, biochemical and clinical data into account it is now possible to establish a genotype/phenotype correlation for MSD and to roughly predict the clinical course for patients with the studied SUMF1 mutations. Missense mutations affecting crucial functional or structural residues in FGE, as well as nonsense mutations, will cause severe forms of the disease, whereas missense mutations not fully abrogating a functional conformation of the FGE protein will lead to attenuated forms. This does not exclude that other factors (mostly putative as yet) act as disease modifier, for example, proteins interacting with FGE for ER retention, intracellular trafficking and degradation, redox protection, substrate presentation or cofactor regeneration as well as unknown factors regulating expression levels of FGE (and sulfatases). Such factors can be expected to have an impact on the clinical presentation of patients.41, 42, 43 Identification and characterization of these interacting factors as well as understanding their role in the molecular and cell biological functioning of FGE will be the basis for designing therapeutic targets. Likely, these targets will be specific for different groups of mutations with similar functional properties. Clear hints toward the existence of such mutant groups come from the observation that despite early degradation of most protein variants, some of them, at comparable expression levels, are rapidly secreted, thereby escaping quality control. Beside enzyme replacement and gene therapy approaches,10, 39 a promising therapeutic approach would thus be to enhance the stability and/or the intracellular retention, and in turn the total residual FGE activity inside the ER, which represents the bottleneck in MSD patients. The use of small molecule chaperones, meanwhile tested in the therapy of some inborn errors of metabolism, are therefore likely to be beneficial also in MSD.44, 45
References
Hopwood J, Ballabio A : Multiple sulfatase deficiency and the nature of the sulfatase family. in Scriver CR BA, Valle D, Sly WS (eds): The Metabolic and Molecular Bases of Inherited Disease. New York: McGraw-Hill, 2001, pp 3725–3732.
Nyhan WL, Ozand PT : Atlas of Metabolic Diseases. London: Chapman and Hall Medical, 1998, pp 614–621.
Van der Knaap M, Valk J : Magnetic Resonance of Myelination and Myelin Disorders, 3 edn, Berlin, Heidelberg, New York: Springer, 2005, pp 82–86.
Eto Y, Gomibuchi I, Umezawa F, Tsuda T : Pathochemistry, pathogenesis and enzyme replacement in multiple-sulfatase deficiency. Enzyme 1987; 38: 273–279.
Perlmutter-Cremer N, Libert J, Vamos E, Spehl M, Liebaers I : Unusual early manifestation of multiple sulfatase deficiency. Ann Radiol (Paris) 1981; 24: 43–48.
Vamos E, Liebaers I, Bousard N, Libert J, Perlmutter N : Multiple sulphatase deficiency with early onset. J Inherit Metab Dis 1981; 4: 103–104.
Burch M, Fensom AH, Jackson M, Pitts-Tucker T, Congdon PJ : Multiple sulphatase deficiency presenting at birth. Clin Genet 1986; 30: 409–415.
Busche A, Hennermann JB, Burger F et al: Neonatal manifestation of multiple sulfatase deficiency. Eur J Pediatr 2009; 168: 969–973.
Diaz-Font A, Santamaria R, Cozar M et al: Clinical and mutational characterization of three patients with multiple sulfatase deficiency: report of a new splicing mutation. Mol Genet Metab 2005; 86: 206–211.
Dierks T, Schlotawa L, Frese MA, Radhakrishnan K, von Figura K, Schmidt B : Molecular basis of multiple sulfatase deficiency, mucolipidosis II/III and Niemann-Pick C1 disease – Lysosomal storage disorders caused by defects of non-lysosomal proteins. Biochim Biophys Acta 2009; 1793: 710–725.
Mancini GM, van Diggelen OP, Huijmans JG, Stroink H, de Coo RF : Pitfalls in the diagnosis of multiple sulfatase deficiency. Neuropediatrics 2001; 32: 38–40.
Loffeld A, Gray RG, Green SH, Roper HP, Moss C : Mild ichthyosis in a 4-year-old boy with multiple sulphatase deficiency. British J Dermatol 2002; 147: 353–355.
