Abstract
Sami of northern Scandinavia are genetic outliers among European populations and their origin has been difficult to determine. In order to study the genetic origin of the Swedish Sami, we have performed high-resolution typing of the class I HLA-A and -B loci and the class II DRB1, DQB1 and DQA1 loci in the northern and southern Swedish Sami. Several of the common class I alleles in Sami (B*0702, B*1501, B*4002 and A*0301) are found at high frequency in other European populations. However, a number of class I and class II alleles (B*4001, A*2402, DRB1*0901 and DRB1*1101) in the Swedish Sami are characteristic of Asian populations. Admixture analyses indicate that 87% of the Sami gene pool is of European origin and that the Asian contribution is 13%. Our HLA analyses indicate a higher proportion of Asian ancestry in the Sami than shown by previous genetic studies.
Similar content being viewed by others
Introduction
The Sami inhabit the northern parts of Sweden, Finland, Norway and the Kola Peninsula of Russia (Figure 1). Archaeological findings on the west coast of Sweden have been dated to more than 10 200 YBP (years before present),1 but it is unclear whether these represent artefacts from Sami or from a culture that preceded them. The present Scandinavian Sami population is estimated to be less than 100 000 people,2 with the majority residing in Norway.
Since the 1950s, numerous studies have addressed the genetic origin of the Sami.3, 4, 5, 6 Many markers show allele frequency differences between the Sami and non-Sami populations in Sweden, and some markers indicate a similarity with Asian populations.7, 8, 9, 10, 11 In addition, a multilocus analysis has positioned the Sami as outliers among European populations.12 Over 80% of the Sami have one of two mitochondrial DNA (mtDNA) motifs.13 The Sami-specific mtDNA lineage carrying one of those motifs, denoted haplogroup U5b1b1, has been suggested to be of eastern European origin14 but is a subgroup of the more diverse haplogroup U5b1b that is found throughout Europe.14 The second common mtDNA haplogroup in the Sami (haplogroup V) has a likely origin in western Europe and is proposed to have reached Scandinavia via eastern Europe.14, 15 Among the haplogroups that are found at low frequencies in the Sami, both D5 and Z are of Asian origin.14, 16 Three major Y-chromosome haplogroups in the Sami (I, N3 and R1a), accounting for 80% of Sami Y-chromosomes, have been identified. The I and R1a haplogroups are common in many European populations, whereas N3 is common in eastern Europe and northern Asia.14 Accordingly, both mtDNA and Y-chromosome analyses indicate an Asian contribution to the Sami gene pool.
Here, we revisit the hypothesis of a mixed genetic origin of Sami and estimate the contribution of non-European ancestry. In Sweden, there are two separate Sami populations that speak two different languages (‘South Sami’ and ‘North Sami’) out of the 10 discrete languages spoken by different Sami groups in Scandinavia.17 A second aim was to study the genetic similarity of these two populations. In order to study the genetic structure of the Swedish Sami population, we performed high-resolution genetic typing of five HLA class I and class II loci. The human leukocyte antigen (HLA) genes on chromosome 6 (6p21.3) encode polymorphic class I and class II molecules that play a major role in the presentation of foreign antigens to circulating T lymphocytes. The HLA region is characterized by high levels of linkage disequilibrium (LD) between loci and very high levels of polymorphism.18 The polymorphism at HLA loci has been subjected to balancing selection19 but the allele frequency distribution has nevertheless proven to be a valuable tool in genetic studies of human populations. Balancing selection may contribute to the maintenance of new alleles that initially occur at low frequency and may otherwise be easily lost by genetic drift. This may result in a higher number of alleles being maintained in a population and available for tracing the evolutionary history.
Materials and methods
Participants
The participants were from the northern Swedish Sami (County of Norrbotten) and the southern Swedish Sami (County of Västerbotten) (Figure 1). A total of 284 unrelated individuals were genotyped (northern Sami n=154; southern Sami n=130). For comparison, we used HLA data of 252 unrelated non-Sami individuals from the Swedish population.20 This study has been approved by the regional ethics committee.
