Abstract
CRISPR–Cas systems that provide defence against mobile genetic elements in bacteria and archaea have evolved a variety of mechanisms to target and cleave RNA or DNA1. The well-studied types I, II and III utilize a set of distinct CRISPR-associated (Cas) proteins for production of mature CRISPR RNAs (crRNAs) and interference with invading nucleic acids. In types I and III, Cas6 or Cas5d cleaves precursor crRNA (pre-crRNA)2,3,4,5 and the mature crRNAs then guide a complex of Cas proteins (Cascade-Cas3, type I; Csm or Cmr, type III) to target and cleave invading DNA or RNA6,7,8,9,10,11,12. In type II systems, RNase III cleaves pre-crRNA base-paired with trans-activating crRNA (tracrRNA) in the presence of Cas9 (refs 13, 14). The mature tracrRNA–crRNA duplex then guides Cas9 to cleave target DNA15. Here, we demonstrate a novel mechanism in CRISPR–Cas immunity. We show that type V-A Cpf1 from Francisella novicida is a dual-nuclease that is specific to crRNA biogenesis and target DNA interference. Cpf1 cleaves pre-crRNA upstream of a hairpin structure formed within the CRISPR repeats and thereby generates intermediate crRNAs that are processed further, leading to mature crRNAs. After recognition of a 5′-YTN-3′ protospacer adjacent motif on the non-target DNA strand and subsequent probing for an eight-nucleotide seed sequence, Cpf1, guided by the single mature repeat-spacer crRNA, introduces double-stranded breaks in the target DNA to generate a 5′ overhang16. The RNase and DNase activities of Cpf1 require sequence- and structure-specific binding to the hairpin of crRNA repeats. Cpf1 uses distinct active domains for both nuclease reactions and cleaves nucleic acids in the presence of magnesium or calcium. This study uncovers a new family of enzymes with specific dual endoribonuclease and endonuclease activities, and demonstrates that type V-A constitutes the most minimalistic of the CRISPR–Cas systems so far described.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Marraffini, L. A. CRISPR-Cas immunity in prokaryotes. Nature 526, 55–61 (2015)
Carte, J., Wang, R., Li, H., Terns, R. M. & Terns, M. P. Cas6 is an endoribonuclease that generates guide RNAs for invader defense in prokaryotes. Genes Dev. 22, 3489–3496 (2008)
Ebihara, A. et al. Crystal structure of hypothetical protein TTHB192 from Thermus thermophilus HB8 reveals a new protein family with an RNA recognition motif-like domain. Protein Sci. 15, 1494–1499 (2006)
Charpentier, E., Richter, H., van der Oost, J. & White, M. F. Biogenesis pathways of RNA guides in archaeal and bacterial CRISPR-Cas adaptive immunity. FEMS Microbiol. Rev. 39, 428–441 (2015)
Nam, K. H. et al. Cas5d protein processes pre-crRNA and assembles into a cascade-like interference complex in subtype I–C/Dvulg CRISPR-Cas system. Structure 20, 1574–1584 (2012)
Jore, M. M. et al. Structural basis for CRISPR RNA-guided DNA recognition by Cascade. Nature Struct. Mol. Biol. 18, 529–536 (2011)
Mulepati, S., Heroux, A. & Bailey, S. Crystal structure of a CRISPR RNA-guided surveillance complex bound to a ssDNA target. Science 345, 1479–1484 (2014)
Plagens, A., Richter, H., Charpentier, E. & Randau, L. DNA and RNA interference mechanisms by CRISPR-Cas surveillance complexes. FEMS Microbiol. Rev. 39, 442–463 (2015)
van der Oost, J., Westra, E. R., Jackson, R. N. & Wiedenheft, B. Unravelling the structural and mechanistic basis of CRISPR-Cas systems. Nature Rev. Microbiol. 12, 479–492 (2014)
Zhang, J. et al. Structure and mechanism of the CMR complex for CRISPR-mediated antiviral immunity. Mol. Cell 45, 303–313 (2012)
