Infection with the SARS-CoV-2 coronavirus results in diverse outcomes for COVID-19, with the disease tending to be more severe and lethal for older males1,2. Yet some young people can also have severe COVID-19. What determines susceptibility to this disease? Writing in Science, Zhang et al.3 and Bastard et al.4 shed light on a key factor that affects whether life-threatening COVID-19 develops. The studies implicate deficiencies in interferon proteins, specifically, type I interferons (IFN-I). Such deficiencies might arise, as Zhang and colleagues report, through inherited mutations in genes encoding key antiviral signalling molecules, or, as Bastard and colleagues describe, by the development of antibodies that bind to and ‘neutralize’ IFN-I. Among people who developed severe COVID-19, such neutralizing antibodies were mostly in older males.
The IFN-I family includes IFN-α, IFN-β and IFN-ω. These molecules provide innate immune defences — they mount an initial rapid antiviral response. IFN-I proteins are a type of immune-signalling molecule called a cytokine; they are induced when a cell detects viral RNA through sensors, such as the proteins TLR3, TLR7 and TLR8 that are found in cellular organelles called endosomes. The IFN-I molecules then bind to the cell-surface receptor IFNAR (comprised of the proteins IFNAR1 and IFNAR2), resulting in the transcription of hundreds of genes5 that block the replication and spread of the virus.
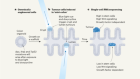
Zhang et al. examined whether people who had life-threatening COVID-19 pneumonia harboured mutations in genes that had previously been associated with severe cases of viral infections such as influenza. These genes belong to the TLR3 and IFN-I signalling pathways. The authors looked for mutations in 13 genes of interest. They found that 3.5% of the individuals (23 of the 659 people tested) had mutations in 8 of these genes, rendering the gene products incapable of producing or responding to IFN-I (Fig. 1).

Figure 1 | A defective antiviral signalling pathway. a, Normally, when the SARS-CoV-2 coronavirus enters human cells, it reaches an organelle called the endosome, where viral RNA is recognized by Toll-like receptors such as TLR7 and TLR3. This recognition drives a pathway (only some pathway proteins are shown) that leads to the expression of genes encoding type I interferon proteins. Zhang et al.3 found that people with severe COVID-19 had mutations in genes that encode components of this process; components associated with such mutations are shown in red. Such individuals do not produce interferon normally. A mutated version of the gene that encodes TLR7 has been reported5 previously in people with severe COVID-19. b, Zhang et al. also identified mutations in genes encoding the receptor for interferon (which consists of the proteins IFNAR1 and IFNAR2). Bastard and colleagues4 report that other individuals with severe COVID-19 have autoantibodies that bind to certain of the body’s type I interferons (IFN-α and IFN-ω, but not IFN-β), and thus block signalling mediated by IFN-α and IFN-ω. Such signalling defects hinder antiviral gene expression.
In vitro studies by Zhang and colleagues confirmed these findings, and indicated that the mutations produce ‘loss-of-function’ versions of proteins. The authors found that people carrying these mutations had low to undetectable levels of IFN-α in their blood plasma during coronavirus infection, linking the mutations to defective IFN-α production in response to viral challenge. By contrast, of 534 individuals with either asymptomatic or mild COVID-19, only one harboured a loss-of-function mutation at one of the 13 sites studied. This individual had a mutation in the IRF7 gene, which encodes a protein required for the production of IFN-I.
None of the people tested who had gene variants in the TLR3 or IFN pathway had previously had severe viral infections. This suggests that although SARS-CoV-2 antiviral defences might rely crucially on IFN-I, other types of viral infection can be controlled by alternative mechanisms in those individuals. Severe COVID-19 infection has also been reported in four young men who had a loss-of-function mutation in the TLR7 gene6, providing further evidence that genetic errors in IFN-I pathways contribute to severe COVID-19.
All eyes on a hurdle race for a SARS-CoV-2 vaccine
Another possible cause of interferon deficiency is the generation of antibodies that target IFN-I — a form of autoimmunity. An individual with an autoimmune disease has autoantibodies that target proteins naturally produced by the body. Autoantibodies that neutralize cytokines might therefore confer a similar susceptibility to infection to that seen in people who have genetic defects affecting cytokine pathways. Anti-IFN-I autoantibodies have been identified in various diseases, including in people with a condition called autoimmune polyglandular syndrome type 1 (APS-1). It was reported7 in June that an individual with APS-1 developed severe COVID-19 pneumonia. However, the roles of such autoantibodies in disease have not been explored in depth.
