- NEWS AND VIEWS
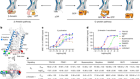
How sex chromosomes break up to get together
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 582, 346-347 (2020)
doi: https://doi.org/10.1038/d41586-020-01483-6
References
Kauppi, L. et al. Genes Dev. 27, 873–886 (2013).
Kauppi, L. et al. Science 331, 916–920 (2011).
Acquaviva, L. et al. Nature 582, 426–431 (2020).
Templado, C., Uroz, L. & Estop, A. Mol. Hum. Reprod. 19, 634–643 (2013).
Zickler, D. & Kleckner, N. Annu. Rev. Genet. 33, 603–754 (1999).
Boekhout, M. et al. Mol. Cell 74, 1053–1068 (2019).
Papanikos, F. et al. Mol. Cell 74, 1069–1085 (2019).
Baudat, F., Imai, Y. & de Massy, B. Nature Rev. Genet. 14, 794–806 (2013).
Brick, K., Smagulova, F., Khil, P., Camerini-Otero, R. D. & Petukhova, G. V. Nature 485, 642–645 (2012).
Thacker, D., Mohibullah, N., Zhu, X. & Keeney, S. Nature 510, 241–246 (2014).

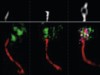 Read the paper: Ensuring meiotic DNA break formation in the mouse pseudoautosomal region
Read the paper: Ensuring meiotic DNA break formation in the mouse pseudoautosomal region
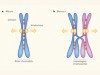 Hold on and let go
Hold on and let go