Staals, R. H. et al. RNA targeting by the type III-A CRISPR-Cas Csm complex of Thermus thermophilus . Mol. Cell 56, 518–530 (2014)
Samai, P. et al. Co-transcriptional DNA and RNA cleavage during type III CRISPR-Cas immunity. Cell 161, 1164–1174 (2015)
Chylinski, K., Le Rhun, A. & Charpentier, E. The tracrRNA and Cas9 families of type II CRISPR-Cas immunity systems. RNA Biol. 10, 726–737 (2013)
Deltcheva, E. et al. CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature 471, 602–607 (2011)
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012)
Zetsche, B. et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 163, 759–771 (2015)
Schunder, E., Rydzewski, K., Grunow, R. & Heuner, K. First indication for a functional CRISPR/Cas system in Francisella tularensis . Int. J. Med. Microbiol. 303, 51–60 (2013)
Makarova, K. S. et al. An updated evolutionary classification of CRISPR-Cas systems. Nature Rev. Microbiol. 13, 722–736 (2015)
Shmakov, S. et al. Discovery and functional characterization of diverse class 2 CRISPR-Cas systems. Mol. Cell 60, 385–397 (2015)
Haurwitz, R. E., Jinek, M., Wiedenheft, B., Zhou, K. & Doudna, J. A. Sequence- and structure-specific RNA processing by a CRISPR endonuclease. Science 329, 1355–1358 (2010)
Dong, D. et al. The crystal structure of Cpf1 in complex with CRISPR RNA. Naturehttp://dx.doi.org/10.1038/nature17944 (20 April 2016)
Jiang, F., Zhou, K., Ma, L., Gressel, S. & Doudna, J. A. A. Cas9-guide RNA complex preorganized for target DNA recognition. Science 348, 1477–1481 (2015)
Nishimasu, H. et al. Crystal structure of Cas9 in complex with guide RNA and target DNA. Cell 156, 935–949 (2014)
Sternberg, S. H., Redding, S., Jinek, M., Greene, E. C. & Doudna, J. A. DNA interrogation by the CRISPR RNA-guided endonuclease Cas9. Nature 507, 62–67 (2014)
Szczelkun, M. D. et al. Direct observation of R-loop formation by single RNA-guided Cas9 and Cascade effector complexes. Proc. Natl Acad. Sci. USA 111, 9798–9803 (2014)
Zhang, Y., Rajan, R., Seifert, H. S., Mondragon, A. & Sontheimer, E. J. DNase H activity of Neisseria meningitidis Cas9. Mol. Cell 60, 242–255 (2015)
Yang, W. An equivalent metal ion in one- and two-metal-ion catalysis. Nature Struct. Mol. Biol. 15, 1228–1231 (2008)
Vasu, K. et al. Increasing cleavage specificity and activity of restriction endonuclease KpnI. Nucleic Acids Res. 41, 9812–9824 (2013)
Nam, K. H. et al. Double-stranded endonuclease activity in Bacillus halodurans clustered regularly interspaced short palindromic repeats (CRISPR)-associated Cas2 protein. J. Biol. Chem. 287, 35943–35952 (2012)
Punetha, A., Sivathanu, R. & Anand, B. Active site plasticity enables metal-dependent tuning of Cas5d nuclease activity in CRISPR-Cas type I–C system. Nucleic Acids Res. 42, 3846–3856 (2014)
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010)
Robinson, J. T. et al. Integrative genomics viewer. Nature Biotechnol. 29, 24–26 (2011)
Thorvaldsdóttir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform. 14, 178–192 (2013)
Sampson, J. R. & Uhlenbeck, O. C. Biochemical and physical characterization of an unmodified yeast phenylalanine transfer RNA transcribed in vitro . Proc. Natl Acad. Sci. USA 85, 1033–1037 (1988)
McClelland, M., Hanish, J., Nelson, M. & Patel, Y. KGB: a single buffer for all restriction endonucleases. Nucleic Acids Res. 16, 364 (1988)
Herbert, S., Barry, P. & Novick, R. P. Subinhibitory clindamycin differentially inhibits transcription of exoprotein genes in Staphylococcus aureus . Infect. Immun. 69, 2996–3003 (2001)