Schlotawa L, Steinfeld R, von Figura K, Dierks T, Gartner J : Molecular analysis of SUMF1 mutations: stability and residual activity of mutant formylglycine-generating enzyme determine disease severity in multiple sulfatase deficiency. Hum Mutat 2008; 29: 205.
al Aqeel A, Ozand PT, Brismar J et al: Saudi variant of multiple sulfatase deficiency. J Child Neurol 1992; 7 (Suppl): S12–S21.
Blanco-Aguirre ME, Kofman-Alfaro SH, Rivera-Vega MR et al: Unusual clinical presentation in two cases of multiple sulfatase deficiency. Pediatr Dermatol 2001; 18: 388–392.
Diez-Roux G, Ballabio A : Sulfatases and human disease. Annu Rev Genomics Hum Genet 2005; 6: 355–379.
Dierks T, Schmidt B, von Figura K : Conversion of cysteine to formylglycine: a protein modification in the endoplasmic reticulum. Proc Natl Acad Sci USA 1997; 94: 11963–11968.
Schmidt B, Selmer T, Ingendoh A, von Figura K : A novel amino acid modification in sulfatases that is defective in multiple sulfatase deficiency. Cell 1995; 82: 271–278.
Dierks T, Lecca MR, Schlotterhose P, Schmidt B, von Figura K : Sequence determinants directing conversion of cysteine to formylglycine in eukaryotic sulfatases. EMBO J 1999; 18: 2084–2091.
Dierks T, Dickmanns A, Preusser-Kunze A et al: Molecular basis for multiple sulfatase deficiency and mechanism for formylglycine generation of the human formylglycine-generating enzyme. Cell 2005; 121: 541–552.
Mariappan M, Radhakrishnan K, Dierks T, Schmidt B, von Figura K : ERp44 mediates a thiol-independent retention of formylglycine-generating enzyme in the endoplasmic reticulum. J Biol Chem 2008; 283: 6375–6383.
Cosma MP, Pepe S, Annunziata I et al: The multiple sulfatase deficiency gene encodes an essential and limiting factor for the activity of sulfatases. Cell 2003; 113: 445–456.
Dierks T, Schmidt B, Borissenko LV et al: Multiple sulfatase deficiency is caused by mutations in the gene encoding the human C(alpha)-formylglycine generating enzyme. Cell 2003; 113: 435–444.
Settembre C, Annunziata I, Spampanato C et al: Systemic inflammation and neurodegeneration in a mouse model of multiple sulfatase deficiency. Proc Natl Acad Sci USA 2007; 104: 4506–4511.
Annunziata I, Bouche V, Lombardi A, Settembre C, Ballabio A : Multiple sulfatase deficiency is due to hypomorphic mutations of the SUMF1 gene. Hum Mutat 2007; 28: 928.
Cosma MP, Pepe S, Parenti G et al: Molecular and functional analysis of SUMF1 mutations in multiple sulfatase deficiency. Hum Mutat 2004; 23: 576–581.
Artigalas OA, da Silva LR, Burin M et al: Multiple sulfatase deficiency: clinical report and description of two novel mutations in a Brazilian patient. Metabolic brain disease 2009; 24: 493–500.
Steinmann B, Mieth D, Gitzelmann R : A newly recognized cause of low urinary estriol in pregnancy: multiple sulfatase deficiency of the fetus. Gynecol Obstet Invest 1981; 12: 107–109.
Preusser-Kunze A, Mariappan M, Schmidt B et al: Molecular characterization of the human Calpha-formylglycine-generating enzyme. J Biol Chem 2005; 280: 14900–14910.
Rommerskirch W, von Figura K : Multiple sulfatase deficiency: catalytically inactive sulfatases are expressed from retrovirally introduced sulfatase cDNAs. Proc Natl Acad Sci USA 1992; 89: 2561–2565.
Voznyi YV, Keulemans JL, van Diggelen OP : A fluorimetric enzyme assay for the diagnosis of II MPS (Hunter disease). J Inherit Metab Dis 2001; 24: 675–680.
Gieselmann V, Schmidt B, von Figura K : In vitro mutagenesis of potential N-glycosylation sites of arylsulfatase. A effects on glycosylation, phosphorylation, and intracellular sorting. J Biol Chem 1992; 267: 13262–13266.
Mariappan M, Gande SL, Radhakrishnan K, Schmidt B, Dierks T, von Figura K : The non-catalytic N-terminal extension of formylglycine-generating enzyme is required for its biological activity and retention in the endoplasmic reticulum. J Biol Chem 2008; 283: 11556–11564.
