How might the emergence of SARS-CoV-2 variants affect efforts to control the COVID-19 pandemic? The threat posed by such variants is focusing attention on vaccination and therapeutic options to grapple with the evolving coronavirus. Writing in Nature, Xu et al.1 describe the development of a genetically engineered mouse that can generate antibodies similar to those produced by camelids (an animal grouping that includes camels and llamas). These antibodies recognize targets using a single, small protein domain called a nanobody, also known as a VHH domain. Vaccination of these mice using proteins based on the SARS-CoV-2 spike protein resulted in the generation of antiviral nanobodies. These nanobodies could be produced in formats that were highly effective against COVID-19 variants that are impervious to many conventional antibodies being developed as therapies.
Conventional antibodies such as those produced by humans and mice recognize antigens (protein fragments of disease-causing agents) by means of two variable domains (VH and VL), which are components of separate heavy- and light-chain proteins (Fig. 1). By contrast, camelids and cartilaginous fishes (such as sharks) can make heavy-chain-only antibodies that recognize antigens using single, variable VHH domains, or nanobodies. One advantage of nanobodies is their small size, which enables them to penetrate tissues and recognize epitopes (the region of an antigen to which an antibody binds) that are normally inaccessible to conventional antibodies.
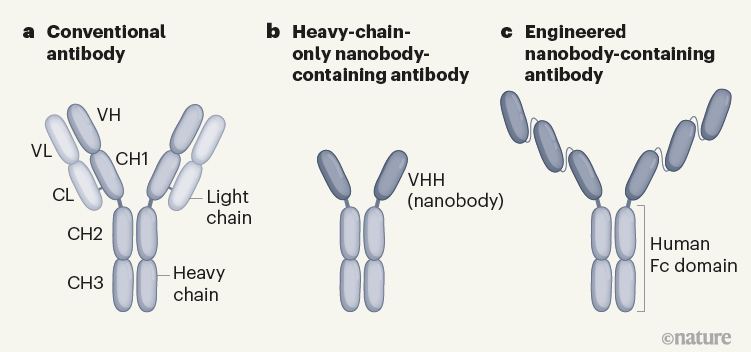
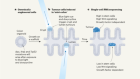
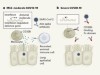
Figure 1 | Different types of antibody can target SARS-CoV-2. a, Human and mouse antibodies bind to their targets using two variable domains (VH and VL) on separate light- and heavy-chain proteins. These light and heavy chains pair through a connection between the light-chain CL domain and the heavy-chain CH1 domain. The paired domains are part of the constant regions of the heavy and light chains. The heavy-chain constant region consists of two other domains (CH2 and CH3, which form what is called the Fc domain) that help antibodies to travel around the body and interact with other components of the immune system. b, Camelid animals, such as camels and llamas, can produce heavy-chain-only antibodies because some of their genes encode heavy-chain proteins that lack CH1 domains. These antibodies recognize their targets using only a single variable domain (VHH) region, which is also known as a nanobody. Xu et al.1 have developed mice that make a similar type of antibody. c, The authors genetically engineered mice to produce antibodies consisting of heavy chains containing three tandem copies of camelid nanobodies and a human Fc domain. They report that these antibodies could prevent SARS-CoV-2 from infecting human cells when tested in vitro, and that such ‘neutralizing’ antibodies were effective against SARS-CoV-2 variants of concern that emerged during the second and third waves of the pandemic.
Nanobodies are generally extremely stable and soluble, and their modular nature means they can be readily expressed alone or in a variety of formats: for example, fused to the human antibody Fc domain that boosts defence responses2. These features make nanobody-based therapeutics a promising alternative to conventional monoclonal antibodies (antibodies with heavy and light chains that have a particular amino-acid sequence and antigen specificity). However, although 2021 saw regulatory approval of the 100th monoclonal-antibody treatment3, only one nanobody-based therapy has been approved for clinical use by the US Food and Drug Administration (FDA)4.
Currently, the only human monoclonal antibodies in advanced development as COVID-19 therapies (see go.nature.com/3xt9ku2) are a type called neutralizing antibodies (which block viral entry). Most of these were obtained from the antibody-producing cells of people who were infected during the first wave of the pandemic. Such antibodies target the receptor-binding domain (RBD) of the spike protein; the virus uses this domain to bind to the receptor that enables it to infect cells5. Human monoclonal antibodies are strongly preferred for clinical development because they are highly specific, are easily manufactured and work in concert with, and are well tolerated by, the human immune system6. However, there are strong arguments in favour of developing alternative, nanobody-based therapeutics in response to evolving human respiratory viruses such as SARS-CoV-2.
Read the paper: Nanobodies from camelid mice and llamas neutralize SARS-CoV-2 variants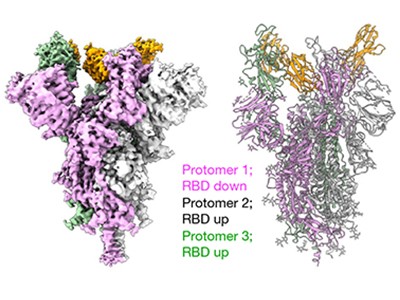
Since the virus began infecting people more than a year ago, the collective human neutralizing-antibody response generated against SARS-CoV-2 has applied a strong selection pressure on the spike-protein RBD. In fact, only two or three amino-acid-residue changes in versions of the virus that became dominant during the second and third waves of the pandemic were enough to render those versions substantially more resistant to neutralization by antibodies generated during the first wave — as assessed by tests on blood samples, known as convalescent serum, from individuals who have had COVID-197.
