Abstract
About 40% of the genetic variance of age-related macular degeneration (AMD) can be explained by a common variation at five common single-nucleotide polymorphisms (SNPs). We evaluated the degree to which these known variants explain the clustering of AMD in a group of densely affected families. We sought to determine whether the actual number of risk alleles at the five variants in densely affected families matched the expected number. Using data from 322 families with AMD, we used a simulation strategy to generate comparison groups of families and determined whether their genetic profile at the known AMD risk loci differed from the observed genetic profile, given the density of disease observed. Overall, the genotypic loads for the five SNPs in the families did not deviate significantly from the genotypic loads predicted by the simulation. However, for a subset of densely affected families, the mean genotypic load in the families was significantly lower than the expected load determined from the simulation. Given that these densely affected families may harbor rare, more penetrant variants for AMD, linkage analyses and resequencing targeting these families may be an effective approach to finding additional implicated genes.
Similar content being viewed by others
Introduction
Age-related macular degeneration (AMD) is the leading cause of blindness among older individuals in developed countries.1 In the past 5 years, several common variants have been reliably and reproducibly associated with AMD.2, 3, 4, 5, 6, 7, 8, 9, 10 In a case–control study drawn from a US-based population of European descent, we found a previously unrecognized, noncoding variant in complement factor H (CFH) and replicated four previously reported alleles associated with AMD: a common coding variant in CFH (rs1061170), LOC387715 (rs10490924), and two variants in the complement component 2 and factor B (C2-CFB) locus.10 Genotypes at these five common single-nucleotide polymorphisms (SNPs) define a broad spectrum of interindividual disease risk and explain about 40% of the classical sibling risk of AMD in the study population.10
As we are now able to explain such a large fraction of the genetic variability of this disease, we were interested in evaluating the degree to which the existing variants explain the clustering of AMD in a group of densely affected families. This information could determine the best approach to finding additional genes implicated in AMD. If densely affected families had less than the expected genetic load at the five known common variants, it would suggest that there were rarer, more penetrant variants in these families to explain their burden of disease. Linkage analysis of the families would then be one of the next steps to finding new variants associated with AMD. If the families had the expected genetic load, it would suggest that additional genetic variants may be similar to those found: common, additive, and with a comparable or smaller magnitude of effect individually. These would be best pursued by whole-genome association studies. To determine whether the actual number of risk alleles at the five variants in our families matched the expected number, we developed a simulated population of families on the basis of the assumption that a fixed proportion of AMD familial risk is based on the five known variants. In this study, we explain a novel analytical technique to develop such a comparison group and its application to determine whether densely affected families with AMD have a genetic profile at the known AMD risk loci that differs from their expected genetic profile, given the density of disease observed.
Materials and methods
Family ascertainment
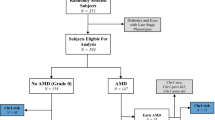
The methods used in this study conformed to the tenets of the Declaration of Helsinki and received approval from appropriate institutional review boards. All participants gave informed consent and signed the relevant forms. Potentially eligible probands with advanced AMD in at least one eye were recruited from several sources as previously described, including from the AMD database of the principal investigator (JMS), from other ophthalmologists throughout the country, from some of the clinical centers participating in a multicenter randomized trial of vitamins and minerals (Age-Related Eye Disease Study, or AREDS), and through various media, such as newsletters, brochures, and newspapers.11 Advanced AMD was defined as either geographic atrophy (grade 4) or neovascular disease (grade 5) as determined by fundus photography and ocular examination using the clinical age-related maculopathy grading system (CARMS) criteria.12 After verifying that the proband had advanced disease, family members of the proband were then asked to participate. Some families were recruited initially as sibling pairs for the purposes of a linkage study, and large families with multiple affected members, as well as discordant siblings, were also recruited. Data collection procedures included standard stereo pair 30-degree ocular fundus photography, blood drawing, and questionnaires regarding risk factors for AMD. AMD status was assigned by CARMS criteria in family members as well.
