Abstract
The human population is undergoing a major transition from a historical metapopulation structure of relatively isolated small communities to an outbred structure. This process is predicted to increase average individual genome-wide heterozygosity (h) and could have effects on health. We attempted to quantify this increase in mean h. We initially sampled 1001 examinees from a metapopulation of nine isolated villages on five Dalmatian islands (Croatia). Village populations had high levels of genetic differentiation, endogamy and consanguinity. We then selected 166 individuals with highly specific personal genetic histories to form six subsamples, which could be ranked a priori by their predicted level of outbreeding. The measure h was then estimated in the 166 examinees by genotyping 1184 STR/indel markers and using two different computation methods. Compared to the value of mean h in the least outbred sample, values of h in the remaining samples increased successively with predicted outbreeding by 0.023, 0.038, 0.058, 0.067 and 0.079 (P<0.0001), where these values are measured on the same scale as the inbreeding coefficient (but opposite sign). We have shown that urbanisation was associated with an average increase in h of up to 0.08–0.10 in this Croatian metapopulation, regardless of the method used. Similar levels of differentiation have been described in many populations. Therefore, changes in the level of heterozygosity across the genome of this magnitude may be common during isolate break-up in humans and could have significant health effects through the established genetic mechanism of hybrid vigour/heterosis.
Similar content being viewed by others
Introduction
The demographic structure of human populations has undergone a dramatic change in the last two centuries. Over this period the total human population has expanded rapidly from 1.5 to more than 6.2 billion and the percentage of the humans inhabiting cities has increased from 2% to over 50%.1This has been associated with migration from smaller and more genetically uniform settlements to much larger settlements. The greatly increased population mobility has led to marriage outside traditional groups and admixture of many genetically differentiated populations in a process referred to as ‘isolate break-up’.2 These processes have led to a decrease in the extent of population substructure and linkage disequilibrium, especially between 1900 and the present day in European populations.3, 4 An immediate predicted effect of urbanisation and admixture is an increase in the level of average individual genome-wide heterozygosity (h) among people born in urban areas. This is because outbreeding of individuals from different populations is expected to increase h in the offspring, just as crossing of plant and animal strains leads to hybrids with increased h.5
In theory, increasing h in human populations due to urbanisation could influence variation in biological traits that show significant dominance variance.6, 7 These include many traits such as systolic blood pressure, total cholesterol and LDL-cholesterol that are major contributors to the global burden of disease.8, 9, 10 Furthermore, increased heterozygosity at the individual level could in theory improve resistance to detrimental factors in the environment, especially infectious diseases, and could delay the onset of many chronic diseases.11, 12 Finally, outbreeding should decrease the burden of recessive monogenic conditions that are characteristic of large global communities with a high prevalence of consanguineous marriages.13, 14 Because of these potential effects of h on health we considered it important to quantify the magnitude of increase in h that is associated with urbanisation relative to traditional rural populations.
Several of our recent findings form the scientific basis for this study. First, we have characterised, in detail, a metapopulation of nine Croatian villages from five Dalmatian islands that may be reasonably representative of historic rural European populations in population genetic terms.15 Second, we used a small subset of individuals from this sample (with varying degrees of consanguinity determined from genealogical records) to define statistical methods that could be used to determine each individual's h-values using both STR and SNP-based genome-wide scans with reasonable precision.16 Third, we applied this approach to investigate the effects of h on a range of biomedically relevant human quantitative traits with two main conclusions: (i) our estimated h-values showed a very high correlation with genealogical expectations; and (ii) we showed statistically significant associations between h and blood pressure and cholesterol levels.17
To further investigate our general hypothesis that global outbreeding could have substantial effects on burden of disease,11 we need to quantify the typical increase in average h that occurs during isolate break-up and urbanisation, which we attempted to do in this study.
Materials and methods
Population sample
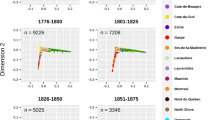
Initially, we examined a targeted sample of 1001 individuals that should have been most informative to understand demographic and genetic histories of the nine isolate populations that were studied. We performed preliminary population genetic analyses of these populations (eg, differentiation between villages based on Wright's fixation index and linkage disequilibrium) using 26 STR markers genotyped in all those individuals (Figure 1).15 The results of a model-based clustering algorithm, implemented in STRUCTURE (http://pritch.bsd.uchicago.edu/software.html), defined six clusters with very distinct genetic signatures, four of which corresponded to single villages – Susak (red), Lopar (blue) Mljet (yellow) and Barbat (green) (Figure 1).
