Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 592, 509-510 (2021)
doi: https://doi.org/10.1038/d41586-021-00888-1
Updates & Corrections
-
Clarification 15 April 2021: The original article did not specify that HRASLS proteins are also known as PLAAT proteins.
References
GBD 2019 Blindness and Vision Impairment Collaborators. Lancet Glob. Health 9, e144–e160 (2021).
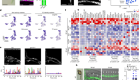
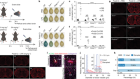
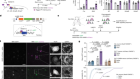
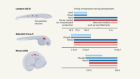
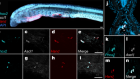
Morishita, H. et al. Nature 592, 634–638 (2021).
Wistow, G. J. & Piatigorsky, J. Annu. Rev. Biochem. 57, 479–504 (1988).
Cvekl, A. & Ashery-Padan, R. Development 141, 4432–4447 (2014).
Wride, M. A. Phil. Trans. R. Soc. Lond. B 366, 1219–1233 (2011).
Matsui, M., Yamamoto, A., Kuma, A., Ohsumi, Y. & Mizushima, N. Biochem. Biophys. Res. Commun. 339, 485–489 (2006).
Morishita, H. et al. J. Biol. Chem. 288, 11436–11447 (2013).
Bu, L. et al. Nature Genet. 31, 276–278 (2002).
Hämälistö, S. et al. Nature Commun. 11, 229 (2020).
Sargeant, T. J. et al. Nature Cell Biol. 16, 1057–1068 (2014).
Cui, X. et al. Biochim. Biophys. Acta Gen. Subj. 1864, 129496 (2020).
Wang, F., Gómez-Sintes, R. & Boya, P. Traffic 19, 918–931 (2018).
Hummasti, S., Hong, C., Bensinger, S. J. & Tontonoz, P. J. Lipid Res. 49, 2535–2544 (2008).

 Read the paper: Organelle degradation in the lens by PLAAT phospholipases
Read the paper: Organelle degradation in the lens by PLAAT phospholipases
 Visionary stem-cell therapies
Visionary stem-cell therapies
 Mitochondria are mixed during cell division
Mitochondria are mixed during cell division