Genotyping
A total of 1265 probands and siblings from 322 families with a proband with advanced AMD were genotyped. For 67 of these families, we were able to obtain phenotypes and genotypes for all probands' siblings. For the remainder of the families, either phenotype or genotype for one or more of the siblings was missing due to death, illness, or other reasons for nonparticipation. Genotyping was performed for the following five SNPs that were previously found to be associated with AMD: CFH (rs1061170 and rs1410996), C2-CFB (rs641153 and rs9332739), and LOC387715 (rs10490924).9, 10 Genotyping was performed on Illumina BeadArray (San Diego, CA, USA) and Sequenom (San Diego, CA, USA) platforms at the Broad Institute Center for Genotyping and Analysis. The process for Illumina and Sequenom genotyping is found at their respective websites (http://www.illumina.com/technolology/tech_overview.ilmn and http://www.sequenom.com/applications/hme_assay.php). One individual (typically the proband) from each of the families was used in our previous case–control association study.10 These individuals were genotyped using the Illumina platform for all variants, except the C2 and CFB variants, which were genotyped on the Sequenom platform. All other individuals not genotyped as part of the association study were genotyped on the Sequenom platform for all the variants. For all genotyping, several duplicates were added to each sample plate for quality assurance and quality control validation of interplate discordance. The data completion rate was 98.4% for individuals who were part of the initial case–control study and 95.2% for the subsequent Sequenom genotyping.
Simulation of comparison family population
We used the statistical package R for the creation of a comparison population of families (www.R-project.org). We simulated phenotypes in the offspring of two-generation families assuming the heritability, environmental variance of AMD and allele frequencies of the five known risk SNPs (Supplementary Table 1). To accomplish this, we first simulated genotypes for a population of parents on the basis of the known European (CEU) population allele frequencies from the HapMap database (www.hapmap.org). We ‘mated’ parents to create families with a varying number of siblings; the offspring's genotypes at the five loci were assigned according to random transmission at meiosis.
We used a liability threshold model to assign phenotypes (CARMS grade 4 or 5 AMD vs others) to all probands and siblings. In this model, we assume an underlying normal distribution of liability for developing a disease.13 Above a certain threshold of liability, all individuals are affected with advanced AMD (grade 4 or 5). The total liability score for each individual is the sum of the known genetic effects from the five risk variants, the residual polygenic effects (which correlate across each family), and the specific environmental effects. To convert the effects on the risk scale into the effect size on the liability scale, we invoked the Fisherian biometrical model. To generate an estimate of the effect size, we assumed that the liability distribution had a mean shift, such that the percentage of the liability distribution above the fixed threshold for affection (taken as the inversion of the normal at the prevalence) matches the expectation based on the odds ratio from the primary analysis.
The magnitude of the known genetic effects was obtained from estimates of the effects for the five loci as determined in our association study.10 Figure 1 shows the estimated prevalence of advanced AMD in the general population according to genotypic load as derived from our previous case–control study.10 Our subsequent simulation is based on this model, which assumes that the prevalence of disease increases as the number of risk genotypes for the five known SNPs increases. Thirty-six genotype bins are defined by the six possible genotypes for the two CFH variants, the three possible genotypes for the LOC387715 variant, and the two possible genotypes at the C2-CFB locus. Owing to the low frequency and nearly identical risk of the protective alleles at C2-CFB, we considered individual risk at C2-CFB in two categories: high risk (individuals carrying neither protective variant) and low risk (individuals carrying one or two protective variants). We estimated that the five common variants contribute about 29% of the total variance in AMD, or about 40% of the total additive genetic variance on the liability scale.10
Estimated prevalence of advanced AMD cases as a function of genetic load. Using genotyping data from a case–control study of AMD, 36 risk categories were determined on the basis of genotypes at the five common risk loci.10 The estimated risk of developing advanced AMD for each of these risk categories was calculated. These risk categories are sorted by the estimated risk of advanced AMD. The table below the figure shows the specific genotypes for each bin. For CFH, the L/M/H designations correspond to low-, medium- and high-risk haplotypes. The LOC387715 L/H designation corresponds to low- and high-risk alleles. For C2/BF, the L or H designation corresponds to low- or high-risk genotypes.
