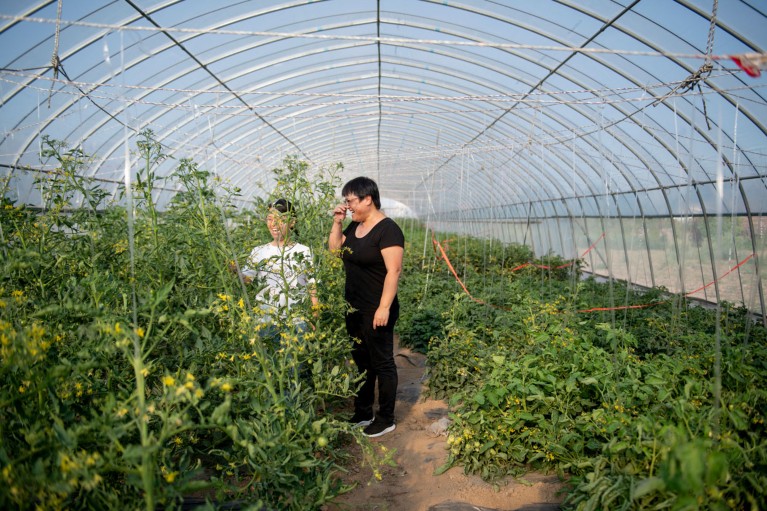
Caixia Gao and a member of her team inspect CRISPR-modified tomato plants in a greenhouse at their growing facility in Beijing. Credit: Stefen Chow
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 604, 790-792 (2022)
doi: https://doi.org/10.1038/d41586-022-01117-z
Updates & Corrections
-
Correction 10 May 2022: An earlier version of this Technology feature article erroneously implied that Pairwise’s leafy green vegetable was being base-edited to improve the nutritional content. In fact, the firm used standard CRISPR and targeted flavour. Also, the stoneless cherry is not yet in field testing.
References
Xing, S. et al. Genome Biol. 21, 230 (2020).
Komor, A. et al. Nature 533, 420–424 (2016).
Gaudelli, N. et al. Nature 551, 464–471 (2017).
Bastet, A. et al. Plant Biotechnol. J. 17, 1736–1750 (2019).
Zhang, R. et al. Nature Plants 5, 480–485 (2019).
Li, C. et al. Nature Biotechnol. 38, 875–882 (2020).
Sretenovic, S. et al. Front. Genome Ed. 3, 756766 (2021).
Anzalone, A. V. et al. Nature 576, 149–157 (2019).
Zong, Y. et al. Nature Biotechnol. https://doi.org/10.1038/s41587-022-01254-w (2022).
Lin, Q. et al. Nature Biotechnol. 39, 923–927 (2021).
Kong, J. et al. Front. Plant Sci. https://doi.org/10.3389/fpls.2020.572319 (2020).
Li, T. et al. Nature Biotechnol. 36, 1160–1163 (2018).

 Super-precise new CRISPR tool could tackle a plethora of genetic diseases
Super-precise new CRISPR tool could tackle a plethora of genetic diseases
 Got mutation? ‘Base editors’ fix genomes one nucleotide at a time
Got mutation? ‘Base editors’ fix genomes one nucleotide at a time
 NatureTech hub
NatureTech hub