Abstract
Microsatellite repeats are frequently found to be mutated in microsatellite-instable colorectal tumours. This suggests that these mutations are important events during tumour development. We have observed frequent mutations in microsatellite-instable (MSI-H) tumours and cell lines of a conserved A14 repeat within the 3′-untranslated region of the interferon-γ receptor 1 gene (IFNGR1). The repeat was mutated in 59% (41 of 70) of colon carcinomas and in all four MSI-H colon cancer cell lines tested. In-vitro analysis of these cell lines did not show a decreased responsiveness to standard IFNγ concentrations when compared to microsatellite-stable colon cancer cell lines. A functional consequence of the frequently found microsatellite instability in IFNGR1 is therefore not evident.
Similar content being viewed by others
Introduction
DNA mismatch repair deficiency in tumours is characterized by a high frequency of microsatellite instability (MSI-H).1 Microsatellite instability is associated with the hereditary non-polyposis colorectal cancer syndrome and also appears in approximately 15% of sporadic colorectal tumours.2 A subset of genes that encompass coding microsatellites may be specifically targeted in MSI-H tumours. Systemic sequence database searches have identified in the human genome up to 17 000 intra-exon coding repeat sequences with a length of six or more nucleotides.3, 4
Frameshift mutations in MSI-H colorectal tumours have been demonstrated for genes involved in signal transduction (TGFBRII, IGFIIR, PTEN), apoptosis (BAX, CASPASE 5), DNA repair (hMSH3, hMSH6, MBD4), transcriptional regulation (TCF-4) and immune surveillance (B2M), with mutation frequencies ranging from 4 to 80%.5
Non-coding microsatellite repeats, however, may just as well be mutated. Of those with potential functional significance are those that lie within the untranslated regions (UTR) of genes. Elements in such regions can regulate mRNA degradation, translation and localization.6, 7, 8 However, such elements have not been extensively studied for and their identification is complicated by the fact that their activity often depends on specific secondary RNA structures.9 Suraweera et al10 characterized frequent mutations in a number of conserved mononucleotide repeats within UTRs and high levels of mutations were found in a novel T25 mononucleotide marker in the 3′-UTR of the CASP2 gene.11
Interferon-γ (IFNγ) is important in regulating cell-mediated immune responses. It directly increases the sensitivity of a target cell to CD8+ cytotoxic lymphocyte attack by upregulating the expression of a variety of genes, including the human leucocyte antigen (HLA) class I genes and Fas/CD95.12 Mismatch repair-deficient tumours potentially exhibit a large number of tumour-specific ‘frameshift’ antigens due to instability of coding microsatellite sequences. This has been suggested to render them more susceptible to both native and therapeutically induced anti-tumour immune responses.13 Indeed, these tumours are associated with an elevated CD8+ intra-epithelial infiltrate.14, 15 Tumour IFNγ responsiveness is important in regulating anti-tumour immune response in vivo, partly depending on antigen presentation.16, 17 Downregulation of the IFNγ receptor has been shown in hepatocellular carcinoma,18 basal cell carcinoma19 and virus-associated tumours,20, 21 and was correlated with larger tumour size and a higher frequency of metastases.18 IFNγ also increases the sensitivity to chemotherapy-induced Fas-mediated apoptosis.22 Although the use of IFNγ solely has yielded somewhat disappointing results in clinical anti-cancer treatment trials,23 its use in combination with different chemotherapeutics, such as 5-fluorouracil, indomethacin, phenyl butyrate, mitomycin C, cyclosporine A or clarithromycin has shown synergetic anti-tumour effects.24, 25, 26, 27, 28, 29, 30 Thus, loss of IFNγ responsiveness would denote a loss of potential for future immuno- and chemo-based adjuvant therapies.
In this study, we describe frequent mutation of the A14 repeat within the 3′-UTR of IFNGR1. As will be discussed, it is difficult to draw conclusions regarding its importance on basis of frequencies alone. Therefore, we also analysed IFNγ responsiveness of colon cancer cells bearing these mutations.