HLA genotyping
For class I HLA-A and HLA-B loci and the class II DRB1, DQB1 and DQA1 loci, DNA (50 ng) was amplified by PCR using biotinylated primers and hybridized to arrays of immobilized oligonucleotide probes.21, 22 Genotypes were assigned using a pattern recognition algorithm implemented in StripScan version 5.7.1 (Roche Molecular Systems).
Allele and haplotype frequencies
Allele and haplotype frequencies were estimated using Arlequin software.23 HLA data specifically collected for anthropological studies from a worldwide selection of populations and available in dbMHC database (http://www.ncbi.nlm.nih.gov/projects/mhc) and in Allelefrequencies.net database (http://www.allelefrequencies.net) were used for comparison. The assignment of populations to geographic regions refers to the original inhabitants of these regions. Data from the Finnish population was included due to the strategic position of this population. As HLA data from all regions were not available in dbMHC, we used DQB1 and DQA1 allele frequencies from the South Korean population (pop 1) and DQA1 allele frequencies from Urumqi Han (China) to represent the Northeast Asian populations. Both datasets are available from the worldwide population allele frequency database (http://www.allelefrequencies.net). Also, the DRB1–DQB1 haplotype frequencies from the Khoton Tarialan population (Mongolia) were chosen to represent the Northeast Asian populations. The following populations from different geographic regions were used in this study – European populations: Bulgarian, Croatian, Czech, Danish, English, Finnish, French, Georgians, German, Irish, Kurds, non-Sami Swedish, northern Swedish Sami, Polish, Portuguese, Romanian, Russian, southern Swedish Sami, Spanish and Svaneti; Northeast Asian populations: Ami, Atayal, Chukchi, Evenks, Han Chinese, Japanese, Ket, Khalkh, Khanty, Khoton Tarialan, Koryaks, Kushun, Mongolian, South Korean, Thao, Tibetans, Tofalar, Tuva and Yami; Southeast Asian populations: Dai Lue, East Timorese, Fijian, Filipino, Indonesian, Ivatan, Javanese, Malay, Moluccan and Thai; American populations: Athabaskans, Canoncito, Guarani-Kaiowa, Pima and Yupik; Australian population: Yuendumu; African populations: Doggon, Kenyan, Pygmy Biaka, Shona and Zulu.
Statistical and phylogenetic analysis
Population differentiation among samples was tested by analysis of molecular variance (AMOVA)24 and observed and expected Hardy–Weinberg genotypic proportions were compared using an exact test,25 both implemented in the Arlequin population genetics software package.23 MDS analyses (for two dimensions, based on the Euclidian distance matrix computed for pairwise populations using allele frequencies) and hierarchical clustering were performed using MATLAB 7.3.0 (statistic toolbox). Nei's standard genetic distance between populations26 was calculated from the allele frequencies and used to reconstruct neighbor-joining (NJ) trees27 using the PHYLIP package28 and the trees were plotted as networks using TreeView.29
The model for uncorrelated allele frequencies between populations, implemented in STRUCTURE,30 was used to construct four population clusters based on the HLA-A, HLA-B and DRB1 genotypes of 100 randomly sampled individuals from each of 14 populations (when 100 individuals were not available from a population, all individuals were sampled). The fraction of genotypes from a population as well as for each individual of a population belonging to each of the four clusters was then visualized. The proportion of admixture in the Swedish Sami was estimated by LEADMIX31 based on HLA-A, HLA-B and HLA-DRB1 allele frequencies.