(a) Geographic location of the nine studied villages on islands of the Eastern Adriatic, Northern and Middle Dalmatia, Croatia. (b) Representation, in two-dimensional space, of genetic distances between villages based on allele frequencies at 26 autosomal short tandem repeat (STR) markers – modified from Vitart et al.15 (c) Population structure in the nine Croatian villages analysed based on 26 STR markers – modified from Vitart et al.15
Then, on a limited subsample of 385 of the 1001 individuals, we performed genome-wide scans with 1184 STR/indel markers to investigate the association of h with a range of human quantitative traits, which we published recently.17 This subset of 385 individuals provided the basis for allele frequency for all 1184 marker loci for computation of the individual levels of heterozygosity.
However, of the 385 individuals with genome-wide scans, only 166 were highly informative for testing the hypothesis of the present paper, ie, global admixture through urbanisation is leading to a substantial increase of heterozygosity in human populations. We selected those 166 individuals with highly specific personal genetic histories based on their clear affiliation to one of the six subsamples, which could be ranked a priori by their predicted level of outbreeding. Eventually, the six subsamples comprised:
-
a)
24 individuals from Susak village, who were completely endogamous in the second ancestral generation (all four grandparents were born in the village),
-
b)
26 endogamous individuals from Lopar village
-
c)
17 endogamous individuals from Mljet village
-
d)
31 endogamous individuals from Komiza village
-
e)
33 individuals having one endogamous parent from any of the villages, and the second who immigrated into that village
-
f)
35 individuals who migrated into any of the four investigated villages (a–d) from mainland Croatian settlements of a size greater than 25 000 (but typically 100 000 or more) and for whom it could be confirmed that the ancestry of each of their parents differed
Our predictions of the ranking of h-values in subsamples a–d were based on their current population size (increasing from a to d), the prevalence of individuals in the village that were isonymous in the second parental generation (ie, have the same surname, specific for the village, as that of one of the parents of the mother and the father predicting consanguinity) – 96, 54, 41 and 35%, respectively; and estimated gene diversity from 26 autosomal STR markers in a random sample of 100 individuals from the four study villages (0.72, 0.74, 0.75 and 0.76, respectively).3, 15
Measurements
A total of 1184 STR markers were genotyped at the Marshfield genotyping facility in Wisconsin, USA. They were all assessed for Hardy–Weinberg equilibrium with 198 showing some evidence of departure from Hardy–Weinberg equilibrium (at P<0.05 level). We genotyped control DNA samples ‘blind’ as a means of estimating typing error and found the averaged genotype error rate to be 0.69%. Further details of the genotyping methods are given elsewhere.17
We determined values of h based on 1184 STR/indel markers by two parallel approaches. Both measures yield valid comparisons of heterozygosity across groups, as they control for differing allele frequencies across different communities.16, 17 The first approach was to measure the standardised multilocus heterozygosity (sMLH), the ratio of the proportion of the heterozygous loci of an individual to the mean heterozygosity of those loci at which the individual was typed.18, 19 Weighted averages of chromosome-based heterozygosity estimates were calculated. The scale of this measure is essentially identical to the scale of the inbreeding coefficient, but reversed in sign.
The second approach, denoted as FA, estimated the excess of homozygosity over Hardy–Weinberg expectation, with greater weight being given to homozygotes at loci with lower expected homozygosity. We have described this method in detail in Carothers et al.16 The variance of FA was estimated from the spread of each individual's 22 separate autosomal values, weighted by the number of markers per autosome. Estimates take negative values when homozygosity is lower than Hardy–Weinberg expectation. FA values correspond exactly to inbreeding coefficients of individuals based on observed allelic frequencies except that, as we explained in detail in our previous work,16, 17 the origin (zero-point) on this scale is arbitrary.
The observed allelic frequencies for both methods were based on all successful genotypes in the sample of 385 genotyped recruits, as this was methodologically more appropriate to obtain sMLH and FA measurements.16
Each sMLH and FA estimate was further adjusted to the number of typed loci to avoid any bias due to the positive correlation between the number of STR markers successfully typed and heterozygosity (relative loss of heterozygotes in individuals with the highest proportion of missing marker data).16, 20 This was performed by regressing the sMLH and FA values on the number of genotypes, weighted by the inverse of their variance, and using the residuals in the further analysis.
