Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 579, 183 (2020)
doi: https://doi.org/10.1038/d41586-020-00660-x
Updates & Corrections
-
Correction 11 March 2020: An earlier version of this story misstated a result from David Vessler's study on ACE2.
References
Walls, A. C. et al. Preprint at bioRxiv https://doi.org/10.1101/2020.02.19.956581 (2020).
Li, H. et al. Preprint at ChinaXiv http://chinaxiv.org/abs/202002.00062 (2020).
Jaimes, J. A., André, N. M., Millet, J. K. & Whittaker, G. R. Preprint at bioRxiv https://doi.org/10.1101/2020.02.10.942185 (2020).
Coutard, B. et al. Antiviral Res. https://doi.org/10.1016/j.antiviral.2020.104742 (2020).
Wrapp, D. et al. Science https://doi.org/10.1126/science.abb2507 (2020).

 Coronavirus latest: global infections pass 90,000
Coronavirus latest: global infections pass 90,000
 Time to use the p-word? Coronavirus enters dangerous new phase
Time to use the p-word? Coronavirus enters dangerous new phase
 When will the coronavirus outbreak peak?
When will the coronavirus outbreak peak?
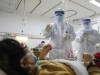 More than 80 clinical trials launch to test coronavirus treatments
More than 80 clinical trials launch to test coronavirus treatments