Urban, J. H. & Vogel, J. Translational control and target recognition by Escherichia coli small RNAs in vivo . Nucleic Acids Res. 35, 1018–1037 (2007)
Pall, G. S. & Hamilton, A. J. Improved northern blot method for enhanced detection of small RNA. Nature Protocols 3, 1077–1084 (2008)
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997)
Edgar, R. C. MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinformatics 5, 113 (2004)
Waterhouse, A. M., Procter, J. B., Martin, D. M., Clamp, M. & Barton, G. J. Jalview Version 2–a multiple sequence alignment editor and analysis workbench. Bioinformatics 25, 1189–1191 (2009)
Robinson, J. T. et al. Integrative genomics viewer. Nature Biotechnol. 29, 24–26 (2011)
Gruber, A. R., Lorenz, R., Bernhart, S. H., Neubock, R. & Hofacker, I. L. The Vienna RNA websuite. Nucleic Acids Res. 36, 3429–3431 (2008)
Darty, K., Denise, A. & Ponty, Y. VARNA: Interactive drawing and editing of the RNA secondary structure. Bioinformatics 25, 1974–1975 (2009)
Acknowledgements
We thank K. Schmidt, F. Hille and A. Escalera Maurer for technical help. This work was funded by the Alexander von Humboldt Foundation (AvH Professorship), the German Federal Ministry for Education and Research, the Helmholtz Association, the German Research Foundation, the Max Planck Society, the Göran Gustafsson Foundation (Göran Gustafsson Prize from the Royal Swedish Academy of Sciences), the Swedish Research Council and Umeå University (all to E.C.), and the Helmholtz Postdoc Programme (to H.R.).
Author information
Authors and Affiliations
Contributions
I.F. and H.R. conducted the biochemical characterization of the DNase and RNase activities, M.B. performed binding studies and seed sequence characterization and A.L.R. performed and analysed RNA sequencing. I.F., H.R. and E.C. designed the research. I.F., H.R., M.B., A.L.R. and E.C. analysed and interpreted the data; I.F., H.R., and E.C. wrote the paper, which M.B and A.L.R. commented on.
Corresponding author
Ethics declarations
Competing interests
E.C. is a co-founder of CRISPR Therapeutics AG and ERS Genomics, and is a member of the scientific advisory board of CRISPR Therapeutics AG and Horizon Discovery Group.
Extended data figures and tables
Extended Data Figure 1 F. novicida U112 expresses short mature type V-A crRNAs composed of repeat-spacer.
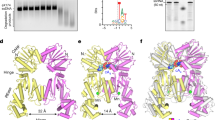
a, In-scale representation of type II-B (cas9) and type V-A (cpf1) CRISPR–Cas loci in F. novicida U112. Blue arrows, cas genes; black arrows, putative pre-crRNA promoters; thick black line, CRISPR leader sequence; black rectangles, CRISPR repeats; green diamonds, CRISPR spacers; red arrows, tracrRNA or scaRNA (small, CRISPR–Cas-associated RNA). b, Expression of type V-A crRNAs determined by sRNA sequencing is represented with a grey bar chart. The coverage of the reads is indicated in brackets and reads starting (5′ end) and ending (3′ end) at each position are shown (image captured from Integrative Genomics Viewer)33,42. The genomic coordinates and size of the CRISPR array in base pairs are indicated. Above the chart bar, a schematic representation shows repeats (black boxes) and spacers (green diamonds) of the F. novicida U112 CRISPR array. The sequence of the type V-A CRISPR array from the leader sequence to the last repeat is shown below. Black bold sequences, repeats; green sequences, spacers. The boxed sequences correspond to the mature crRNAs for which we could determine the 5′ and 3′ ends. The mature crRNAs are composed of part of the repeat in 5′ and part of the spacer in 3′.
