- NEWS AND VIEWS
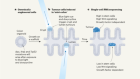
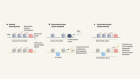
Tumour metabolites hinder DNA repair
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 582, 492-494 (2020)
doi: https://doi.org/10.1038/d41586-020-01569-1
References
Sulkowski, P. L. et al. Nature 582, 586–591 (2020).
King, A., Selak, M. A. & Gottlieb, E. Oncogene 25, 4675–4682 (2006).
Ye, D., Guan, K.-L. & Xiong, Y. Trends Cancer 4, 151–165 (2018).
Rose, N. R., McDonough, M. A., King, O. N. F., Kawamura, A. & Schofield, C. J. Chem. Soc. Rev. 40, 4364–4397 (2011).
Chowdhury, R. et al. EMBO Rep. 12, 463–469 (2011).
Xu, W. et al. Cancer Cell 19, 17–30 (2011).
Xiao, M. et al. Genes Dev. 26, 1326–1338 (2012).
Sun, Y. et al. Nature Cell Biol. 11, 1376–1382 (2009).
Cairncross, J. G. et al. J. Clin. Oncol. 32, 783–790 (2014).
Knijnenburg, T. A. et al. Cell Rep. 23, 239–254.e6 (2018).
Sulkowski, P. L. et al. Sci. Transl. Med. 9, eaal2463 (2017).
Inoue, S. et al. Cancer Cell 30, 337–348 (2016).

 Read the paper: Oncometabolites suppress DNA repair by disrupting local chromatin signalling
Read the paper: Oncometabolites suppress DNA repair by disrupting local chromatin signalling
 Metabolic vulnerability in tumours illuminated
Metabolic vulnerability in tumours illuminated
 One rogue agent suffices for genomic chaos
One rogue agent suffices for genomic chaos