Results
Allele, genotype and haplotype frequencies
We compared the allele and haplotype frequencies between the two Sami populations and between Sami and non-Sami populations. There was no significant difference in heterozygosity between the northern and southern Sami populations (Table 1). For all loci, the allele frequencies were significantly different (P<0.05) in pairwise tests among the northern Sami, southern Sami and non-Sami Swedish populations. Fewer class I alleles were found in northern Sami than in southern Sami (Table 1), and southern Sami showed a higher allelic overlap with other European populations than shown by northern Sami (Supplementary material, Supplementary Table S1–S3). Similarly, there were fewer alleles at class II loci in the northern Sami (Table 1). Observed genotype frequencies did not deviate from expected binomial (Hardy–Weinberg) proportions (P>0.05) at any locus in the two Sami populations. The Ewens–Watterson homozygosity test of neutrality32, 33 showed statistically significant negative values for the normal deviate of the homozygosity (Fnd) for DQA1 and DQB1 in both the southern and northern Sami, consistent with balancing selection acting on the HLA polymorphism at these two loci (Table 1). LD (D′) was high for all combinations of loci, with the northern Sami showing higher D′ values than the southern Sami (Table 1).
Population affinities
Several of the alleles (B*0702, B*1501, B*4002, A*0301) that are uncommon in Asian populations (Supplementary Tables S1–S3) showed similar frequencies in northern Sami as in other European populations, consistent with a predominantly European contribution to the Sami gene pool. Other alleles (B*4001, A*2402, DRB1*0901, DRB1*1101) showed similar frequencies between northern Sami and several Asian populations, while they occurred at a lower frequency in other European populations, indicative of an Asian influence in the Swedish Sami. Three of the alleles in the latter group were found on one northern Sami class I–class II haplotype (A*2402–B*4001–DRB1*0901). Finally, for two alleles (DRB1*0801, B*2705) the frequency in northern Sami was much higher than in any other population used for comparison. This pattern may either be due to selection for these alleles in Swedish Sami or genetic drift.
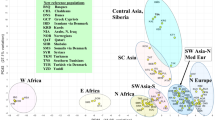
The MDS analysis resulted in stress values of 0.063, 0.091 and 0.077 for the class I, class II and class I+DRB1 data, respectively. In the MDS plots of the class I data from 14 worldwide populations, both Sami populations were located together with other European populations (Figure 2a). For class II loci (18 populations), both Sami populations were located somewhat outside other European populations and in the vicinity of two Siberian populations (Tuva, Khalkh; Figure 2b). In the combined analysis of HLA-A, HLA-B and DRB1 (15 populations), both Sami populations were found close to other European populations (Figure 2c). In MDS plots with only European populations (data not shown), southern Sami clustered with other European populations, whereas northern Sami were located outside this group, consistent with a different genetic contribution to this population.
In the phylogenetic network for class I data from 29 populations, both Sami populations were found in close proximity to other European populations (Figure 3a), whereas for class II loci (35 populations), the southern Sami clustered with the other European populations and the northern Sami were again found in the group with Northeast Asian populations (Figure 3b). Interestingly, the northern Sami were found in between south (Tuva and Mongolian) and north (Ket and Evenk) Siberian populations.
Using the combined HLA-A, HLA-B and HLA-DRB1 allele frequencies of 21 populations, both the northern and southern Sami were found on the same branch, located between Northeast Asian and European populations (Figure 3c).
The STRUCTURE analysis showed African populations mainly belonging to the first cluster (green), the European populations to the second (blue) cluster, the Asian and Australian populations to the third (yellow) cluster, and the Native American populations to the fourth (red) cluster (Figure 4a). The two Sami populations belonged predominantly to the blue (European) cluster and to a lesser degree to the yellow (Asian and Australian) cluster. On the individual level, about 60% of the Sami individuals belonged mainly (more than 90%) to the blue (European) cluster. A few Sami belonged almost entirely to the yellow (Asian) cluster and about 30% of the individuals represented various mixtures of the two clusters (Figure 4b).
Genetic structure of the Sami and a worldwide set of populations. STRUCTURE plots, based on HLA class I and class II data when divided into 4 clusters. The clusters, represented by different colours, are defined on the basis of the allele frequencies in the samples independent of geographic origin of the individuals. In (a) the average values for 14 worldwide populations is shown and in (b) the individuals from each of the populations is shown. The abbreviations used in the figure are: Af – African; Eu – European; NE As – Northeast Asian; SE As – Southeast Asian; Au – Australian; Am – American. The populations include 1 Shona (Zimbabwe, southern Africa), 2 Doggon (Mali, West Africa), 3 Irish (North Ireland, Europe), 4 non-Sami Swedish (Sweden, Europe), 5 southern Sami (Sweden, Europe), 6 northern Sami (Sweden, Europe), 7 Tuva (Russia, Northeast Asia), 8 Korean (South Korea, Northeast Asia), 9 Australian (Cape York, Australia), 10 Filipino (Philippine, Southeast Asia), 11 Atayal (Taiwan, Northeast Asia), 12 Ami (Taiwan, Northeast Asia), 13 Yupik (Alaska, North America), and 14 Guarani-Kaiowa (Brazil, South America).