Extended Data Figure 2 Wild-type Cpf1 purifies as a monomer in solution.
Recombinant Cpf1 of F. novicida U112 purified via affinity and cation-exchange chromatography was applied to a Superdex 200 size-exclusion column. a, SDS–PAGE of protein eluates obtained by nickel-affinity purification (left panel), which were further purified by cation-exchange chromatography (right panel). b, Protein samples obtained by size-exclusion chromatography were separated by SDS–PAGE (8% polyacrylamide) and visualized with Coomassie staining. c, Elution profile of the size-exclusion chromatography of wild-type Cpf1. The partition coefficient Kav for Cpf1 was calculated as 0.0538 by using the equation Kav = (Ve −V0)/(Vt −V0), with Ve, elution volume; V0, void volume (elution volume of blue dextran, 45.171 ml) and Vt, geometric column volume (482.5 ml). d, Calibration curve of proteins with known molecular weights (thyroglobulin (669 kDa), ferritin (443 kDa), β-amylase (200 kDa), alcohol dehydrogenase (ADH; 150 kDa), bovine serum albumin (BSA; 66 kDa), carbonic anhydrase (29 kDa); Molecular Weight Marker Kit, Sigma-Aldrich). The molecular weight of these proteins was plotted against their calculated Kav and fitted by exponential regression analysis. On the basis of the calculation, the Kav of Cpf1 results in a molecular weight of 187 kDa, indicating a monomeric form of Cpf1 in solution. Assuming that the protein does not adopt a perfect globular shape, this result is in accordance with the theoretical molecular weight of Cpf1 (153 kDa).
Extended Data Figure 3 The endoribonucleolytic activity of Cpf1 is dependent on the presence of an intact repeat sequence.
a, Cleavage assays were performed by incubating 100 nM of internally labelled RNA constructs corresponding to different repeat and spacer sequence variants of pre-crRNA-sp5 (pre-crRNA-containing spacer 5) with 1 μM of Cpf1 for 30 min at 37 °C. The cleavage reaction was analysed by denaturing polyacrylamide gel electrophoresis and phosphorimaging. The cleavage products are represented schematically. The sequence compositions of the RNAs used as substrates are shown. RNA structures were generated with RNAfold43 and visualized using VARNA44 software. Cpf1 cleaved only the RNA templates containing a full-length repeat sequence. The substrate containing two repeats was cleaved twice resulting in more than two fragments, whereas cleavage of RNA with only one repeat resulted in two fragments, consistent with the determined cleavage site (see Fig. 1). b, Northern blot analysis of total RNA extracted from E. coli co-transformed with a plasmid encoding pre-crRNA and either the empty vector or overexpression vectors encoding wild-type (wt) Cpf1 and variants. Cpf1 expression was induced (+) or not induced (−) with IPTG. The northern blot was probed against the spacer sequence of the tested pre-crRNA. In the absence of Cpf1 (empty vector or not induced), the amount of transcript is reduced compared to reactions with Cpf1 present, suggesting stabilization of pre-crRNA upon binding of Cpf1. Expression of Cpf1, Cpf1(K852A) and Cpf1(K869A) results in the production of a distinct processed transcript of 65 nt, whereas expression of Cpf1(H843A), Cpf1(K852A) or Cpf1(K869A) results in the production of additional higher transcripts. Expression of Cpf1(F873A) results in almost no detectable processed transcript.
