- NEWS AND VIEWS
Multiple genomes give switchgrass an advantage
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 590, 394-395 (2021)
doi: https://doi.org/10.1038/d41586-021-00212-x
References
Casler, M. D. in Switchgrass (ed. Monti, A.) Ch. 2 (Springer, 2012).
Augustin, L. et al. Nature 429, 623–628 (2004).
Diffenbaugh, N. S. & Field, C. B. Science 341, 486–492 (2013).
Lovell, J. T. et al. Nature 590, 438–444 (2021).
Barabaschi, D. et al. Mol. Biotechnol. 50, 250–266 (2012).
Nieto Feliner, G., Casacuberta, J. & Wendel, J. F. Front. Genet. 11, 792 (2020).
Triplett, J. K., Wang, Y., Zhong, J. & Kellogg, E. A. PLoS ONE 7, e38702 (2012).

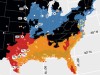 Read the paper: Genomic mechanisms of climate adaptation in polyploid bioenergy switchgrass
Read the paper: Genomic mechanisms of climate adaptation in polyploid bioenergy switchgrass
 From sea to sea
From sea to sea
 Keen insights from quinoa
Keen insights from quinoa