Admixture analyses
The proportions of European and Asian ancestry were calculated for the northern and southern Sami using (1) the non-Sami Swedish population as descendents of an ancestral European population, (2) the Japanese population as descendents of an ancestral Asian population and (3) the northern and southern Sami populations as descendents of an admixture event between the ancestral European and ancestral Asian populations. Using these assumptions, the proportion of European ancestry in the northern Sami population was estimated to be 0.87 (95% CI: 0.76–0.94) with the remaining 0.13 contributed by Asian populations (Table 2). Using the same populations, the European contribution to the southern Sami was estimated to be 0.92 (95% CI: 0.87–0.97) and the remaining 0.08 Asian input. Similarly, the amount of admixture between the ancestral Sami and non-Sami Swedish populations was calculated using (1) the non-Sami Swedish population as descendents of an ancestral Swedish population, (2) the northern Sami population as descendents of an ancestral Sami population and (3) the southern Sami population as descendents of an admixture event between the ancestral Swedish and the ancestral Sami populations. The ancestral Swedish contribution in the southern Sami was estimated to be 0.58 (95% CI: 0.43–0.68).
LEADMIX also provides estimates relating to the effective population size and time since divergence/admixture of the populations.31 We made the assumptions that (1) an ancestral European and an ancestral Asian population were parental to the Sami, (2) European and Asian populations separated 60 000 years ago34 and (3) the admixture event creating the present Sami population occurred 10 000 years ago.14 We do not have an estimated time point for the putative admixture event between the ancestral Sami and non-Sami Swedish populations. Using the effective population size values from the previous calculation, we set the average effective population size of the non-Sami Swedish population to 100 000 both before and after the admixture event. Based on this, we estimated the average effective population sizes as 8900 (95% CI: 6618–13 910) for the ancestral Sami population, 22 300 for the northern Sami and 18 600 (95% CI: 13 912–26 696) for the southern Sami. The time between the separation of the ancestral European population (into the ancestral Sami and non-Sami populations) to the subsequent admixture event between these two populations, were estimated to be 14 924 (95% CI: 9538–24 472) years. The time point for this admixture event was estimated to be 5256 YBP (95% CI: 40–12 232).
Discussion
The peopling of Scandinavia and origin of the Sami
Archaeological studies have shown that humans were present in the northernmost parts of Europe soon after the end of the last glaciation, about 10 000 YBP. The finding of a western European origin of most mitochondrial haplogroups in Sami, dated at around the time of the last glacial maximum about 18 000 YBP,14, 15 suggests that these haplogroups have arisen in the populations inhabiting small refuges in Europe during that time. The colonization of the European continent after the last glaciation allowed the spread of mitochondrial lineages eastwards, possibly leading to the eastern European origin of some of the Sami mtDNA haplogroups, dated at about 8000 YBP.14, 15 It has been suggested that during the same time period, there was an influence from Mongolian and Finno-Ugric people in eastern Europe, consistent with the finding of mitochondrial haplogroups with an Asian origin in the Sami population.16 Although previous data from protein polymorphisms, blood groups, mtDNA and the Y-chromosome have indicated a contribution from Asian populations, the extent of this contribution to the Sami gene pool has not been quantified. A previous study of HLA class II variation indicated a closer relationship between a Sami population from north-western Russia and Oriental (Japanese and Korean) populations compared to non-Sami European populations.35 As that study was based on HLA typing with a lower genetic resolution than employed in our study, the two datasets are difficult to compare.