The residual background polygenic variance is obtained by subtracting the known genetic variance from total heritability. The estimated heritability for advanced AMD is ∼70%.14 The residual background polygenic variance is therefore ∼41% on the liability scale. For polygenicity, we estimated the amount of heritability that was conferred by the loci, and reduced the heritable component of the phenotype accordingly. With this residual heritable component, we can then draw random normals and multiply by the square root of the residual variance, as that will provide a liability score that will contribute to the appropriate amount of variance (that is, var(k*X)=k^2*var(X), where k is a constant and X is a randomly drawn variable). In simulating the residual polygenic component, we assumed a multivariate, normal model with a correlation of 0.5 between siblings and between parents and their offspring. Thus, half of the variance of the residual polygenic component was between families and half was within families.
The remaining 30% of the variance in AMD is attributable to nongenetic factors (that is, environment and chance).14 The liability score was then transformed into diagnosis of advanced AMD if the score fell above a threshold determined by the prevalence of advanced AMD in persons 75–79 years of age, or ∼5%,1, 15 as the individuals in our family study cohort had a median age of 79 years and a mean age of 77.7 years.
As we recognize that rare Mendelian forms of AMD could also exist in a population of densely affected families, we repeated the simulation described above with the addition of a risk allele with a 0.1% frequency and an odds ratio for advanced AMD of 20. We determined the distribution of slopes of mean genotypic load from the actual families vs the mean genotypic load from this additional simulation with a rare, penetrant allele.
Comparison of actual families to simulated families
Once we had a simulated population of sibships, with each sibling assigned a phenotype, we randomly selected families who had at least one sibling with advanced AMD, the ‘proband.’ For each of the simulated families and the actual families, we calculated a genotypic load, a measure of the number of risk alleles present in the five known genetic variants. The average genotypic load was determined for families that shared the same number of affected siblings and total siblings; this was carried out separately for simulated and actual families. Empirical P-values were calculated on the basis of simulation results. We determined individual simulation family deviations from the average genotypic load for simulation families of the same configuration and then recorded the observed deviation from the mean score of actual families with the same configuration. We calculated significance by counting the number of instances in which the simulated family was more deviant that the actual family and divided this by the total number of simulations.
In addition, the distribution of the individual families' genotypic loads was plotted against their expected mean genotypic loads from the simulation. The slope of the line for these data points was determined. To determine whether this slope was significantly different from the expected slope, we determined the slopes for genotypic loads from each set of 322 simulation families plotted against the mean genotypic loads derived from all simulation sets. We also calculated 2.5 and 97.5% confidence intervals for the mean genotypic load for each family configuration. These confidence bounds were based on individual simulated families. In all, 100 000 simulated families in each configuration were generated and sorted by their mean genotypic load. The 2500th and 97500th averages were set as the confidence intervals for each configuration. The slope of the line of individual families was determined.
For families who had one or more siblings missing either genotype or both genotype and phenotype information (due to death, illness, and so on), siblings with missing information were excluded from the analysis. To ensure the validity of this approach, genetic load distribution was determined separately for the set of 65 families in which all siblings had complete phenotype and genotype information and for the 257 families in which genotype and/or phenotype information was unavailable on all siblings.
Results
Figure 2 shows the distribution of advanced AMD by genotype score in all 65 fully genotyped and phenotyped families (Figure 2a) and in the other 257 families who did not have all siblings phenotyped and/or genotyped (Figure 2b). The distribution of genotypic load was not different to a statistically significant degree between the two groups (P=0.11, Kolmogorov–Smirnov test). The lack of difference between the two distributions suggests that combining these two groups into one comparison group for simulated families is valid.