Roeser D, Preusser-Kunze A, Schmidt B et al: A general binding mechanism for all human sulfatases by the formylglycine-generating enzyme. Proc Natl Acad Sci USA 2006; 103: 81–86.
Fey J, Balleininger M, Borissenko LV, Schmidt B, von Figura K, Dierks T : Characterization of posttranslational formylglycine formation by luminal components of the endoplasmic reticulum. J Biol Chem 2001; 276: 47021–47028.
Vembar SS, Brodsky JL : One step at a time: endoplasmic reticulum-associated degradation. Nat Rev Mol Cell Biol 2008; 9: 944–957.
Christis C, Lubsen NH, Braakman I : Protein folding includes oligomerization – examples from the endoplasmic reticulum and cytosol. FEBS J 2008; 275: 4700–4727.
Fraldi A, Zito E, Annunziata F et al: Multistep, sequential control of the trafficking and function of the multiple sulfatase deficiency gene product, SUMF1 by PDI, ERGIC-53 and ERp44. Hum Mol Genet 2008; 17: 2610–2621.
Zito E, Buono M, Pepe S et al: Sulfatase modifying factor 1 trafficking through the cells: from endoplasmic reticulum to the endoplasmic reticulum. EMBO J 2007; 26: 2443–2453.
Peng J, Schmidt B, von Figura K, Dierks T : Identification of formylglycine in sulfatases by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. J Mass Spectrom 2003; 38: 80–86.
Conary JT, Hasilik A, von Figura K : Synthesis and stability of steroid sulfatase in fibroblasts from multiple sulfatase deficiency. Biol Chem Hoppe Seyler 1988; 369: 297–302.
Steckel F, Hasilik A, von Figura K : Multiple sulfatase deficiency: degradation of arylsulfatase A and B after endocytosis in fibroblasts. Eur J Biochem 1985; 151: 147–152.
Steckel F, Hasilik A, von Figura K : Synthesis and stability of arylsulfatase A and B in fibroblasts from multiple sulfatase deficiency. Eur J Biochem 1985; 151: 141–145.
Sawkar AR, Adamski-Werner SL, Cheng WC et al: Gaucher disease-associated glucocerebrosidases show mutation-dependent chemical chaperoning profiles. Chem Biol 2005; 12: 1235–1244.
Ulloa-Aguirre A, Janovick JA, Brothers SP, Conn PM : Pharmacologic rescue of conformationally-defective proteins: implications for the treatment of human disease. Traffic 2004; 5: 821–837.
Acknowledgements
We would like to thank N Hollstein, N Eiselt, A Winter, M Balleininger, J Kaiser, T Wilke, M-A Frese and E Wiegmann for excellent technical assistance. We are grateful to M Henneke, H Rosewich, P Huppke, A Ohlenbusch, M Mariappan and B Mueksch for helpful discussions and K von Figura for his support to implement the project. This work was supported by the Research program of the Faculty of Medicine, Georg August University Göttingen (LS), by the Deutsche Forschungsgemeinschaft, the Fonds der Chemischen Industrie and Shire Human Genetic Therapies, Inc. (Cambridge, MA, USA).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on European Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Schlotawa, L., Ennemann, E., Radhakrishnan, K. et al. SUMF1 mutations affecting stability and activity of formylglycine generating enzyme predict clinical outcome in multiple sulfatase deficiency. Eur J Hum Genet 19, 253–261 (2011). https://doi.org/10.1038/ejhg.2010.219
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2010.219
Keywords
This article is cited by
-
Late infantile form of multiple sulfatase deficiency with a novel missense variant in the SUMF1 gene: case report and review
BMC Pediatrics (2023)
-
Association between SUMF1 polymorphisms and COVID-19 severity
BMC Genomic Data (2023)
-
Expression, activity and localization of lysosomal sulfatases in Chronic Obstructive Pulmonary Disease
Scientific Reports (2019)
-
Genome-wide expression analysis reveals six contravened targets of EZH2 associated with breast cancer patient survival
Scientific Reports (2019)
-
Sulfatase modifying factor 1 (SUMF1) is associated with Chronic Obstructive Pulmonary Disease
Respiratory Research (2017)