The high frequency of some class I and class II alleles in our Sami samples that are characteristic of European populations, supports the theory that the Sami gene pool is predominantly of European origin. At the same time, the high frequency of some alleles and haplotypes in the Sami population that are rare in other European populations but common in Northeast Asian (particularly Siberian) and Native American populations, provides strong evidence of a genetic contribution to the Sami from Asian populations. Our MDS and phylogenetic network analyses both place the Swedish Sami in close vicinity to other European populations, whereas some analyses indicate a similarity to some Siberian populations. The relationship among populations differs somewhat between the class I and class II loci, emphasizing the need to use multiple loci for studying the genetic relationships between populations. In the phylogenetic network for the class II loci, northern Sami were found in the group with Northeast Asian populations, in between South and North Siberian populations. Interestingly this phylogeny is similar to a previous study of SNP variation in the non-recombining part of the Y-chromosome.36
The admixture analysis also indicates that the main contribution to the Swedish Sami is from European populations, but shows a surprisingly high Asian influence (13%). The admixture estimates might depend on the choice of parental populations and we presently lack complete HLA data on most Asian populations. However, since using different contemporary Asian populations in the analyses does not change the admixture proportion substantially (data not shown), this does not appear to be a very serious limitation. In the STRUCTURE analyses based on the genotypes of individuals, about 30% of the northern Sami individuals have a genotype composition which resembles that of Northeast Asian populations. In our STRUCTURE analysis we chose to infer four population clusters. Estimating the most supported number of clusters (K) for a dataset is not trivial. Since, for our dataset, the likelihood continues to increase with increasing K (eg, due to isolation by distance or inbreeding37), we focused on a value of K=4, which captures most of the structure in the data and seems biologically relevant.
Although the estimates for time of admixture and effective population size are tentative and depend on a number of assumptions, they are interesting in relation to hypotheses for the origin of the Sami. The time estimated for admixture in the southern Sami (5300 YBP) is earlier than the presumed colonization of northern Sweden by other European populations. This might suggest that people from Central Europe reached the southern part of the Sami range earlier than previously assumed, or may reflect a migration of Sami to the southern part of their range, which at that time was already populated by other European tribes. The wide confidence interval for the estimates includes both the possibility of recent admixture or an early migration of non-Sami populations to northern Scandinavia.
The estimated effective population sizes indicate that the Sami population has existed at a size roughly 20% of the effective size of the non-Sami Swedish population. The Sami are not known to have experienced a dramatic bottleneck in historic times, but are believed to have maintained a relatively constant, but limited, population size over time. The LD between microsatellite markers has been shown to be 2–3 times higher in the southern Sami than in the non-Sami Swedish population, consistent with a more limited population size.38 Interestingly, Kauppi and collegues39 found the LD in 75 kb of MHC class II region to be similar between Sami and UK Europeans, but that many haplotypes were population specific. Our data shows that some of the HLA alleles (eg, DRB1*0801) have a much higher frequency in the northern Sami than in any other human population used for comparison. This could reflect demographic events, such as a bottleneck or genetic drift, or be the result of specific selection for this or other alleles (eg, B*2705) on the same haplotype. Indeed, our Ewens–Watterson analysis indicates that selection has been acting on two of the class II loci, consistent with the finding that balancing selection has been acting on some HLA polymorphisms.19 In addition, the smaller effective population size in both the northern and southern Sami populations makes them more sensitive to genetic drift. However, due to lack of information on nature of the putative selection, we cannot distinguish between selection and drift (or a combination of the two) as explanations for the unusually high frequency of these alleles.
Both the higher number of HLA alleles in the southern Sami and the larger allelic overlap with alleles present in the non-Sami Swedish population, are indicative of recent admixture between the southern Sami and the non-Sami Swedish population. Under this assumption of admixture between a putative ancestral Sami population (represented by the northern Sami) and the non-Sami Swedish population, the proportion of non-Sami Swedish influence in the southern Sami was estimated to be 58%. An independent estimate based on mtDNA also indicated considerable (48%) admixture of non-Sami European ancestry in the southern Sami.16
In summary, the HLA class I and class II analyses show that the main genetic contribution to the Swedish Sami has come from European populations. However, the estimated Asian influence in the northern Sami is higher than that indicated by other genetic markers. The genetic contribution from Asian populations to the southern Sami is lower compared to the northern Sami, most likely due to significant admixture with the non-Sami Swedish population.