Distribution of mean genotypic load among families. We calculated the mean familial genotypic load for individuals in the 65 families who have genotype and phenotype information available on all siblings and in the 257 families who did not have genotype and/or phenotype information available on all siblings. The number of individuals in each genotypic load category is shown here for the 65 (a) and 257 (b) families. There was no statistically significant difference between these two distributions (P=0.11, χ2-test).
For the primary analysis, we simulated 5000 sets of 322 families. Table 1 shows the proportion of expected (based on simulated families) and observed (based on actual families) affected siblings in families of different sizes. For most family configurations, there was no statistically significant difference between the actual average genotypic load and the average genotypic load from the simulation. There was, however, a statistically significant difference in some of the larger, more densely affected families and in the 2-, 3- and 4-sibling families with only one affected sibling. For these less-densely affected families, the average genotypic load was higher than that expected by simulation (P=0.015, P=0.009 and P=0.004, respectively). It was also higher in the more densely affected families with three out of five and four out of six affected family members (P=0.020 and P=0.048, respectively). The actual average genotypic load was lower than that expected by simulation in families with four out of four and four out of five siblings affected (P=0.05 and P=0.020, respectively).
Figure 3 shows the distribution of mean genotypic loads for individual families in relation to the expected average genotypic load from the simulation. Most of the families' genotypic loads fall within the 95% confidence interval. There are 28 outlying families, which is slightly higher than what would be predicted by chance alone. The families that fall below the 2.5% confidence interval include two families from the four out of four configuration and one family from the four out of five configuration. In Figure 3, the slope of the line for the individual families' genotypic loads vs mean genotypic loads, by family configuration, is 0.4594. Figure 4a shows the distribution of the slopes when the genotypic loads from each set of simulated families are plotted against the mean genotypic loads averaged among all sets of simulated families. The slope for our actual families is smaller than that expected from simulation (slope=1) to a statistically significant degree (P<0.001). This reflects the fact that the observed genotypic load for some of our families is less than the expected genotypic load from the simulation.
Observed vs expected genotypic load among families. The mean genotypic load for each family was plotted against the mean genotypic load for a family of the same configuration, as determined by simulation. Each dot represents one family. Triangles mark the lower 2.5% confidence bound, whereas diamonds mark the upper 97.5% confidence bound. The slope of the line for the individual families' genotypic loads is 0.4594.
(a) Distribution of slopes for plots of mean genotypic load across all simulation families vs individual families' genotypic loads from each simulation subset of 322 families, by family configuration. The arrow indicates the location of the slope from Figure 3 within the tail of this distribution. (b) Distribution of slopes for plots of mean genotypic load across all simulation families, in which the simulation included a rare, high-penetrance allele, vs genotypic loads from each simulation subset of 322 families, by family configuration. Please note the difference in the x axis scale between panels a and b. The distribution of values in panel b is shifted to values lower than that of panel a.
Discussion
In this study, the majority of densely affected families had average genotypic loads for the five common SNPs associated with AMD that would be expected on the basis of a simulation of genotypic load. This assumes that the five variants account for the known familial variance in AMD.
The ability of simulation to closely mirror the actual average genotype distribution also supports the validity of this novel analytical approach and its underlying assumptions regarding allele frequency, disease prevalence, and estimates of the effect sizes of individual risk SNPs. As gene discovery proceeds for other common diseases to the point at which a significant amount of familial variance can be explained by known genetic variation, this approach may be useful to try to understand the genetic architecture of undiscovered familial variance. This could provide a basis on which to decide whether to pursue additional common additive variants or rarer, more penetrant variants. It may also serve to identify families who are particularly likely to harbor additional, undiscovered variants. On the basis of this approach alone, however, we cannot distinguish a priori for a specific family whether the remainder of the unexplained genetic variance for AMD is because of highly penetrant loci or because of multiple common ones.