Materials and methods
Colon tumour samples and cell culture
Tumour samples were derived from formalin-fixed, parafin-embedded material sent to our department for MSI analysis between 1999 and 2002. Cases were analysed following the medical ethnical guidelines described in the Code Proper Secondary Use of Human Tissue established by the Dutch Federation of Medical Sciences. The present study falls under approval by the Medical Ethical Committee of the LUMC (protocol P01–019).
Cell lines HCT 15, LoVo, LS180, LS411N, SW480 and SW837 were obtained from the American Type Culture Collection (Atlanta, GA, USA) and cultured and harvested according to standard methods.
Microsatellite instability and sequence analysis
MSI and sequence analyses were performed as described previously.31. MSI analysis was performed using the NCI markers BAT25, BAT26, D2S123, D5S346, D17S250,32 supplemented with BAT40, MSH3 and MSH6. Tumours were classified as MSI-H (instability of at least 30% of the markers) or MSS (no instability). IFNGR1 was analysed using forward primer 5′-GAGGATGTGTGGCATTTTCA-3′ and reverse primer 5′-TGCTATACCAAGGCAGAGAAAAG-3′.
IFNGR1 mRNA expression
Total RNA was isolated using TriZol (Ambion, Austin, TX, USA) following the manufacturer's recommendations. RNA (2 μg) was used for cDNA synthesis using Oligo dT primers and AMV Reverse Transcriptase (Roche Applied Science, Penzberg, Germany). We performed quantitative real-time PCR using SYBR Green (Eurogentec, Seraing, Belgium) on an iCycler real-time PCR detection system (Bio-Rad, Hercules, CA, USA). Ct values were calibrated to an amount of Human Total Colon RNA (Clontech, BD Biosciences, Palo Alto, CA, USA) and normalized to the level of expression of three stably expressed housekeeping genes HNRPM, CPSF6 and TBP.33 Calibrated Ct values were divided by Ct values of 2 μg Human Total Colon RNA. Primers for amplification of IFNGR1 mRNA used were 5′-TCCTCAGTGCCTACACCAACTAA-3′ (nucleotides 79–101, exon 1–2), and 5′-CTCGTCACAATCATCTTCCTTCTG-3′ (nucleotides 568–591, exon 5).
Flow cytometry of IFNγ-stimulated cells
HLA I and Fas expression was analysed by flow cytometry of viable cells as described previously34, 35 using the primary antibodies W6/32 (supernatant), 2R2 (Alexis Biochemicals, Carlsbad, CA, USA), and mouse isotype controls (Dako Cytomation, Glostrup, Denmark). Samples containing 1 μ M propidium iodide (Calbiochem, San Diego, CA, USA) were collected on an FACSCalibur (Becton Dickinson Immunocytometry Systems, San Jose, CA, USA). Linearized mean fluorescence intensity values were determined by WinList (Verity Software House Inc., Topsham, ME, USA) and subtracted by control values. Means derived from duplicate experiments were used; single experiments encompassed one-session analyses of all cell lines using the same instrument settings. We constructed a dose–response curve using 8, 40, 200 and 1000 U/ml IFNγ (PeproTech Inc., Rocky Hill, NJ, USA) for 48 h. All cell lines were already maximally responsive to a concentration of 8 U/ml. The extent of upregulation was further analysed using 200 U/ml IFNγ.
Fas-induced apoptosis
To quantify Fas-induced apoptosis using 1 μg/ml 2R2 (Alexis Biochemicals) and 1 μg/ml protein A for 6 h, percentages of apoptotic cells were determined using the Nicoletti assay as described previously.22, 36.For analysis we used a modified ModFit algorithm previously described37 and kindly provided by Verity Software House Inc. Means derived from duplicate experiments were used.