References
Nordqvist B : Coastal adaptations in the Mesolitic. A study of costal sites with organic remains from the Boreal and Atlantic periods in Western Sweden. GOTARC, Series B 13 Göteborg 2000.
Hassler S, Sjölander P, Ericsson AJ : Construction of a database on health and living conditions of the Swedish Sami population. In: Befolkning och bosättning i norr – Etnicitet, identitet och gränser i historiens sken. Umeå: Centre for Sami Research, Umeå University, 2004; 107–124.
Beckman L, Broman B, Jonsson B, Mellbin T : Further data on teh blood groups of teh Swedish Lapps. Acta Genet 1959; 9: 1–8.
Allison AC, Broman B, Mourant AE, Ryttinger L : The blood groups of Swedish Lapps. J R Anthropol Inst 1956; 86: 87–94.
Allison AC, Hartmann O, Brendemoen OJ, Mourant AE : The blood groups of the Norwegian Lapps. Acta Pathol Microbiol Scand 1952; 31: 334–338.
Beckman L : Samerna en genetiskt unik urbefolkning. Umeå: Solfjädern Offset AB, 1996.
Beckman L, Beckman G, Nylander PO : Gc subtypes in Finns, Swedes and Swedish Lapps. Hum Hered 1988; 38: 18–21.
Beckman G, Beckman L, Sikstrom C : Serum complement (C3, BF, C4) types in Swedish Saamis. Hum Hered 1993; 43: 362–365.
Fan C, Sikstrom C, Beckman G, Beckman L : Orosomucoid polymorphism in Finns, Swedes and Swedish Saamis. Hum Hered 1993; 43: 272–275.
Sikstrom C, Nylander PO : Transferrin C subtypes and ethnic heterogeneity in Sweden. Hum Hered 1990; 40: 335–339.
Larsen LA, Vuust J, Nystad M, Evseeva I, Van Ghelue M, Tranebjaerg L : Analysis of FMR1 (CGG)(n) alleles and DXS548-FRAXAC1 haplotypes in three European circumpolar populations: traces of genetic relationship with Asia. Eur J Hum Genet 2001; 9: 724–727.
Cavalli-Sforza LL, Piazza A : Human genomic diversity in Europe: a summary of recent research and prospects for the future. Eur J Hum Genet 1993; 1: 3–18.
Sajantila A, Lahermo P, Anttinen T et al.: Genes and languages in Europe: an analysis of mitochondrial lineages. Genome Res 1995; 5: 42–52.
Tambets K, Rootsi S, Kivisild T et al.: The western and eastern roots of the Saami – the story of genetic ‘outliers’ told by mitochondrial DNA and Y chromosomes. Am J Hum Genet 2004; 74: 661–682.
Torroni A, Bandelt HJ, Macaulay V et al.: A signal, from human mtDNA, of postglacial recolonization in Europe. Am J Hum Genet 2001; 69: 844–852.
Ingman M, Gyllensten U : A recent genetic link between Sami and the Volga-Ural region of Russia. Eur J Hum Genet 2007; 15: 115–120.
Gordon RG : Ethnologue: Languages of the World. Dallas: SIL International, 2005.
Marsh SG : Nomenclature for factors of the HLA system, update September 1999. Marsh for the WHO Nomenclature Committee for Factors of the HLA System. Tissue Antigens 2000; 55: 100.
Hughes AL, Nei M : Pattern of nucleotide substitution at major histocompatibility complex class I loci reveals overdominant selection. Nature 1988; 335: 167–170.
Engelmark M, Beskow A, Magnusson J, Erlich H, Gyllensten U : Affected sib-pair analysis of the contribution of HLA class I and class II loci to development of cervical cancer. Hum Mol Genet 2004; 13: 1951–1958.