It therefore follows that monoclonal-antibody therapies mined from antibody responses generated during the first wave of the pandemic might quickly become obsolete. RBD epitopes of the spike protein that are recognized by neutralizing nanobodies, and that are not under selection pressure as a consequence of epitope recognition by human antibodies, might provide COVID-19 antivirals that do not quickly become ineffective as viral variants emerge. Broadly neutralizing nanobodies — nanobodies that recognize evolutionarily conserved epitopes of the spike protein — might even be useful against other coronaviruses with the capacity to drive a future pandemic.
The small and soluble nature of nanobodies means that they should be inexpensive to produce and easy to administer directly by inhalation to target key initial sites of viral replication in the respiratory tract. A study8 in hamsters, assessing the intranasal aerosol delivery of neutralizing nanobodies targeting SARS-CoV-2, reports that nanobodies were effectively deposited throughout the animals’ respiratory tract, and that this treatment notably reduced the level of virus. As long as nanobody therapies are administered only once over a short period of time to people with an acute infection, then strong immune responses should not be directed towards the nanobody itself, rendering it ineffective. The issue of such immune responses is generally a concern in relation to monoclonal-antibody therapies being developed to treat diseases that require repeated antibody administration over longer periods of time.
Some antibodies can dampen antiviral defences in people with severe COVID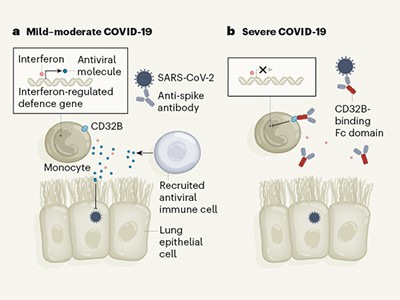
Although there has been some hesitancy in moving nanobodies to the clinic, many studies have found that potently neutralizing nanobodies can be elicited in camelids using SARS-CoV-2 vaccines based on the spike protein9–13. Xu and colleagues now offer a way forward through their generation of heavy-chain-only antibody-producing ‘nanomice’. The approach offers a system that should make nanobody discovery easier, faster and less expensive than was previously possible. Laboratory facilities to care for mice are inexpensive and ubiquitous, the mouse immune system is well understood, and high-quality tools such as those needed for cell sorting are readily available. Furthermore, immunizations in mice can occur on a much faster timescale than is possible for larger animal models — an important consideration when a rapid response to a newly emerging pandemic is required.
To generate these nanomice, Xu et al. replaced a large region of genomic DNA, containing all of the mouse heavy-chain variable (V) genes, with a region of DNA comprising 30 heavy-chain V genes derived from alpaca, dromedary and Bactrian camels. Each gene was fused to a DNA sequence that enabled the gene to form the usual connection (through a process termed recombination) to mouse heavy-chain D and J genes to make complete VHH genes. The V genes were also fused to promoter DNA sequences so that the VHH genes could be expressed in mouse antibody-producing B cells. Each developing B cell could indeed recombine a single camelid V, mouse D and mouse J gene to generate B-cell populations expressing different VHH-gene sequences as heavy-chain-only antibodies. The authors demonstrated that these cells could respond normally to immunization, undergoing a process (termed affinity maturation) that boosts the potency and specificity of antibodies for the antigen that they respond to.
All for one and one for all to fight flu
Xu and colleagues then immunized three nanomice and one llama with the SARS-CoV-2 spike protein and RBD. They identified neutralizing nanobodies in both animal models. These nanobodies could be formatted to be expressed as three tandem nanobody copies fused to a human antibody Fc domain (Fig. 1). This domain is a key feature of conventional antibodies: it enables an antibody to transit around the body, improves the antibody’s lifespan and boosts interactions with other components of the immune system. This format using tandem nanobodies should help to boost antigen binding by the antibodies. The authors’ evidence indicates that these engineered antibodies could potently neutralize all of the tested SARS-CoV-2 variants of concern (viruses with RBD mutations and which were associated with the second and third waves of the pandemic). Moreover, the proteins from nanomice recognized evolutionarily conserved epitopes on the RBD that do not overlap with regions commonly recognized by human antibodies.
The COVID-19 pandemic presents a unique opportunity for nanobodies to shine in the clinic, and the nanomouse platform is poised to help bring higher-quality therapeutic-nanobody options to the table, pushing the odds of success even higher. Just as the development of mice with antibodies containing human variable domains (such as Regeneron’s VelocImmune mouse) has helped to deliver the 100th FDA-approved monoclonal antibody, perhaps the nanomouse will give nanobody-based therapeutics a push in the same direction.

 Read the paper: Nanobodies from camelid mice and llamas neutralize SARS-CoV-2 variants
Read the paper: Nanobodies from camelid mice and llamas neutralize SARS-CoV-2 variants
 Some antibodies can dampen antiviral defences in people with severe COVID
Some antibodies can dampen antiviral defences in people with severe COVID
 All for one and one for all to fight flu
All for one and one for all to fight flu