Results and discussion
Frequently found MSI suggests a clonally selective advantage, although passenger mutations due to a possible lack of selection pressure of innocent bystander genes, ie genes whose function is irrelevant during tumourigenesis, cannot be excluded.38 For TCF-4, it has been shown that frameshift mutations due to instability of a coding A9 repeat lack functional consequences.39 Therefore, comprehensive criteria have been proposed at the Bethesda meeting to define valid target genes in MSI-H tumours, including (1) a high frequency of mutation, (2) bi-allelic inactivation, (3) involvement in a tumour suppressor pathway, which is (4) also involved in MSS tumours and (5) functional consequences of the mutation.2 However, few reports have taken these criteria into account40, 41 and statistical methods have been proposed to identify valid target genes by statistical analysis of mutation frequencies only.42, 43
To examine the (in)stability of the IFNGR1 repeat we first analysed 17 MSS colon cancer samples and 2 MSS colon cancer cell lines (SW480 and SW837). In all cases the repeat was conserved, ie monomorphic.10 Subsequently, 70 MSI-H colon tumour specimens were tested from patients complying with the clinical Bethesda criteria,1 38 of whom a germ line mutation was identified in one of the MMR genes. MSI in the repeat was found in tumours from 12 of 20 (60%) hMLH1 patients, 6 of 12 (50%) hMSH2 patients, 2 of 6 (33%) hMSH6 patients, in 21 of 32 (66%) of the residual MSI-H tumours and in 4 of 4 (100%) MSI-H colon cancer cell lines.
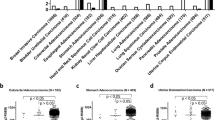
The frequency of mutations in microsatellite tracts is associated with the number and type of repeats.32, 44 Previous studies reported a mutation frequency up to 54% in 29 mononucleotide repeats (8–10 bp) within intronic sequences;38 in conserved repeats instability was found in up to 28% of tumours only.42 Furthermore, G-repeats are considerably more prone to mutations than A-repeats.38 In 19 conserved UTR repeats studied10 a mutation frequency was found up to 95%, but the repeat lengths ranged from 15 up to 32 nucleotides. Led by the Bethesda criteria2 we decided to study whether 59% MSI in the A14 repeat is truly indicative of a valid target.
First, we studied bi-allelicity of the mutations. The microsatellite analysis used cannot discriminate between mono- or bi-allelic mutations of tumour tissue due to contaminating normal cells. Therefore, we analysed the cell lines described above and performed sequence analysis for confirmation of the observed deletions. A bi-allelic A3 deletion was detected in LoVo and LS411N, in LS180 both an A4 and an A5 deletions were seen and in HCT 15 only a mono-allelic deletion of a single A was observed. No mutations in the MSS cell lines SW480 and SW837 were observed in line with the MSI results.
To detect a possible decline in IFNGR1 mRNA expression upon 3′-UTR mutations we applied quantitative RT-PCR. The receptor is normally expressed on colon epithelium45 and no decreased IFNGR1 mRNA expression of the MSI-H colon cancer cell lines was observed when compared to the MSS cell lines (Table 1).
To rule out other regulatory effects of the 3′-UTR mutation we investigated the extent of IFNγ-mediated HLA class I and Fas/CD95 upregulation in MSI-H compared to MSS colon cancer cell lines. HLA class I molecules are not present on the cell surface of LoVo and HCT 15 cells, due to mutations of the light chain β2-microglobulin.46 We analysed dose–response curves to make sure the experiments were performed under conditions leading to maximum responsiveness. No decreased responsiveness to IFNγ was observed for MSI-H cell lines compared to the MSS cell lines (Table 1). To study the effect of IFNγ on Fas-induced apoptosis, we measured the fraction of apoptotic cells as quantified by flow cytometry. Again, no impaired IFNγ response was noted in the MSI-H cell lines (Table 1). Although, the concentrations used are similar to other studies on in-vitro IFNγ responses in colon cancer cell lines,47 we cannot completely rule out a differential response to far lower concentration of IFNγ not yielding maximum response.
In conclusion, we identified a conserved A14 microsatellite repeat within the 3′-UTR of the potential tumour suppressor IFNGR1 that is mutated in 59% of MSI-H colorectal tumours. Compared to MSI of other conserved, non-coding but transcribed short mononucleotide tracts, the high frequency of instability suggests it to be a valid target during colorectal tumourigenesis. However, in-vitro analysis did not show an impaired response to IFNγ in any cell line harbouring the mutated gene questioning the functional consequences of MSI in this repeat. High mutation rates as a consequence of unidentified sequence-dependent factors, chromatin structure or mechanisms associated with transcription may contribute to the high frequency of mutations found in tumours in the absence of selective advantage.38, 40, 48 Studying functional consequences seems indispensable to describe the role of these mutations in tumourigenesis rather than the frequency of genomic instability per se.49, 50 Despite a high frequency of MSI of the 3′-UTR of IFNGR1, it probably does not contribute to colorectal tumour progression.