Extended Data Figure 4 Cpf1 is a sequence- and structure-specific endoribonuclease.
a, Design of various repeat variants of pre-crRNA-sp5 (pre-crRNA with spacer 5) with an altered repeat sequence, a destroyed repeat structure, single nucleotide exchanges (1–4) in the RRS (purple) and changed loop (green) and stem sizes (yellow). Note that the 5′ repeat region of the wild-type repeat is not shown in the different variants. Red circles highlight the mutated or added residues. The RNA structures were generated with RNAfold43 and visualized using VARNA44 software. b, Internally labelled pre-crRNAs containing a wild-type repeat sequence, an altered repeat sequence or a destroyed repeat structure were obtained by in vitro transcription. The 5′ end-labelled wild-type substrate was used to generate an alkaline hydrolysis ladder (OH) and an RNase T1 digest (T1) for size determination of the RNA fragments (Life Technologies). Cpf1 cleaved only the pre-crRNA template containing the wild-type repeat sequence yielding a small 19-nt 5′ repeat fragment and a 50-nt intermediate crRNA. c, Substrates with serial single mutations of the four RRS nucleotides (1–4, counting from the cleavage site) were tested for processing by Cpf1. Changes of the first three nucleotides were not tolerated for Cpf1-mediated processing, whereas changing the fourth nucleotide yielded a substrate that was processed with less efficiency compared to the wild-type substrate. d, The influence of loop variations in the repeat was tested with substrates containing +1 or −1 nucleotide in the loop. Both substrates were processed by Cpf1. Stems with +1 or −1 base pair, or +4 base pairs were used to determine length requirements of the stem. Cpf1 did not cleave any of the three substrates tested. The RNA cleavage reactions were performed by incubating 1 μM of Cpf1 with 200 nM of RNA variant at 37 °C for 5 min in the presence of 10 mM MgCl2. The cleavage products were analysed by denaturing polyacrylamide gel electrophoresis and phosphorimaging. RNA fragments are represented schematically and fragment sizes are indicated in nucleotides.
Extended Data Figure 5 RNA and DNA cleavage activities of Cpf1 are dependent on divalent metal ions.
a, Cleavage assays of pre-crRNA-sp5 (repeat-spacer 5, full-length, RNA 4, Extended Data Fig. 6) by Cpf1 in KGB buffer supplemented with different concentrations of divalent metal ion (indicated in mM) or EDTA (10 mM). Cleavage products were analysed by denaturing polyacrylamide gel electrophoresis and visualized by phosphorimaging. RNA fragments are represented schematically and fragment sizes are indicated in nucleotides. Specific RNA cleavage was observed in the presence of MgCl2. Less specific cleavage was detected with CaCl2, MnCl2 and CoCl2. No cleavage of pre-crRNA-sp5 was detected in presence of NiCl2 and ZnCl2. b, Cleavage assays of supercoiled plasmid DNA containing protospacer 5 by Cpf1 programmed with crRNA-sp5 (repeat-spacer 5, processed, RNA3, Extended Data Fig. 6) in KGB buffer supplemented with different concentrations of divalent metal ions (indicated in mM). Cleavage products were analysed by agarose gel electrophoresis and visualized by EtBr staining. DNA cleavage was observed in the presence of MgCl2 and MnCl2. A more specific cleavage was observed in the presence of CaCl2. The addition of CoCl2, NiCl2 or ZnCl2 to the reaction did not result in DNA cleavage. li, linear; sc, supercoiled; M, 1 kb ladder (Fermentas). Quantification of three independent experiments shown in Extended Data Table 1b.
Extended Data Figure 6 Cpf1 requires crRNA with an intact repeat structure to specifically cleave DNA.
a, Cleavage assays of protospacer 5 containing supercoiled plasmid DNA by Cpf1 programmed with different RNA constructs (1, RNA construct containing spacer 4; 2–8, RNA constructs containing spacer 5) in the presence of 10 mM CaCl2. Cleavage products were analysed by agarose gel electrophoresis and visualized by EtBr staining. b, Cleavage of 5′-radiolabelled oligonucleotide duplexes containing protospacer 5 in the presence of 10 mM CaCl2. Cleavage products were analysed by denaturing polyacrylamide gel electrophoresis and visualized by phosphorimaging. Fragment sizes are indicated in nucleotides. RNA structures were generated with RNAfold43 and visualized using VARNA44 software. Only the RNAs containing a full-length repeat and a spacer complementary to the target mediate DNA cleavage by Cpf1.
