- NEWS AND VIEWS
Designer fibre meals sway human gut microbes
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 595, 32-34 (2021)
doi: https://doi.org/10.1038/d41586-021-01601-y
References
Delannoy-Bruno, O. et al. Nature 595,91–95 (2021).
Chen, R. Y. et al. N. Engl. J. Med. 384, 1517–1528 (2021).
Chen, R. Y. et al. N. Engl. J. Med. 383, 321–333 (2020).
Mostafa, I. et al. BMC Public Health 20, 242 (2020).
Patnode, M. L. et al. Cell 179, 59–73 (2019).
Gehrig, J. L. et al. Science 365, 139 (2019).
Zeevi, D. et al. Cell 163, 1079–1094 (2015).
Berry, S. E. et al. Nature Med. 26, 964–973 (2020).
US Department of Health and Human Services and US Department of Agriculture. 2015–2020 Dietary Guidelines for Americans (HHS/USDA, 2015).
Sonnenburg, E. D. et al. Nature 529, 212–215 (2016).
Eilam, O. et al. mBio 5, e01526-14 (2014).
Healey, G. et al. Br. J. Nutr. 119, 176–189 (2018).
Deehan, E. C. et al. Cell Host Microbe 27, 389–404 (2020).
Al-Asmakh, M. & Zadjali, F. J. Microbiol. Biotechnol. 25, 1583–1588 (2015).
Lundberg, R., Toft, M. F., August, B., Hansen, A. K. & Hansen, C. H. F. Gut Microbes 7, 68–74 (2016).
Competing Interests
E.E. is the is the scientific founder of DayTwo and BiomX.

 Read the paper: Evaluating microbiome-directed fibre snacks in gnotobiotic mice and humans
Read the paper: Evaluating microbiome-directed fibre snacks in gnotobiotic mice and humans
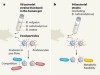 Food for thought about manipulating gut bacteria
Food for thought about manipulating gut bacteria
 Ancient human faeces reveal gut microbes of the past
Ancient human faeces reveal gut microbes of the past