References
Rodriguez-Bigas MA, Boland CR, Hamilton SR et al: A National Cancer Institute Workshop on Hereditary Nonpolyposis Colorectal Cancer Syndrome: meeting highlights and Bethesda guidelines. J Natl Cancer Inst 1997; 89: 1758–1762.
Boland CR, Thibodeau SN, Hamilton SR et al: A National Cancer Institute workshop on microsatellite instability for cancer detection and familial predisposition: development of international criteria for the determination of microsatellite instability in colorectal cancer. Cancer Res 1998; 58: 5248–5257.
Woerner SM, Gebert J, Yuan YP et al: Systematic identification of genes with coding microsatellites mutated in DNA mismatch repair-deficient cancer cells. Int J Cancer 2001; 93: 12–19.
Park J, Betel D, Gryfe R et al: Mutation profiling of mismatch repair-deficient colorectal cancers using an in silico genome scan to identify coding microsatellites. Cancer Res 2002; 62: 1284–1288.
De Leeuw W : General Introduction; in de Leeuw W: Genetic instability in HNPCC related tumors, PhD thesis, Leiden 2003, pp 9–65.
Wickens M, Bernstein DS, Kimble J, Parker R : A PUF family portrait: 3′UTR regulation as a way of life. Trends Genet 2002; 18: 150–157.
Veyrune JL, Campbell GP, Wiseman J, Blanchard JM, Hesketh JE : A localisation signal in the 3′ untranslated region of c-myc mRNA targets c-myc mRNA and beta-globin reporter sequences to the perinuclear cytoplasm and cytoskeletal-bound polysomes. J Cell Sci 1996; 109 (Part 6): 1185–1194.
Laroia G, Sarkar B, Schneider RJ : Ubiquitin-dependent mechanism regulates rapid turnover of AU-rich cytokine mRNAs. Proc Natl Acad Sci USA 2002; 99: 1842–1846.
Pesole G, Liuni S, D'Souza M : PatSearch: a pattern matcher software that finds functional elements in nucleotide and protein sequences and assesses their statistical significance. Bioinformatics 2000; 16: 439–450.
Suraweera N, Iacopetta B, Duval A, Compoint A, Tubacher E, Hamelin R : Conservation of mononucleotide repeats within 3′ and 5′ untranslated regions and their instability in MSI-H colorectal cancer. Oncogene 2001; 20: 7472–7477.
Findeisen P, Kloor M, Merx S et al: T25 repeat in the 3′ untranslated region of the CASP2 gene: a sensitive and specific marker for microsatellite instability in colorectal cancer. Cancer Res 2005; 65: 8072–8078.
Bergmann-Leitner ES, Abrams SI : Influence of interferon gamma on modulation of Fas expression by human colon carcinoma cells and their subsequent sensitivity to antigen-specific CD8+ cytotoxic T lymphocyte attack. Cancer Immunol Immunother 2000; 49: 193–207.
Linnebacher M, Gebert J, Rudy W et al: Frameshift peptide-derived T-cell epitopes: a source of novel tumor-specific antigens. Int J Cancer 2001; 93: 6–11.
Jass JR : Pathology of hereditary nonpolyposis colorectal cancer. Ann NY Acad Sci 2000; 910: 62–73.
Guidoboni M, Gafa R, Viel A et al: Microsatellite instability and high content of activated cytotoxic lymphocytes identify colon cancer patients with a favorable prognosis. Am J Pathol 2001; 159: 297–304.
Kaplan DH, Shankaran V, Dighe AS et al: Demonstration of an interferon gamma-dependent tumor surveillance system in immunocompetent mice. Proc Natl Acad Sci USA 1998; 95: 7556–7561.