Extended Data Figure 7 Analysis of target DNA cleavage by crRNA-programmed Cpf1 in presence of Mg2+.
a, Cleavage assays of protospacer 5 containing supercoiled plasmid DNA by Cpf1 programmed with crRNA-sp4 or crRNA-sp5 (crRNA-sp4, repeat-spacer 4, processed, RNA 1, Extended Data Fig. 6; crRNA-sp5, repeat-spacer 5, processed, RNA 3, Extended Data Fig. 6) in absence or presence of 10 mM MgCl2. Plasmid DNA cleavage was observed only with Cpf1 programmed with crRNA-sp5 in presence of Mg2+. b, Oligonucleotide cleavage assays using Cpf1 programmed with crRNA-sp5 (repeat-spacer 5, processed, RNA 3, Extended Data Fig. 6) in presence of 10 mM MgCl2. Either the target or the non-target strand was 5′-radiolabelled before annealing to the non-labelled complementary strand to form the duplex substrate. c, Sequencing analysis of the cleavage product obtained in a. The termination of the sequencing reaction indicates the cleavage site. Note that an enhanced signal for adenine is a sequencing artefact. d, Plasmid DNA containing protospacer 5 and the PAMs 1–6, or 5′-radiolabelled double-stranded oligonucleotide containing protospacer 5 and PAMs 1, 7–9 were subjected to cleavage by Cpf1 programmed with crRNA-sp5 (repeat-spacer, full-length, RNA 4, Extended Data Fig. 6) in the presence of 10 mM MgCl2 (upper and lower panel, respectively). Oligonucleotide cleavage products are indicated in nucleotides.
Extended Data Figure 8 Binding studies of Cpf1.
a, EMSAs of 5′-radiolabelled double-stranded oligonucleotides containing protospacer 5 by Cpf1 programmed with RNA 1–6 (see Extended Data Fig. 6). The protein concentrations used were 8, 52 and 512 nM. Reactions were analysed by native PAGE and phosphorimaging. Unbound and bound DNAs are indicated. Higher DNA binding affinities are observed when Cpf1 is programmed with an RNA containing an entire repeat sequence. b, EMSAs of 5′-radiolabelled crRNA-sp5 (repeat-spacer 5, processed, RNA 3, Extended Data Fig. 6) by wild-type Cpf1, Cpf1(H843A), Cpf1(K852A), Cpf1(K869A) and Cpf1(F873A). The protein concentrations used were 2, 4, 8, 12, 16, 24, 32, 48 and 64 nM. Reactions were analysed by native polyacrylamide gel electrophoresis and phosphorimaging. Unbound and bound RNAs are indicated. Shown are representatives of at least three individual experiments. The bound and unbound RNA fractions were quantified, plotted against the enzyme concentration and fitted by nonlinear regression analysis. The calculated Kd values (± s.d.) were 16 ± 1 nM (wild type), 17 ± 0.5 nM (H843A), 12 ± 1 nM (K852A), 10 ± 1 nM (K869A) and 17 ± 1 nM (F873A). There are no differences between the RNA binding affinities of wild-type and mutant Cpf1. c, EMSAs of 5′-radiolabelled double-stranded oligonucleotides containing protospacer 5 targeted by wild-type Cpf1, Cpf1(D917A), Cpf1(E1006A) and Cpf1(D1255A) in complex with crRNA-sp5 (repeat-spacer 5, full-length, RNA 4, Extended Data Fig. 6). The protein concentrations used were 8, 16, 32, 42, 52, 64, 74, 128 and 256 nM. Reactions were analysed by native polyacrylamide gel electrophoresis and phosphorimaging. Unbound and bound DNAs are indicated. Shown are representative of at least three individual experiments. The bound and unbound DNA fractions were quantified, plotted against the enzyme concentration and fitted by nonlinear regression analysis. The calculated Kd values (± s.d.) were 50 ± 3 nM (wild type), 48 ± 8 nM (D917A), 40 ± 8 nM (E1006A) and 52 ± 6 nM (D1255A). There are no differences between the RNA-mediated DNA binding affinities of wild-type and mutant Cpf1. The reduced Kd for E1006A can be explained by the removal of the large negatively charged amino acid, which might facilitate interaction of Cpf1 with the DNA.
