- NEWS AND VIEWS
Pancreatic cancer hidden in plain sight
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 581, 34-35 (2020)
doi: https://doi.org/10.1038/d41586-020-01103-3
References
Yamamoto, K. et al. Nature 581, 100–105 (2020).
Wolchok, J. Cell 175, 1452–1454 (2018).
O’Reilly, E. M. et al. JAMA Oncol. 5, 1431–1438 (2019).
Balachandran, V. P., Beatty, G. L. & Dougan, S. K. Gastroenterology 156, 2056–2072 (2019).
Yao, W., Maitra, A. & Ying, H. EBioMedicine 53, 102655 (2020).
Whittle, M. C. & Hingorani, S. R. Gastroenterology 156, 2085–2096 (2019).
Pommier, A. et al. Science 360, eaao4908 (2018).
Gettinger, S. et al. Cancer Discov. 7, 1420–1435 (2017).
Grasso, C. S. et al. Cancer Discov. 8, 730–749 (2018).
Waddell, N. et al. Nature 518, 495–501 (2015).
Ryschich, E. et al. Clin. Cancer Res. 11, 498–504 (2005).
Santana-Codina, N., Mancias, J. D. & Kimmelman, A. C. Annu. Rev. Cancer Biol. 1, 19–39 (2017).
Yang, S. et al. Genes Dev. 25, 717–729 (2011).
Bryant, K. L. et al. Nature Med. 25, 628–640 (2019).
Kinsey, C. G. et al. Nature Med. 25, 620–627 (2019).

 Read the paper: Autophagy promotes immune evasion of pancreatic cancer by degrading MHC-I
Read the paper: Autophagy promotes immune evasion of pancreatic cancer by degrading MHC-I
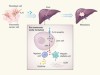 Molecular envoys pave the way for pancreatic cancer to invade the liver
Molecular envoys pave the way for pancreatic cancer to invade the liver
 New predictors for immunotherapy responses sharpen our view of the tumour microenvironment
New predictors for immunotherapy responses sharpen our view of the tumour microenvironment