Bugawan TL, Apple R, Erlich HA : A method for typing polymorphism at the HLA-A locus using PCR amplification and immobilized oligonucleotide probes. Tissue Antigens 1994; 44: 137–147.
Erlich H, Bugawan T, Begovich AB et al.: HLA-DR, DQ and DP typing using PCR amplification and immobilized probes. Eur J Immunogenet 1991; 18: 33–55.
Schneider S, Roessli D, Excoffier L : Arlequin. A software for population genetics data analysis; Ver 2.000., Genetics and Biometry Lab, Department of Anthropology, University of Geneva., 2000.
Excoffier L, Smouse PE, Quattro JM : Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 1992; 131: 479–491.
Guo SW, Thompson EA : Performing the exact test of Hardy–Weinberg proportion for multiple alleles. Biometrics 1992; 48: 361–372.
Nei M : Genetic distance between populations. Am Nat 1972; 106: 283–292.
Saitou N, Nei M : The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 1987; 4: 406–425.
Felsenstein J : PHYLIP – phylogeny inference package (Version 3.2). Cladistics 1989; 5: 164–166.
Page RD : TreeView: an application to display phylogenetic trees on personal computers. Comput Appl Biosci 1996; 12: 357–358.
Pritchard JK, Stephens M, Donnelly P : Inference of population structure using multilocus genotype data. Genetics 2000; 155: 945–959.
Wang J : Maximum-likelihood estimation of admixture proportions from genetic data. Genetics 2003; 164: 747–765.
Slatkin M : A correction to the exact test based on the Ewens sampling distribution. Genet Res 1996; 68: 259–260.
Slatkin M : An exact test for neutrality based on the Ewens sampling distribution. Genet Res 1994; 64: 71–74.
Macaulay V, Hill C, Achilli A et al.: Single, rapid coastal settlement of Asia revealed by analysis of complete mitochondrial genomes. Science 2005; 308: 1034–1036.
Evseeva I, Spurkland A, Thorsby E et al.: HLA profile of three ethnic groups living in the North-Western region of Russia. Tissue Antigens 2002; 59: 38–43.
Wells RS, Yuldasheva N, Ruzibakiev R et al.: The Eurasian heartland: a continental perspective on Y-chromosome diversity. Proc Natl Acad Sci USA 2001; 98: 10244–10249.
Pritchard JK, Wen X, Falush D : Documentation for STRUCTURE software: version 2.2. Department of Human Genetics, University of Chicago, 2007.
Johansson A, Vavruch-Nilsson V, Edin-Liljegren A, Sjolander P, Gyllensten U : Linkage disequilibrium between microsatellite markers in the Swedish Sami relative to a worldwide selection of populations. Hum Genet 2005; 116: 105–113.
Kauppi L, Sajantila A, Jeffreys AJ : Recombination hotspots rather than population history dominate linkage disequilibrium in the MHC class II region. Hum Mol Genet 2003; 12: 33–40.
Acknowledgements
We are grateful for the samples from the northern Sami communities originally collected by the late Professor Lars Beckman of Umeå University. The contribution of samples from the southern Sami communities from the Medical Biobank in Umeå is also greatly appreciated. The study was supported by grants from the Swedish Natural Sciences Research Council, the European Commission/EUROSPAN and the Foundation for Strategic Research (SSF).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on European Journal of Human Genetics website (http://www.nature.com/ejhg)
Supplementary information
Rights and permissions
About this article
Cite this article
Johansson, Å., Ingman, M., Mack, S. et al. Genetic origin of the Swedish Sami inferred from HLA class I and class II allele frequencies. Eur J Hum Genet 16, 1341–1349 (2008). https://doi.org/10.1038/ejhg.2008.88
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2008.88
Keywords
This article is cited by
-
SweHLA: the high confidence HLA typing bio-resource drawn from 1000 Swedish genomes
European Journal of Human Genetics (2020)
-
A genome-wide analysis of population structure in the Finnish Saami with implications for genetic association studies
European Journal of Human Genetics (2011)