- TECHNOLOGY FEATURE
Studying life at the extremes
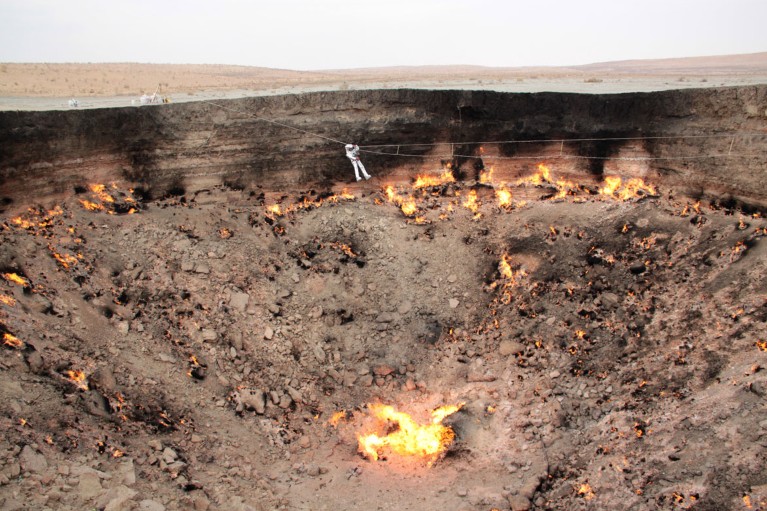

Researchers are looking for microbes in soil samples from this flaming gas crater in Turkmenistan, known as the Door to Hell. Credit: Stefan Green/XMP
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 587, 165-166 (2020)
doi: https://doi.org/10.1038/d41586-020-03055-0
References
Tighe, S. et al. J. Biomol. Tech. 28, 31–39 (2017).
Flusberg, B. A. et al. Nature Methods 7, 461–465 (2010).
Riley, L. A. et al. J. Ind. Microbiol. Biotechnol. 46, 1435–1443 (2019).
Walker, J. E. et al. Metab. Eng. Commun. 10, e00116 (2020).
Pulschen, A. A. et al. Curr. Biol. 30, 2852–2859 (2020).
Eun, Y.-J. et al. Nature Microbiol. 3, 148–154 (2018).
Turkowyd, B. et al. Preprint at bioRxiv https://doi.org/10.1101/2020.07.27.222935 (2020).

 The search for microbial dark matter
The search for microbial dark matter
 Scientists glimpse oddball microbe that could help explain rise of complex life
Scientists glimpse oddball microbe that could help explain rise of complex life
 Extremophiles: Life at the deep end
Extremophiles: Life at the deep end