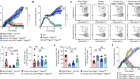
Remarkably, Bastard et al. report that, of the 987 people with severe COVID-19 whom they tested, 135 (13.7%) had antibodies that recognized an IFN-α subtype (IFN-α2), IFN-ω or both, whereas none of the 663 people with asymptomatic or mild COVID-19, and only 0.3% of healthy individuals examined (4 of 1,227) had such autoantibodies.
In addition, plasma samples in 10.2% of the 987 individuals with severe COVID-19 had interferon-neutralizing activity; the authors observed that this could hinder the ability of IFN-α2 to block in vitro infection of human cells with SARS-CoV-2. The finding demonstrates that these autoantibodies have the potential to affect the course of SARS-CoV-2 infection. Notably, 94% of the people who had anti-IFN-I antibodies were male, and they were generally older than most of the other individuals. Bastard and colleagues argue that the autoantibodies were present before the people were exposed to SARS-CoV-2, because these autoantibodies were detected early, within one to two weeks of infection. Furthermore, two of them had confirmed pre-existing autoantibodies against IFN-I.
What leads to the production of these autoantibodies? B cells of the immune system that make autoantibodies are normally selectively eliminated during development. The B cells that produce anti-cytokine autoantibodies in people with APS-1 arise as a result of defects in this selection process8. Thus, the anti-IFN-I antibodies found by the authors might arise as a consequence of faulty B-cell-tolerance checkpoints.
COVID-19 poses a riddle for the immune system
Why is autoantibody production skewed towards a greater occurrence in older men? Such faulty B-cell selection seems different from that of other autoimmune diseases, which tend to affect mainly females. Although it is known that the regulation of developing B cells is similar in young and middle-aged males and females, autoantibody levels in older people have not been investigated9. Many genes on the X chromosome encode molecules, such as FOXP3, BTK and CD40L, that are essential for immune responses and for early B-cell checkpoints9. Perhaps some such genetic mutation on the X chromosome favours the emergence of anti-cytokine autoantibodies. If so, males would be more vulnerable because they depend on a single copy of these genes on their X chromosome, unlike females, who have a back-up gene copy on a second X chromosome.
This leads to a central question. How does a defective IFN-I response lead to life-threatening COVID-19? The most direct explanation is that IFN-I deficiencies lead to uncontrolled viral replication and spread. However, IFN-I deficiencies might also have other consequences for immune-system function, such as the loss of suppression of immune-signalling complexes called inflammasomes and enhanced production of cytokines that are made downstream of these complexes10. Mice engineered to have abnormalities in the IFN-I pathway are more likely to die of influenza as a result of excessive inflammasome activation, not because of high levels of viral replication11, and such a phenomenon might explain severe COVID-19 in IFN-I-deficient people.
Individuals with genetic mutations in the IFN-I-induction pathway would therefore benefit from therapy that provides interferon, but such treatment would not help those with mutations in the genes encoding IFNAR. Furthermore, people who have neutralizing antibodies to IFN-α and IFN-ω might benefit from therapy that provides other types of interferon, such as IFN-β and IFN-λ, if given early during infection.
What other anti-cytokine autoantibodies might people carry? If others are found, it will be interesting to determine whether they also affect the course of infectious diseases. Might such autoreactive antibodies interfere with efforts to achieve vaccine-induced immunity in certain individuals? These latest results also suggest that blood samples (described as convalescent plasma) from people who have recovered from COVID-19, which can offer a source of antibodies targeting coronavirus, should be examined to exclude anti-IFN-I autoantibodies before being given as treatment.
As the search for effective treatment and vaccines continues, these key questions will help in refining the path forward. The new findings also highlight the need to examine both the genetic and the autoantibody-mediated contributors to severe cases of infectious disease.

 COVID-19 poses a riddle for the immune system
COVID-19 poses a riddle for the immune system
 All eyes on a hurdle race for a SARS-CoV-2 vaccine
All eyes on a hurdle race for a SARS-CoV-2 vaccine