Shankaran V, Ikeda H, Bruce AT et al: IFNgamma and lymphocytes prevent primary tumour development and shape tumour immunogenicity. Nature 2001; 410: 1107–1111.
Nagao M, Nakajima Y, Kanehiro H et al: The impact of interferon gamma receptor expression on the mechanism of escape from host immune surveillance in hepatocellular carcinoma. Hepatology 2000; 32: 491–500.
Kooy AJ, Tank B, Vuzevski VD, van Joost T, Prens EP : Expression of interferon-gamma receptors and interferon-gamma-induced up-regulation of intercellular adhesion molecule-1 in basal cell carcinoma; decreased expression of IFN-gamma R and shedding of ICAM-1 as a means to escape immune surveillance. J Pathol 1998; 184: 169–176.
Morrison TE, Mauser A, Wong A, Ting JP, Kenney SC : Inhibition of IFN-gamma signaling by an Epstein–Barr virus immediate-early protein. Immunity 2001; 15: 787–799.
Moore PS, Chang Y : Molecular virology of Kaposi's sarcoma-associated herpesvirus. Philos Trans R Soc Lond B Biol Sci 2001; 356: 499–516.
von Reyher U, Strater J, Kittstein W, Gschwendt M, Krammer PH, Moller P : Colon carcinoma cells use different mechanisms to escape CD95-mediated apoptosis. Cancer Res 1998; 58: 526–534.
Meyskens Jr FL, Kopecky K, Samson M et al: Recombinant human interferon gamma: adverse effects in high-risk stage I and II cutaneous malignant melanoma. J Natl Cancer Inst 1990; 82: 1071.
Huang Y, Horvath CM, Waxman S : Regrowth of 5-fluorouracil-treated human colon cancer cells is prevented by the combination of interferon gamma, indomethacin, and phenylbutyrate. Cancer Res 2000; 60: 3200–3206.
Aquino A, Prete SP, Greiner JW et al: Effect of the combined treatment with 5-fluorouracil, gamma-interferon or folinic acid on carcinoembryonic antigen expression in colon cancer cells. Clin Cancer Res 1998; 4: 2473–2481.
Li G, Kawakami S, Kageyama Y, Yan C, Saito K, Kihara K : IFN gamma-induced up-regulation of PD-ECGF/TP enhances the cytotoxicity of 5-fluorouracil and 5′-deoxy-5-fluorouridine in bladder cancer cells. Anticancer Res 2002; 22: 2607–2612.
Ishihara M, Kubota T, Watanabe M et al: Interferon gamma increases the antitumor activity of mitomycin C against human colon cancer cells in vitro and in vivo. Oncol Rep 1999; 6: 621–625.
Schwartzberg LS, Petak I, Stewart C et al: Modulation of the Fas signaling pathway by IFN-gamma in therapy of colon cancer: phase I trial and correlative studies of IFN-gamma, 5-fluorouracil, and leucovorin. Clin Cancer Res 2002; 8: 2488–2498.
Berek JS : Interferon plus chemotherapy for primary treatment of ovarian cancer. Lancet 2000; 356: 6–7.
Windbichler GH, Hausmaninger H, Stummvoll W et al: Interferon-gamma in the first-line therapy of ovarian cancer: a randomized phase III trial. Br J Cancer 2000; 82: 1138–1144.
de Leeuw WJ, van Puijenbroek M, Merx R et al: Bias in detection of instability of the (C)8 mononucleotide repeat of MSH6 in tumours from HNPCC patients. Oncogene 2001; 20: 6241–6244.
Eichler EE, Holden JJ, Popovich BW et al: Length of uninterrupted CGG repeats determines instability in the FMR1 gene. Nat Genet 1994; 8: 88–94.
van Wezel T, Lombaerts M, van Roon EH et al: Expression analysis of candidate breast tumour suppressor genes on chromosome 16q. Breast Cancer Res 2005; 7: R998–R1004.
Sasaki DT, Dumas SE, Engleman EG : Discrimination of viable and non-viable cells using propidium iodide in two color immunofluorescence. Cytometry 1987; 8: 413–420.