Although the genotypic load for most of our AMD families did not deviate significantly from the expected load, we did identify some families, particularly in the four out of four and four out of five affected configurations, who had a lower than expected genotypic load. There are several explanations that could explain this finding. There may be rare, more penetrant genetic variants in these families contributing to their disease. To further explore this possibility, we also performed a simulation that included an additional rare, penetrant risk allele. We show that the distribution of slopes of the mean genotypic load from the actual families vs the mean genotypic load from this additional simulation is shifted to lower values (Figure 4b). This is because, under these conditions, a family with a particular disease density now has a possible explanation for their increased burden of disease that does not involve the five common SNPs. If our genotypic load score recognized and accounted for the existence of this rare allele, the median slope in the distribution of slopes would once again be 1. Therefore, our current genotypic load, which only takes into account the five known common SNPs, is incomplete in its ability to explain some of the disease burden in densely affected families. One possible explanation for the difference between the observed vs simulated genotypic loads is an undiscovered, rare, high-penetrance variant.
Alternatively, these families could also share a common environmental risk factor so that the density of disease is not explained on a genetic basis. This is less likely, given that environmental risk factors were well documented as part of the selection of these families, and no strong, common, environmental risk factor that can explain these differences has been identified in these families. A third possibility is that the AMD in these families is a distinct subphenotype of AMD, with an underlying genetic architecture that is different from the AMD in the general population. However, this is unlikely, as the phenotypes observed represent the typical phenotypes seen in other AMD populations, and all families had a proband with advanced AMD. To further explore the possibility that environmental risk factors or subphenotypes could account for our findings, we compared environmental risk factors (smoking and body mass index)16, 17 and the two main subtypes of AMD, advanced dry (geographic atrophy) and wet (choroidal neovascularization) AMD, in families whose genotypic load was as expected and in families with a lower than expected genotypic load (Table 2). There is no statistically significant difference between these two groups for any of these variables. A final possibility for explaining our findings is that the assumption of a liability model or the assumption of additivity of the known loci and the polygenic portion on the liability scale may be incorrect.
It should be noted that our method of selecting densely affected families is subject to ascertainment bias, because some siblings were selected as affected sibling pairs for a linkage analysis. Therefore, to design a study that was independent of the ascertainment scheme strategy, we adopted an ascertainment-free analysis approach by considering the mean genetic load according to family configuration.
There are some limitations to our study. Genotype and/or phenotype information was unavailable on some siblings primarily because of illness or death of the sibling. There is a possibility of misclassification bias. Some unaffected siblings were ascertained at a younger age than their affected proband sibling. We did, however, follow all siblings prospectively to obtain the most recent AMD grade used in this study, and all subjects had a grade assigned after an age of 60 years. It is still possible that a few of these siblings could develop AMD or progress with time. We have limited numbers of densely affected families for some of the family configurations. This limits our power to detect true differences between actual vs simulated families. Finally, we recognize that some of the reported P-values are marginal and may be due to chance.
Despite these limitations, we believe that the novel analytic approach used in this study is a valid way to determine the expected genotypic load for densely affected families for diseases in which a significant proportion of the genetic variance is already known. In our AMD families, this method allowed us to identify a subset of densely affected families who have a lower than expected genotypic load. Given that these families may harbor rare and more penetrant variants for AMD, linkage analyses targeting these families would be one way to search for potential additional implicated genes. Resequencing of the known associated genes could reveal additional implicated variants.
References
Friedman DS, O'Colmain BJ, Munoz B et al: Prevalence of age-related macular degeneration in the United States. Arch Ophthalmol 2004; 122: 564–572.
Montezuma SR, Sobrin L, Seddon JM : Review of genetics in age related macular degeneration. Semin Ophthalmol 2007; 22: 229–240.
Klein RJ, Zeiss C, Chew EY et al: Complement factor H polymorphism in age-related macular degeneration. Science 2005; 308: 385–389.