Standardised multilocus heterozygosity is defined as the ratio of the proportion of typed loci for which an individual is heterozygous and the mean heterozygosity of those loci at which the individual was typed, ie:

where, sMLHI is standardised multilocus heterozygosity of individual I; NIHL is the number of heterozygous loci in the genome of individual I; NIATL is the number of all typed loci in the genome of individual I; ΣL=1 → NIATL([h(L)]/S) is the sum of mean heterozygosities of all individuals in the population sample typed at locus L over all loci that were typed in individual I. Thereby, h(L) is the heterozygosity at every marker locus L for which any individual can have a value of 0 (homozygote) or 1 (heterozygote), and S is the number of individuals in population sample typed at locus L.
The formula used to compute individual values for genome-wide FA was as described in Appendix 2 of Campbell et al,17 using the unbiased estimate of heterozygosity at the kth locus, Hk, given in Equation (2.11), and with nkl, the number of l-alleles at marker locus k, set equal to the corresponding number observed in the set of 385 individuals referred to above. We did not use the approximate weighting variant of the formula, referred to at the end of this Appendix.
Statistical analyses
The overall difference in sMLH and FA values among the six investigated subsamples was explored by ANOVA and significance levels adjusted using a Bonferroni correction.
Results
We defined six subsamples (Figure 1), hypothesised a priori and without the knowledge of genotype data, which would show an ordered ranking of h-values. Figure 2 shows the distribution of sMLH and FA values by subgroups. We show that the mean sMLH increases (Figure 2a) and FA decreases (Figure 2b) steadily from subgroup a to subgroup f in a stepwise fashion in line with our a priori expectations. Overall, the variation around the mean value of FA in each subgroup is substantial, with 50% of the subgroup typically lying within ±2 and 100% within ±6 percentage points from the mean. The full range of the FA estimate between the highest and the lowest of the 166 individual values observed in all six subgroups is 21 percentage points, which implies that the most ‘outbred’ individual had an additional 21% of his/her genome heterozygous in comparison to the most ‘inbred’ individual. Despite the modest sample sizes in each subgroup the overall difference between the six investigated village samples was statistically highly significant (ANOVA P<0.001, F=29.06 for FA and P<0.001, F=10.38 for sMLH). The differences between all 15 possible pairs of the six subgroups were investigated by ANOVA with Bonferroni correction (data not shown). This showed that the differences between any two neighbouring subsamples were not significant, but all other investigated differences were highly significant (P<0.001 in most cases).
(a) Mean level of standardised multilocus heterozygosity (sMLH) by study villages. (b) Mean level of FA by study villages. We found a positive correlation between the number of markers successfully typed and heterozygosity (negative correlation with h).16, 18 To control for this we excluded marker data from individuals for whom there were data on less than 750 markers, and corrected for any remaining effect statistically. This resulted in negative mean FA values in all six subsamples (as a proportion of the failed genotypes were likely to be heterozygotes).16, 18
Compared to the value of mean FA in the least outbred sample, values of FA in the remaining samples decreased successively with predicted outbreeding by 0.023, 0.038, 0.058, 0.067 and 0.079 (P<0.0001). All the results achieved through application of the sMLH yielded similar results, with successive increase among investigated subsamples of 0.037, 0.043, 0.079, 0.064 and 0.101 (P<0.001). This shows that the two methods are highly comparable, although opposite in sign (direction) of their measurement scale. We have shown that urbanisation was associated with an average increase in h of up to 0.08–0.10 in this Croatian metapopulation, regardless on the method used.
Discussion
The observed ranking in sMLH and FA values from a to f can partly be explained by the proportion of inbred individuals in our subgroups. With decreasing village population size, mate choice becomes limited and the prevalence of consanguinity increases. This is shown in the increasing proportion of individuals likely to be consanguineous based on grandparental isonymy data, increasing from 35% (Komiza), to 96% (Susak).15 However, a more detailed analysis of the proportion of consanguineous individuals within our four subsamples from the chosen villages based on further genealogical analyses showed the following proportion of persons having an expected inbreeding coefficient greater than 1.56% (found in offspring of second cousins) within our final subsamples: 100% in Susak, 50% in Lopar, 35% in Mljet and 32% in Komiza. Therefore, the proportion of consanguineous individuals in the subsamples alone cannot explain the pattern of variation found in sMLH and FA values between examinees shown in Figure 2. It is likely that loss of genetic diversity in the most isolated communities, owing primarily to genetic drift, further contributes to the decrease in h. Analogously, the increase in genetic diversity that occurs through introduction of many new rare variants into urban areas contributes to an overall increase in h.
