- TECHNOLOGY FEATURE
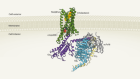
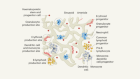
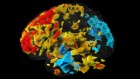
AI spots cell structures that humans can’t

Illustration by The Project Twins
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 592, 154-155 (2021)
doi: https://doi.org/10.1038/d41586-021-00812-7
References
Ounkomol, C., Seshamani, S., Maleckar, M. M., Collman, F. & Johnson, G. R. Nature Meth. 15, 917–920 (2018).
Christiansen, E. M. et al. Cell 173, 792–803 (2018).
Kandel, M. E. et al. Nature Commun. 11, 6256 (2020).
Nguyen, T. C. et al. J. Biomed. Opt. 25, 096009 (2020).
Wieslander, H., Gupta, A., Bergman, E., Hallström, E. & Harrison, P. J. Preprint at bioRxiv https://doi.org/10.1101/2021.01.18.427121 (2021).

 Deep learning takes on tumours
Deep learning takes on tumours
 Sharper signals: how machine learning is cleaning up microscopy images
Sharper signals: how machine learning is cleaning up microscopy images
 Deep learning for biology
Deep learning for biology
 NatureTech hub
NatureTech hub