- TECHNOLOGY FEATURE
Artificial intelligence powers protein-folding predictions
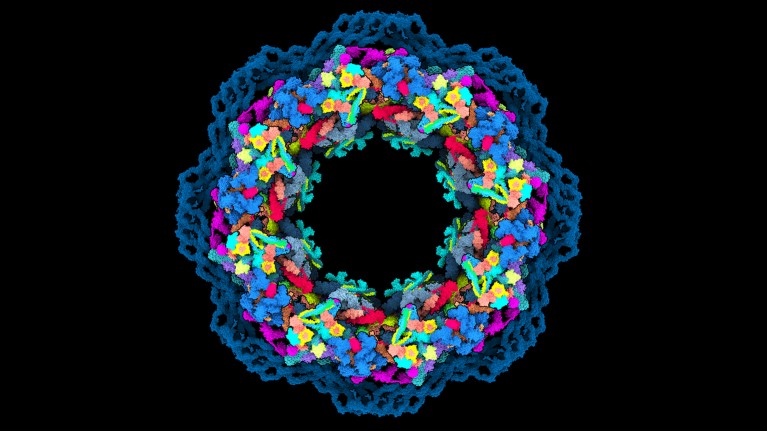
A model of the human nuclear pore complex, built using AlphaFold2 and structural data. Credit: Agnieszka Obarska-Kosinska
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 599, 706-708 (2021)
doi: https://doi.org/10.1038/d41586-021-03499-y
References
Porta-Pardo, E., Ruiz-Serra, V. & Valencia, A. Preprint at bioRxiv https://doi.org/10.1101/2021.08.03.454980 (2021).
Jumper, J. et al. Nature 596, 583–589 (2021).
Tunyasuvunakool, K. et al. Nature 596, 590–596 (2021).
Zheng, W. et al. Proteins https://doi.org/10.1002/prot.26193 (2021).
Yang, J. et al. Proc. Natl Acad. Sci. USA 117, 1496–1503 (2020).
Xu, J. Proc. Natl. Acad. Sci. USA 116, 16856–16865 (2019).
Marks, D. S. et al. PLoS ONE 6, e28766 (2011).
Baek, M. et al. Science 373, 871–876 (2021).
Mirdita, M. et al. Preprint at bioRxiv https://doi.org/10.1101/2021.08.15.456425 (2021).
Bryant, P., Pozzati, G. & Elofsson, A. Preprint at bioRxiv https://doi.org/10.1101/2021.09.15.460468 (2021).
Humphreys, I. R. et al. Science https://doi.org/10.1126/science.abm4805 (2021).
Evans, R. et al. Preprint at bioRxiv https://doi.org/10.1101/2021.10.04.463034 (2021).
Akdel, M. et al. Preprint at bioRxiv https://doi.org/10.1101/2021.09.26.461876 (2021).

 DeepMind’s AI predicts structures for a vast trove of proteins
DeepMind’s AI predicts structures for a vast trove of proteins
 DeepMind’s AI for protein structure is coming to the masses
DeepMind’s AI for protein structure is coming to the masses
 ‘It will change everything’: DeepMind’s AI makes gigantic leap in solving protein structures
‘It will change everything’: DeepMind’s AI makes gigantic leap in solving protein structures
 Cryo-electron microscopy shapes up
Cryo-electron microscopy shapes up
 NatureTech hub
NatureTech hub