The degree to which the Dalmatian situation is a general one depends on how similar the effective population sizes and genetic structuring in the Dalmatian metapopulation are to other rural communities undergoing urbanisation over the last century. The villages studied here vary in size from 200–1800 inhabitants today and were historically considerably larger. Endogamy rates are high with 42–98% (mean 75%) of individuals having four grandparents from their village, reflected in a mean FST of approximately 2% for autosomal microsatellite markers. Comparisons among the European countries give FST values of about 1%, but there is a paucity of studies focusing on rural areas within countries. Pairwise FST values up to 1.2% for autosomal microsatellite markers were observed among rural populations in Scotland,15 which now mix and mate in urban centres across the country. Classical marker studies in Italy, such as those of 37 villages in the Parma valley, have also revealed very fine scale structure with an overall FST of 2.6%.21 Such differentiation was probably the norm in rural Europe and the changes in heterozygosity inferred here are thus likely to be commonplace. Apportioning the genetic variation among continents and populations on a global scale reveals that Europeans have the smallest among-population variance component.22 Urbanisation in other continents, thus, has the scope to generate larger outbreeding effects than are seen here, which will reach a maximum in the case of admixture between continental populations in the New World and elsewhere.
Although we used two different h estimation methods in this paper, we tend to give more weight to the FA over sMLH, as we consider it more suitable in substructured population with significant inbreeding. This is because sMLH is a robust method that does not distinguish between an individual's homozygosity for rare and common alleles, whereas FA gives more weight to homozygosity of rare than common alleles.16 Therefore, although sMLH has been consistently shown to be a very useful method for studying heterozygosity levels in natural animal and plant populations with fewer number of markers used in genotyping,18, 19 we gave more weight to results based on FA measure in this manuscript as it provides more insight into homozygosity in rare genetic variation, which is one of the key features of recent inbreeding that could be affecting human quantitative traits with relevance to health.
This study, of a metapopulation of genetic isolates, illustrates the measurable and substantial differences in h that exist across modern human populations. We show that levels of h are increasing through the processes of urbanisation, isolate break-up and admixture. The wide range of variation observed in h is unlikely to be random or stochastic, as it is closely correlated with individual genetic histories, based on knowledge of the birthplace of the parents and grandparents of the participants. We suggest, therefore, that population substructure, recent inbreeding and population admixture are the underlying mechanisms of the variation in h that we have observed. Positive effects of heterosis and hybrid vigour have been amply demonstrated in animals and plants, especially in the F1 generation.23 Thus, the breakdown of the past population substructures and greater outbreeding resulting from increased human population mixing and urbanisation may be leading to increasing h and beneficial effects on a range of traits associated with human health and disease. It has also been proposed that the traits showing marked secular trends during the 20th century, such as IQ, height and age at menarche, could also have been influenced to some extent by increasing h over this period.17, 24, 25
References
Clark D : Urban World, Global City. UK: Routledge, 2004.
Darvasi A, Shifman S : The beauty of admixture. Nat Genet 2005; 37: 118–119.
Vitart V, Carothers AD, Suffolk R et al: Increased level of linkage disequilibrium in rural compared to urban communities: a factor to consider in association-study Design. Am J Hum Genet 2005; 76: 763–772.
Helgason A, Ingvadottir B, Hrafnkelsson B, Gulcher J, Stefansson K : An Icelandic example of the impact of population structure on association studies. Nat Genet 2005; 37: 90–95.
Falconer DS, Mackay TFC : Introduction to Quantitative Genetics, 4th edn. Harlow, UK: Prentice Hall, 1996.
Charlesworth B, Hughes KA : Age-specific inbreeding depression and components of genetic variance in relation to the evolution of senescence. Proc Natl Acad Sci USA 1996; 93: 6140–6145.
Wright A, Charlesworth B, Rudan I, Carothers A, Campbell H : A polygenic basis for late-onset disease. Trends Genet 2003; 19: 97–106.
World Health Organisation: World Health Report 2002: Reducing Risks, Promoting Healthy Life. Geneva: WHO, 2002.
Abney M, McPeek MS, Ober C : Broad and narrow heritabilities of quantitative traits in a founder population. Am J Hum Genet 2001; 68: 1302–1307.
Rudan I, Campbell H, Carothers A et al: Inbreeding and the genetic complexity of human hypertension. Genetics 2003; 163: 1011–1021.
Rudan I, Campbell H, Carothers AD, Hastie ND, Wright AF : Contribution of consanguinity to polygenic and multifactorial diseases. Nat Genet 2006; 38: 1224–1225.