Corver WE, Cornelisse CJ, Fleuren GJ : Simultaneous measurement of two cellular antigens and DNA using fluorescein-isothiocyanate, R-phycoerythrin, and propidium iodide on a standard FACScan. Cytometry 1994; 15: 117–128.
Nicoletti I, Migliorati G, Pagliacci MC, Grignani F, Riccardi C : A rapid and simple method for measuring thymocyte apoptosis by propidium iodide staining and flow cytometry. J Immunol Methods 1991; 139: 271–279.
Rorke EA, Jacobberger JW : Transforming growth factor-beta 1 (TGF beta 1) enhances apoptosis in human papillomavirus type 16-immortalized human ectocervical epithelial cells. Exp Cell Res 1995; 216: 65–72.
Zhang L, Yu J, Willson JK, Markowitz SD, Kinzler KW, Vogelstein B : Short mononucleotide repeat sequence variability in mismatch repair-deficient cancers. Cancer Res 2001; 61: 3801–3805.
Ruckert S, Hiendlmeyer E, Brueckl WM et al: T-cell factor-4 frameshift mutations occur frequently in human microsatellite instability-high colorectal carcinomas but do not contribute to carcinogenesis. Cancer Res 2002; 62: 3009–3013.
Duval A, Hamelin R : Mutations at coding repeat sequences in mismatch repair-deficient human cancers: toward a new concept of target genes for instability. Cancer Res 2002; 62: 2447–2454.
Perucho M : Tumors with microsatellite instability: many mutations, targets and paradoxes. Oncogene 2003; 22: 2223–2225.
Duval A, Reperant M, Hamelin R : Comparative analysis of mutation frequency of coding and non coding short mononucleotide repeats in mismatch repair deficient colorectal cancers. Oncogene 2002; 21: 8062–8066.
Woerner SM, Benner A, Sutter C et al: Pathogenesis of DNA repair-deficient cancers: a statistical meta-analysis of putative Real Common Target genes. Oncogene 2003; 22: 2226–2235.
Chen J, Heerdt BG, Augenlicht LH : Presence and instability of repetitive elements in sequences the altered expression of which characterizes risk for colonic cancer. Cancer Res 1995; 55: 174–180.
Valente G, Ozmen L, Novelli F et al: Distribution of interferon-gamma receptor in human tissues. Eur J Immunol 1992; 22: 2403–2412.
Bicknell DC, Rowan A, Bodmer WF : Beta 2-microglobulin gene mutations: a study of established colorectal cell lines and fresh tumors. Proc Natl Acad Sci USA 1994; 91: 4751–4756.
Woerner SM, Kloor M, Schwitalle Y et al: The putative tumor suppressor AIM2 is frequently affected by different genetic alterations in microsatellite unstable colon cancers. Genes Chromosomes Cancer 2007; 46: 1080–1089.
Bellacosa A : Functional interactions and signaling properties of mammalian DNA mismatch repair proteins. Cell Death Differ 2001; 8: 1076–1092.
Sieber OM, Heinimann K, Tomlinson IP : Genomic instability – the engine of tumorigenesis? Nat Rev Cancer 2003; 3: 701–708.
Rajagopalan H, Nowak MA, Vogelstein B, Lengauer C : The significance of unstable chromosomes in colorectal cancer. Nat Rev Cancer 2003; 3: 695–701.
Acknowledgements
We thank Dr JP Medema for assistance with the Nicoletti assays and Dr WE Corver for assistance with ModFit analyses. This work was supported by grant 2000/2135 of the Dutch Cancer Society (KWF) and by the Centre for Medical Systems Biology (CMSB), a centre of excellence approved by the Netherlands Genomic Initiative/Netherlands Organisation for Scientific Research (NWO). No potential conflicts of interest are declared.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Dierssen, J., van Puijenbroek, M., Dezentjé, D. et al. Frequent mutations in the 3′-untranslated region of IFNGR1 lack functional impairment in microsatellite-unstable colorectal tumours. Eur J Hum Genet 16, 1235–1239 (2008). https://doi.org/10.1038/ejhg.2008.81
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2008.81