- NEWS AND VIEWS
A molecular connection hints at how a genetic risk factor drives Crohn’s disease
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 593, 201-203 (2021)
doi: https://doi.org/10.1038/d41586-021-00979-z
References
Ng, S. C. et al. Lancet 390, 2769–2778 (2017).
Nayar, S. et al. Nature 593, 275–281 (2021).
Rieder, F., Zimmermann, E. M., Remzi, F. H. & Sandborn, W. J. Gut 62, 1072–1084 (2013).
Cuthbert, A. P. et al. Gastroenterology 122, 867–874 (2002).
Sidiq, T., Yoshihama, S., Downs, I. & Kobayashi, K. S. Front. Immunol. 7, 367 (2016).
Franchi, L., Warner, N., Viani, K. & Nuñez, G. Immunol. Rev. 227, 106–128 (2009).
Brugman, S. Dev. Comp. Immunol. 64, 82–92 (2016).
Duggan, S. T. & McKeage, K. Drugs 71, 2193–2212 (2011).
Hilmi, I., Tan, Y. M. & Goh, K. L. World J. Gastroenterol. 12, 1435–1438 (2006).
Mondal, K. & Kugathasan, S. Nature Rev. Gastroenterol. Hepatol. 14, 266–268 (2017).

 Read the paper: A myeloid–stromal niche and gp130 rescue in NOD2-driven Crohn’s disease
Read the paper: A myeloid–stromal niche and gp130 rescue in NOD2-driven Crohn’s disease
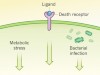 Stressful genetics in Crohn’s disease
Stressful genetics in Crohn’s disease
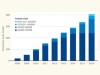 A decade of shared genomic associations
A decade of shared genomic associations