Rudan I, Campbell H : Five reasons why inbreeding may have considerable effect on post-reproductive human health. Coll Antropol 2004; 28: 943–950.
Bittles AH, Mason WM, Greene J, Rao NA : Reproductive behaviour and health in consanguineous marriages. Science 1991; 252: 789–794.
Bittles AH, Neel JV : The costs of human inbreeding and their implications for variations at the DNA level. Nat Genet 1994; 8: 117–121.
Vitart V, Biloglav Z, Hayward C et al: 3000 years of solitude: extreme level of differentiation in the island isolates of the Dalmatian coast. Eur J Hum Genet 2006; 14: 478–487.
Carothers AD, Rudan I, Hayward C et al: Estimating human individual inbreeding coefficients: comparison of genealogical and marker heterozygosity approaches. Ann Hum Genet 2006; 70: 666–676.
Campbell H, Carothers AD, Rudan I et al: Effects of genome-wide heterozygosity on a range of biomedically relevant human quantitative traits. Hum Mol Genet 2007; 16: 233–241.
Hoffman JI, Boyd IL, Amos W : Exploring the relationship between parental relatedness and male reproductive success in the Antarctic fur seal, Arctocephalus gazelle. Evolution 2004; 58: 2087–2099.
Coltman DW, Pilkington JG, Smith JA, Pemberton JM : Parasite-mediated selection against inbred Soay sheep in a free-living, island population. Evolution 1999; 53: 1259–1267.
Hirschhorn JN, Daly MJ : Genome-wide association studies for common diseases and complex traits. Nat Rev Genet 2005; 6: 95–108.
Cavalli-Sforza LL, Feldman MW : Spatial subdivision of populations and estimates of genetic variation. Theor Popul Biol 1990; 37: 3–25.
Rosenberg NA, Pritchard JK, Weber JL et al: Genetic structure of human populations. Science 2002; 298: 2381–2385.
Charpentier M, Setchell JM, Prugnolle F : Genetic diversity and reproductive success in mandrills (Mandrillus sphinx). Proc Natl Acad Sci USA 2005; 102: 16723–16728.
Altukhov YP, Sheremet'eva VA, Rychkov YG : Heterosis as the cause of the secular trend in humans. Dokl Biol Sci 2000; 370: 43–46.
Mingroni MA : The secular rise in IQ: giving heterosis a closer look. Intelligence 2004; 32: 65–83.
Acknowledgements
Igor Rudan and Ozren Polasek were supported by Overseas Research Scheme and by the scholarship from the University of Edinburgh. Ozren Polasek is supported by the PhD scholarship from the University of Edinburgh. Igor Rudan and Zrinka Biloglav were supported by the British Scholarship Trust Fellowship. Zrinka Biloglav was supported by ‘Miroslav Cackovic’ fellowship of the Faculty of Medicine, University of Zagreb. The study was partially supported through the grants from the National Institutes of Health to Igor Rudan, Harry Campbell and James L Weber; Ministry of Science, Education and Sport of the Republic of Croatia to Igor Rudan (number 108-1080315-0302), and the grants from The British Council, The Wellcome Trust, The Royal Society and Medical Research Council to Harry Campbell and Igor Rudan and European Commission FP6 STRP grant number 018947 (LSHG-CT-2006-01947). The authors collectively thank medical students of the Faculty of Medicine, University of Zagreb, Croatia; local general practitioners and nurses in study populations; the employees of several other Croatian institutions, including but not limited to the Institute for Anthropological Research in Zagreb, Croatia; the University of Rijeka and Split, Croatia; Croatian Institute of Public Health; Institutes of Public Health in Split and Dubrovnik, Croatia for their individual help in planning and carrying out the field work related to the project.
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests statement
The authors declare that they have no competing financial interests.
Rights and permissions
About this article
Cite this article
Rudan, I., Carothers, A., Polasek, O. et al. Quantifying the increase in average human heterozygosity due to urbanisation. Eur J Hum Genet 16, 1097–1102 (2008). https://doi.org/10.1038/ejhg.2008.48
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2008.48
Keywords
This article is cited by
-
Genome-wide meta-analysis identifies novel loci associated with parathyroid hormone level
Molecular Medicine (2018)
-
Runs of homozygosity: windows into population history and trait architecture
Nature Reviews Genetics (2018)
-
Pairomics, the omics way to mate choice
Journal of Human Genetics (2013)
-
Does inbreeding affect N-glycosylation of human plasma proteins?
Molecular Genetics and Genomics (2011)
-
Genes predict village of origin in rural Europe
European Journal of Human Genetics (2010)