Extended Data Figure 9 Processing activity of Cpf1 is specific for pre-crRNA and crRNA-mediated targeting of Cpf1 is directed only against single- and double-stranded DNA.
a, Cpf1 processing activity was tested against pre-crRNA and pre-crDNA. Wild-type Cpf1 or Cpf1(D917A) (1 μM) was incubated with 200 nM internally labelled pre-crRNA-sp5 (repeat-spacer 5, full-length, RNA 4, Extended Data Fig. 6) or a 5′-labelled ssDNA (pre-crDNA-sp5) construct with the same sequence as the RNA in KGB buffer with 10 mM MgCl2 for 5 min at 37 °C. Incubation of wild-type Cpf1 and DNase inactive mutant (Cpf1(D917A)) with the RNA construct, but not the DNA construct, resulted in the expected cleavage products of a 19-nt repeat fragment and a 50-nt intermediate crRNA, indicating that the processing activity of Cpf1 is specific for RNA. b, crRNA-mediated DNA cleavage activity of Cpf1. Cpf1 (100 nM) in complex with crRNA-sp5 (repeat-spacer 5, full-length, RNA 4, Extended Data Fig. 6) was incubated with 10 nM of 5′-radiolabelled ssRNA, dsRNA, ssDNA, dsDNA or RNA–DNA hybrids in KGB buffer with either MgCl2 (10 mM; upper panel) or CaCl2 (10 mM; lower panel) for 1 h at 37 °C. The oligonucleotide DNA substrates contained the sequence for protospacer 5 targeted by the tested crRNA. For DNA–RNA hybrids, the 5′-radiolabelled target strand is indicated with an asterisk. Only ssDNA and dsDNA substrates were cleaved, indicating that the crRNA-mediated cleavage activity of Cpf1 is only directed against DNA substrates. The cleavage products for ssDNA, however, vary from those expected or observed for dsDNA. Cleavage reactions were analysed by denaturing polyacrylamide gel electrophoresis and phosphorimaging. RNA cleavage products are indicated schematically. RNA and DNA fragment sizes are given in nucleotides.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-2, Supplementary Table 1 and Supplementary References. (PDF 2039 kb)
Rights and permissions
About this article
Cite this article
Fonfara, I., Richter, H., Bratovič, M. et al. The CRISPR-associated DNA-cleaving enzyme Cpf1 also processes precursor CRISPR RNA. Nature 532, 517–521 (2016). https://doi.org/10.1038/nature17945
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature17945
This article is cited by
-
An Update on the Application of CRISPR Technology in Clinical Practice
Molecular Biotechnology (2024)
-
Development of a CRISPR/Cpf1 system for multiplex gene editing in Aspergillus oryzae
Folia Microbiologica (2024)
-
Expansion of the CRISPR/Cas Genome-Sculpting Toolbox: Innovations, Applications and Challenges
Molecular Diagnosis & Therapy (2021)
-
The working dead: repurposing inactive CRISPR-associated nucleases as programmable transcriptional regulators in plants
aBIOTECH (2020)
-
Cas12a mediates efficient and precise endogenous gene tagging via MITI: microhomology-dependent targeted integrations
Cellular and Molecular Life Sciences (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.