Edwards AO, Ritter III R, Abel KJ, Manning A, Panhuysen C, Farrer LA : Complement factor H polymorphism and age-related macular degeneration. Science 2005; 308: 421–424.
Gold B, Merriam JE, Zernant J et al: Variation in factor B (BF) and complement component 2 (C2) genes is associated with age-related macular degeneration. Nat Genet 2006; 38: 458–462.
Hageman GS, Anderson DH, Johnson LV et al: A common haplotype in the complement regulatory gene factor H (HF1/CFH) predisposes individuals to age-related macular degeneration. Proc Natl Acad Sci USA 2005; 102: 7227–7232.
Haines JL, Hauser MA, Schmidt S et al: Complement factor H variant increases the risk of age-related macular degeneration. Science 2005; 308: 419–421.
Rivera A, Fisher SA, Fritsche LG et al: Hypothetical LOC387715 is a second major susceptibility gene for age-related macular degeneration, contributing independently of complement factor H to disease risk. Hum Mol Genet 2005; 14: 3227–3236.
Haddad S, Chen CA, Santangelo SL, Seddon JM : The genetics of age-related macular degeneration: a review of progress to date. Surv Ophthalmol 2006; 51: 316–363.
Maller J, George S, Purcell S et al: Common variation in three genes, including a noncoding variant in CFH, strongly influences risk of age-related macular degeneration. Nat Genet 2006; 38: 1055–1059.
Seddon JM, Santangelo SL, Book K, Chong S, Cote J : A genomewide scan for age-related macular degeneration provides evidence for linkage to several chromosomal regions. Am J Hum Genet 2003; 73: 780–790.
Seddon JM, Sharma S, Adelman RA : Evaluation of the clinical age-related maculopathy staging system. Ophthalmology 2006; 113: 260–266.
Pearson KLA : On the inheritance of characteristics not capable of exact quantitative measurement. Philos Trans R Soc Lond A 1901; 195: 79–150.
Seddon JM, Cote J, Page WF, Aggen SH, Neale MC : The US twin study of age-related macular degeneration: relative roles of genetic and environmental influences. Arch Ophthalmol 2005; 123: 321–327.
Klein R, Peto T, Bird A, Vannewkirk MR : The epidemiology of age-related macular degeneration. Am J Ophthalmol 2004; 137: 486–495.
Seddon JM, Ajani UA, Sperduto RD et al: Dietary carotenoids, vitamins A, C, and E, and advanced age-related macular degeneration. JAMA 1994; 272: 1413–1420.
Seddon JM, Cote J, Davis N, Rosner B : Progression of age-related macular degeneration: association with body mass index, waist circumference, and waist-hip ratio. Arch Ophthalmol 2003; 121: 785–792.
Acknowledgements
This research was supported by EY16335-02 (National Eye Institute K12 Harvard Vision Clinician Scientist Development Award); EY11309 from the US National Institutes of Health (National Eye Institute RO1 Grant, JMS) ; the Foundation Fighting Blindness; Massachusetts Lions Research Fund, Inc; the Macular Degeneration Research Fund, Ophthalmic Epidemiology and Genetics Service, Tufts Medical Center; and by the Broad Institute Center for Genotyping and Analysis, supported by Grant U54 RR020278 from the NCRR. LS is a recipient of a Research to Prevent Blindness Career Development Award and JMS was a recipient of the Research to Prevent Blindness Lew R Wasserman Award.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on European Journal of Human Genetics website (http://www.nature.com/ejhg)
Supplementary information
Rights and permissions
About this article
Cite this article
Sobrin, L., Maller, J., Neale, B. et al. Genetic profile for five common variants associated with age-related macular degeneration in densely affected families: a novel analytic approach. Eur J Hum Genet 18, 496–501 (2010). https://doi.org/10.1038/ejhg.2009.185
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2009.185